Quotation: Edwards F, Fantozzi G, Simon AY, Morretton J-P, Herbette A, Tijhuis AE, et al. (2024) Centrosome amplification primes ovarian most cancers cells for apoptosis and potentiates the response to chemotherapy. PLoS Biol 22(9):
e3002759.
https://doi.org/10.1371/journal.pbio.3002759
Educational Editor: Ana J. Garcia-Saez, Universitat zu Koln, GERMANY
Acquired: November 23, 2023; Accepted: July 17, 2024; Printed: September 5, 2024
Copyright: © 2024 Edwards et al. That is an open entry article distributed below the phrases of the Creative Commons Attribution License, which allows unrestricted use, distribution, and replica in any medium, offered the unique creator and supply are credited.
Information Availability: All related knowledge are throughout the paper and its Supporting Information file. Information underlying this research are from the Organic Useful resource Heart (BRC) of Institut Curie (certification quantity: 2009/33837.4; AFNOR NF S 96 90). The entry to the tumor cohort is restricted and subjected to the moral analysis by the Institut Curie. To inquiry about this cohort please seek the advice of https://institut-curie.org/, or contact the pinnacle of the pathology division at Institut Curie: Dr Anne Salomon: anne.salomon@curie.fr.
Funding: Labex CelTisPhyBio (https://labex-cellnscale.institut-curie.org/) offered the post-doctoral fellowship and Agence pour la Recherche contre le Most cancers (www.arcfoundation.org) offered the postdoctoral fellowship PDF20190508563 to FE. La Ligue contre le most cancers (www.ligue-cancer.net) PhD fellowship 17562 to GF. This work was supported by an InCA-Bio (www.e-cancer.fr) (2015‐PLBIO15‐237 grant, a WWC (www.worldwidecancerresearch.org) grant, (21-0042), an InCA (www.e-cancer.fr) (2021-1-PREV-Bio grant), the Institut Curie and the CNRS. The Basto lab is a member of the CelTisphybio labex (https://labex-cellnscale.institut-curie.org/). The funders didn’t have any function in research design, knowledge assortment and evaluation, resolution to publish, or preparation of the manuscript.
Competing pursuits: I’ve learn the journal’s coverage and the authors of this manuscript have the next competing pursuits: RB is a member of the PLOS Biology Editorial Board.
Abbreviations:
BRC,
Organic Useful resource Heart; CART,
Classification and Regression Timber; CNR,
centrosome to nuclei ratio; HGSOC,
high-grade serous ovarian most cancers; HPF,
high-power subject; MOMP,
mitochondria outer membrane permeabilization; MTOC,
microtubule organizing middle
Introduction
Centrosomes are the main microtubule organizing facilities (MTOCs) in proliferating animal cells, whose construction and quantity are tightly regulated through the cell cycle [1]. The centrosome is duplicated throughout S-phase in a PLK4 kinase-dependent method and the two centrosomes contribute to the well timed and useful meeting of a bipolar spindle throughout mitosis. In most cancers cell strains, centrosome structural and numerical defects are frequent [2] and specifically centrosome amplification—greater than 2 centrosomes per cell—has additionally been noticed in situ in tumor samples [3–5]. Cells with centrosome amplification carry out bipolar mitosis through centrosome clustering mechanisms throughout spindle meeting [6–10]. Centrosome amplification is nonetheless related to chromosome instability [11,12] and elevated invasive behaviors [13–15]. As first postulated by T. Boveri [16], centrosome amplification can drive tumorigenesis in vivo [7,17–19]. Defects in centrosome clustering capability are related to deadly multipolar divisions [8,11,20], motivating the seek for inhibitors that restrict centrosome clustering and eradicate cells with centrosome amplification [20–25].
Carboplatin and Paclitaxel are cytotoxic brokers utilized in remedy of assorted strong cancers, and their mixture is the usual of look after remedy of superior stage epithelial ovarian most cancers sufferers [26]. Carboplatin induces DNA injury and Paclitaxel stabilizes microtubules resulting in cell dying through mitotic disaster which is outlined as dying throughout or following irregular mitosis [27,28]. Regardless of the central function of centrosomes in spindle meeting, how centrosome amplification influences the response to mixed Carboplatin and Paclitaxel stays unexplored. Paclitaxel has been proven to induce multipolar divisions [29] and this may be elevated by centrosome amplification [30]. The influence of centrosome amplification on the response to DNA damaging brokers has nevertheless not been explored regardless of centrosomes regulating a number of signaling pathways that might affect chemotherapy responses [31–35]. A number of penalties of centrosome amplification might due to this fact synergize with mixed Carboplatin and Paclitaxel to induce environment friendly most cancers cell elimination.
Right here, we selected to check how centrosome amplification influences the response to mixed Carboplatin and Paclitaxel within the context of epithelial ovarian most cancers, a illness with poor scientific final result associated to late analysis, and frequent relapse [36]. Centrosome amplification is noticed in ovarian most cancers cell strains, and we lately additionally characterised its incidence in situ in affected person samples [5]. We use an inducible PLK4 overexpression system in ovarian most cancers cell strains to induce centrosome amplification in isogenic backgrounds. We carry out single-cell live-imaging of cells to evaluate the correlations between mitotic behaviors and cell destiny throughout chemotherapy. We present that centrosome amplification potentiates the response to mixed Carboplatin and Paclitaxel through a number of mechanisms. Past multipolar divisions related to Paclitaxel publicity, we discovered that centrosome amplification additionally enhances cell dying independently of mitotic behaviors. We present that centrosome amplification, though effectively tolerated by ovarian most cancers cells, results in mitochondria outer membrane permeabilization (MOMP) priming. We assess the extent of centrosome amplification in a beforehand characterised ovarian most cancers affected person cohort and observe an affiliation between excessive centrosome numbers and the affected person time to relapse in addition to their general survival. Collectively, our work reveals for the primary time that centrosome amplification can synergize with mixed chemotherapy, advancing our understanding of its penalties in most cancers.
Outcomes
Centrosome amplification enhances cell dying in response to mixed Paclitaxel and Carboplatin
To check the affect of centrosome amplification on the response to Carboplatin and Paclitaxel, we used an inducible PLK4 overexpression system (PLK4OE) within the epithelial ovarian most cancers cell line OVCAR8. Exposing cells to doxycycline for 72 h at 1 μg/ml induced centrosome amplification (greater than 2 centrosomes per cell) in round 80% of cells, in comparison with 4% in management OVCAR8 (DMSO handled, PLK4Ctl) (S1A and S1B Fig). We used MTT viability assays to find out Paclitaxel and Carboplatin IC50s over the 72 h following PLK4OE induction (S1C Fig). For each medicine, the IC50 was decrease for PLK4OE (67 μm and three,4 nM, respectively) in comparison with PLK4Ctl (136 μm and 5,1 nM, respectively), already suggesting that centrosome amplification sensitizes cells to those chemotherapeutic brokers. We additionally decided working mixture concentrations (100 μm Carboplatin + 3,3 nM Paclitaxel) that induce 60% development inhibition in PLK4Ctl and 89% in PLK4OE (S1D Fig).
We subsequent used live-imaging to research how centrosome amplification influences the response to chemotherapy and specifically if it will increase catastrophic mitosis. We used H2B-RFP expressing OVCAR8 permitting us to watch chromosome behaviors throughout mitosis and chromatin compaction that happens when cells die (Fig 1A and S1 Movie). We imaged cells through the 72 h of publicity to 100 μm Carboplatin + 3,3 nM Paclitaxel and carried out evaluation of full cell lineages, counting the variety of cells produced per lineage (proliferation) and the destiny of those cells (viability). In untreated OVCAR8 cells, PLK4OE lowered proliferation in comparison with PLK4Ctl, however independently of a rise in cell dying (Fig 1B and S2 Movie). Mixed chemotherapy lowered proliferation of each PLK4Ctl and PLK4OE cells and this discount was related to a rise in cell dying (Fig 1B and S3 Movie) in addition to cell cycle lengthening (S1E Fig). In settlement with the MTT dose-response assays, mixed chemotherapy induced the next proportion of cell dying in PLK4OE with 75% of cells dying, in comparison with PLK4Ctl the place 33% died (Fig 1B, 1E and 1F). By inspecting cell destiny in consecutive generations, we noticed that cell dying was primarily occurring in generations 2 and three suggesting that passage by way of mitosis or prolonged publicity time to chemotherapy is detrimental for the progeny (Fig 1C).
Fig 1. Centrosome amplification enhances cell dying in response to mixed chemotherapy.
(A) Single-cell live-imaging workflow. OVCAR8 cells expressing H2B-RFP and inducible for PLK4 overexpression are uncovered to DMSO or 1 μg/ml Doxycycline (DOX) for 72 h to induce centrosome amplification. PLK4Ctl and PLK4OE cells are then filmed throughout 72 h of chemotherapy and lineages are tracked over a number of generations. Consultant pictures of mitotic behaviors and cell fates are proven with the color-coded legends used within the subsequent panels. White arrows level to defects. Lineage evaluation consists in counting for every beginning cell, the variety of cells adopting the completely different fates (see panel B). Technology evaluation consists in figuring out the share of a technology that may undertake the completely different fates (see panel C). Mitosis and destiny correlation consist in figuring out the share of cells adopting the completely different fates, relying on the habits of the mom cell throughout mitosis (see panel D). (B) Bar graphs displaying the averages and SEM of the variety of cells per lineage adopting the indicated fates (legends in panel A). A minimal of 20 lineages had been analyzed from 2 unbiased experiments, statistical assessments: Fisher’s actual check on the variety of cell dying occasions (pooling dying in interphase and in mitosis). Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (C) Bar graphs displaying the common and SEM of the odds of cells present process indicated fates (legends in panel A). Two unbiased experiments, statistical assessments: Fisher’s actual check on the variety of cell dying occasions (pooling dying in interphase and dying in mitosis). Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (D) Vertical axis: Bar graphs displaying the common and SEM of the odds of mitotic phenotypes (legends in panel A); 137 and 133 cell divisions had been analyzed from 2 unbiased experiments, statistical check: Fisher’s actual check on the variety of multipolar divisions. Horizontal axis: Bar graphs displaying the common and SEM of the odds of cells present process indicated fates (legends in panel A) based on the mitotic habits of mom cells, with bar width relying on the proportion of cells displaying a given mitotic phenotype. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of No mis-segregation progeny (progeny of blue mitosis) dying in mitosis and interphase, p-value = 0,0005. (E, F) Single-cell profiles of PLK4Ctl (E) and PLK4OE (F) present process Carboplatin + Paclitaxel publicity. Every row corresponds to 1 cell (legends in panel A). Information for Fig 1 could be present in S1 Data.
We due to this fact characterised the mitotic behaviors within the first technology specializing in chromosome mis-segregation (Fig 1A and S4 Movie). In untreated PLK4Ctl OVCAR8, we noticed a big proportion of chromosome instability with round 20% of mitosis occurring with both chromosome mis-alignment, one lagging chromosome, or one chromatin bridge (Figs 1A, black arrows, S1F and S1G). These behaviors, that are generally tough to discriminate on the low spatiotemporal decision utilized in our long-term live-imaging strategy, had been pooled collectively and thought of as Slight mis-segregation occasions (Fig 1A). These occasions had been extra frequent in PLK4OE (36%), in settlement with centrosome amplification inducing merotelic attachments [11]. Nonetheless, the proportion of Multipolar divisions was negligible regardless of the presence of supernumerary centrosomes (S1F and S1G Fig). Importantly, mixed chemotherapy induced a rise in 2 behaviors related to robust chromosome mis-segregation: Multipolar divisions and bipolar divisions related to a number of chromosome mis-segregation occasions (combos of bridges, lagging and misaligned chromosomes—termed right here Excessive mis-segregation) (Fig 1A 1D vertical axis and 1E and 1F). Occasions of full division failure—both through cytokinesis failure, mitotic slippage, or dying in mitosis—had been noticed however remained rare upon mixed chemotherapy (Fig 1D, vertical axis). PLK4OE cells uncovered to mixed chemotherapy current near 47% of multipolar divisions in comparison with 6% noticed in PLK4Ctl (Fig 1D vertical axis and Fig 1E and 1F). We subsequent centered on the destiny of the progeny produced by the completely different cell division classes and noticed that inside cells finishing cell division, increased ranges of chromosome mis-segregation had been related to elevated cell dying within the progeny (Fig 1D, horizontal axis). Particularly, multipolar divisions had been related to at the very least 50% cell dying within the progeny, and the rise of those multipolar divisions in PLK4OE cells uncovered to mixed chemotherapy due to this fact contributes to the decreased viability noticed on this situation. Paclitaxel has been prompt to induce multipolar spindles in cells that current centrosome amplification [30], and exposing PLK4OE cells to Paclitaxel alone additionally induces multipolar divisions (S1H Fig), suggesting the elevated multipolarity we observe in presence of the mixed chemotherapy is attributable to Paclitaxel. Certainly, staining mitotic spindles for α-Tubulin and centrosome markers revealed that whereas bipolar and pseudo-bipolar spindles are essentially the most frequent inside untreated PLK4OE prometaphases and metaphases, Paclitaxel remedy induces the formation of extra multipolar spindles in addition to irregular wanting spindles that we named undefined spindles (S1I and S1J Fig).
Nonetheless, and surprisingly, we additionally noticed that independently of the kind of mitosis induced by mixed chemotherapy, the proportion of cell dying within the progeny was increased in PLK4OE in comparison with PLK4Ctl (Fig 1D, horizontal axis). This was specifically the case for the progeny of cells that don’t present any chromosome mis-segregation, the place 40% cell dying is noticed in PLK4OE in comparison with solely 12% cell dying in PLK4Ctl (Fig 1D, horizontal axis). These outcomes recommend that centrosome amplification will increase cell dying in response to mixed chemotherapy independently of multipolarity and chromosome segregation errors.
Centrosome amplification enhances cell dying in response to Carboplatin independently of catastrophic mitosis
We had been subsequent keen on understanding why cell dying is enhanced in PLK4OE in comparison with PLK4Ctl when related mitotic behaviors are noticed. We observed that the IC50 of Carboplatin is decrease for PLK4OE in comparison with PLK4Ctl (S1C Fig, proper). We due to this fact hypothesized that PLK4OE cells might reply in a different way to DNA injury induced by Carboplatin. We carried out live-imaging to higher perceive how OVCAR8 cells reply to 136 μm Carboplatin, and the way centrosome amplification modifies this response (S5 Movie). First specializing in the lineage evaluation, we noticed that Carboplatin-treated cells have a discount in proliferation in comparison with untreated cells and a rise in cell dying (Fig 2A–2D evaluate with S2F Fig for controls and S1E Fig). In settlement with PLK4OE cells being extra delicate to Carboplatin, fewer viable cells had been produced in Carboplatin uncovered PLK4OE in comparison with PLK4Ctl, and this was related to a rise in cell dying with 47% in Plk4OE and 30% in PLK4Ctl (Fig 2A–2D). Much like the mixed chemotherapy, most cell dying occurred in technology 2 (Fig 2E) and we due to this fact centered on chromosome mis-segregation through the first mitosis. In each PLK4Ctl and PLK4OE handled with Carboplatin, the principle phenotype was a rise in Excessive mis-segregation divisions primarily occurring after 24 h of Carboplatin publicity (Fig 2A, 2B, 2F and 2G evaluate with S2I Fig for untreated controls). In contrast to within the response to mixed chemotherapy, Multipolar divisions remained negligible (Fig 2A, 2B and 2F). We additional characterised mitosis with Excessive mis-segregation by staining cells mounted at 48 h of Carboplatin remedy, observing α-Tubulin in addition to centrosomal, centriolar, and centromeric markers. In settlement with our live-imaging experiment which characterised these divisions as being bipolar we noticed primarily bipolar spindles primarily based on the α-Tubulin staining, each in PLK4OE and in PLK4Ctl (S2A and S2B Fig). These spindles had been nevertheless regularly characterised by the presence of extra MTOCs even in PLK4Ctl cells that importantly don’t appear to contribute to forming useful spindle poles. In these cells, these extra MTOCs diversified primarily based on the presence of centrosomal and centriolar proteins (S2C Fig) and prompt Carboplatin could also be inducing centriole over-duplication in addition to untimely centriole disengagement as beforehand described for different sources of replication stress [37]. Regardless of bipolar spindle group, these mitosis had been characterised by in depth chromosome mis-segregation and in each PLK4OE and PLK4Ctl, a majority of those anaphases are characterised by the presence of acentric chromosomes (S2D and S2E Fig), and probably generated aneuploidy at deadly ranges.
On the low spatial and temporal decision used to correlate mitotic fates and cell dying, the proportions of the various kinds of mitosis had been related in Carboplatin handled PLK4OE and PLK4Ctl (Fig 2A, >2B and 2F, vertical axis). The elevated cell dying noticed in PLK4OE in comparison with PLK4Ctl once more appears to stem from a distinction within the affiliation between cell dying and mitotic phenotypes (Fig 2F, horizontal axis). Certainly, in PLK4Ctl, cell dying happens primarily within the progeny of Excessive mis-segregation mitosis, whereas in PLK4OE appreciable ranges of cell dying are additionally noticed following different mitotic behaviors (Fig 2A, 2B and 2F, horizontal axis). Particularly, round 35% of cell dying was noticed for the progeny of cells that had No mis-segregation or Slight mis-segregation defects, in distinction with PLK4Ctl the place lower than 10% of those cells died (Fig 2F, horizontal axis). In an effort to verify that our low spatiotemporal decision imaging set-up just isn’t hindering the detection of mitotic defects in PLK4OE that may clarify this elevated cell dying, we carried out live-imaging of Carboplatin-treated cells at increased spatiotemporal decision. This imaging was carried out through the first 24 h of Carboplatin remedy throughout which No mis-segregation or Slight mis-segregation occasions happen (Fig 2G). These occasions remained the dominating inhabitants noticed in each PLK4Ctl and PLK4OE, and Excessive mis-segregation of chromosomes remained rare within the first 24 h of Carboplatin remedy and at comparable ranges in PLK4OE and PLK4Ctl (S2F Fig). We had been capable of decide the proportions of lagging chromosomes, chromatin bridges, and misaligned chromosomes and noticed that PLK4OE primarily induces a nonsignificant tendency to extend chromosome mis-alignment in comparison with PLK4Ctl (S2G Fig). Nonetheless, this habits can also be frequent in untreated PLK4OE cells the place little cell dying is noticed (Fig 2D) suggesting it’s not chargeable for the noticed enhance in cell dying in response to Carboplatin. For the reason that anticipated penalties of PLK4OE in cells is to advertise multipolarity throughout mitosis, which can not have been detected on the low-resolution films or in mounted preparations, we examine whether or not multipolarity was detected in cells with further centrosomes handled with carboplatin or DMSO. To take action, we generated iOVCAR8 cells expressing GFP-Tubulin and H2B-RFP to observe concomitantly mitotic spindle and chromosome habits. Imaging cells at increased decision and at brief time intervals in time-lapse approaches revealed nevertheless, that the massive majority of each PLK4OE and PLK4OE carboplatin-treated cells divide in a bipolar method (S2H and S2I Fig). As seen in mounted cells, bipolar spindles include clustered and un-clustered centrosomes and divide with/with out chromosome mis-segregation errors (S2H Fig, top-second and third rows). Multipolar divisions are rare and fascinating even introduced centrosome clustering of sure further centrosomes (S2H Fig, fourth row). It’s price noticing that sure cell divisions had been extraordinarily lengthy, which can even be influenced by the co-expression of GFP-tubulin and H2B. Lastly, a rise in cell dying is noticed on this cell line, suggesting that GFP-Tubulin expression might sensitize these cells to the results of centrosome amplification and carboplatin (S2H Fig, fifth row and S2I Fig).
The temporal sample of mitotic phenotypes with Excessive mis-segregation occasions occurring when mitosis is initiated after an extended time spent in Carboplatin (Fig 2A and 2G) prompt an affect of the cell cycle on Carboplatin responses. We thus analyzed the cell cycle in response to Carboplatin utilizing a FUCCI expressing OVCAR8 cell line. This technique allowed us to discriminate cells in G1 from cells in S/G2 (Figs 2H, 2I, S3A and S3B), whereas nonetheless observing cell dying occasions which happen primarily in S/G2 (S3C Fig). First, we noticed that Carboplatin induced a rise in S/G2 size, suggesting DNA injury in OVCAR8 cells prompts the intra S and/or the G2/M checkpoint (S3D Fig), whereas G1 size was not strongly various in any noticed situation (S3E Fig). Subsequent, we noticed that S/G2 lengthening and subsequent delayed mitotic entry occurred primarily in cells that had been in G1 on the onset of Carboplatin publicity, and due to this fact in cells uncovered to Carboplatin for a whole S-phase (Fig 2H and 2J). Regardless of this delay, these cells probably entered mitosis with unrepaired injury, driving Excessive mis-segregation of chromosomes throughout mitosis and finally resulting in dying of the progeny. In PLK4OE cells uncovered to Carboplatin, the timing and proportions of mitotic and cell cycle behaviors was much like PLK4Ctl. Excessive mis-segregation occasions occurred in cells with a robust delay in mitotic entry (Fig 2B, 2F and 2G), which concurred with cells that had been in G1 on the onset of Carboplatin publicity (Fig 2I and 2J). We did word a slight lengthening in G1 period in Carboplatin-treated PLK4OE cells in comparison with PLK4Ctl (S3E Fig), however we didn’t observe a big affiliation between cell dying and G1 size (S3F Fig).
These observations due to this fact recommend that PLK4Ctl cells are primarily killed by catastrophic mitosis induced by excessive ranges of DNA injury and subsequent Excessive mis-segregation of chromosomes. PLK4OE cells then again additionally die following mitosis characterised by No or Slight mis-segregation of chromosomes occurring early upon Carboplatin remedy, suggesting they die independently of deadly mitotic behaviors. In settlement with cell dying occurring independently of mitosis in PLK4OE, 17% cells died within the first technology in comparison with solely 6% in PLK4Ctl (Fig 2A, 2B and 2E). Particularly, cells that had been in G1 in the beginning of Carboplatin publicity and due to this fact can accumulate DNA injury of their first cell cycle, had been preferentially killed in PLK4OE with 25% of cells dying in comparison with 8% in PLK4Ctl (Fig 2H, 2I and 2K). Our findings suggest that centrosome amplification sensitizes cells to the impact of Carboplatin in a single cell cycle and independently of catastrophic mitotic behaviors.
The centrosome has beforehand been concerned in regulating the DNA injury response through recruitment of ATR, ATM, Chk1, and Chk2 [31,32]. We investigated whether or not centrosome amplification modifies the signaling downstream of DNA injury, explaining elevated cell dying in response to Carboplatin. We detected phosphorylation and activation of Chk1 and p53 in response to Carboplatin, however no distinction is noticed in PLK4OE in comparison with PLK4Ctl cell extracts (S3G and S3H Fig). In settlement with DNA injury ranges and response being unchanged by PLK4OE, we additionally noticed no distinction within the ranges of staining for the early marker of DNA injury response γ-H2AX, or for the recruitment of DNA injury restore elements FANCD2, 53BP1, and Rad51 (S3I and S3J Fig). Collectively, these observations recommend that centrosome amplification will increase cell dying in response to DNA injury, independently of catastrophic mitosis, but additionally independently of the DNA injury response.
Fig 2. Centrosome amplification enhances cell dying in response to Carboplatin independently of chromosome mis-segregation.
(A, B) Single-cell profiles of PLK4Ctl (A) and PLK4OE (B) present process Carboplatin publicity. For comparability with untreated controls—S2F Fig. Shade coding of mitosis and fates legends in panel C. (C) Legends for panels A, B, D, E, F, and G, as outlined in Fig 1A. (D) Bar graphs displaying the averages and SEM of the variety of cells produced per lineage, adopting the indicated fates (legends in panel 1A). A minimal of 31 lineages had been analyzed from 2 unbiased experiments, statistical assessments: Fisher’s actual check on the variety of cell dying occasions (pooling dying in interphase and in mitosis). Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (E) Bar graphs displaying the common and SEM of the odds of cells present process indicated fates (legends in Fig 1A and 1C). Two unbiased experiments, statistical assessments: Fisher’s actual check on the variety of cell dying occasions (pooling dying in interphase and dying in mitosis). Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (F) Vertical axis: Bar graphs displaying the common and SEM of the odds of mitotic phenotypes (legends in Fig 1A and 1C); 142 and 146 cell divisions analyzed from 2 unbiased experiments, statistical check: Chi-square check. Horizontal axis: Bar graphs displaying the averages and SEM of the odds of cells present process indicated fates (legends in Fig 1A and 1C) based on the mitotic habits of the mom cell, with bar width relying on the proportion of cells based on their mitotic habits. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of No mis-segregation progeny (progeny of blue mitosis) dying in mitosis and interphase, p-value = 0,0003. (G) Scatter dot plot graphs displaying time of mitotic entry, with median and interquartile vary. Cells had been labeled relying on mitotic phenotypes with color-code outlined in panel C. Two unbiased experiments with a minimal of 48 mitosis analyzed per class. Statistical assessments: Kruskal–Wallis with Dunn’s a number of comparisons assessments. (H, I) Single-cell profiles of FUCCI PLK4Ctl (H) and PLK4OE (I) cells present process Carboplatin publicity. Instances in G1, S/G2 and mitosis, in addition to dying in every of those phases are color-coded as indicated. (J) Scatter dot plots of the time of mitotic entry relying on cell cycle section at film begin, with median and interquartile vary. Two unbiased experiments with a minimal of 10 instances analyzed per class. Statistical assessments: Kruskal–Wallis with Dunn’s a number of comparisons assessments. (Ok) Bar graphs displaying the common and SEM of the share of cells adopting the indicated fates (legends in panel I). Two unbiased experiments, statistical check: Fischer’s actual check on the variety of cells dying (regardless of the cell cycle section). Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. Information for Fig 2 could be present in S2 Data.
Centrosome amplification modulates mitochondrial apoptosis independently of p53 and the PIDDosome
In an effort to higher perceive how centrosome amplification will increase cell dying in response to Carboplatin independently of mitotic errors, we characterised the kind of cell dying and the related signaling community. We noticed that Carboplatin remedy induced apoptosis characterised by Caspase-3 cleavage (Fig 3A and 3B), in addition to cells changing into optimistic for Annexin-V by move cytometry, which was fully suppressed by the pan-caspase inhibitor Q-VD-Oph (Fig 3C). PLK4OE induces each a untimely cleavage of Caspase-3 (t = 48 h in comparison with t = 72 h in PLK4Ctl), in addition to a rise within the Annexin-V optimistic cell inhabitants (53% in comparison with 32% in PLK4Ctl), confirming that centrosome amplification potentiates the apoptotic response to Carboplatin (Fig 3A–3C). To find out if the MOMP-dependent apoptotic pathway was activated in response to Carboplatin, we stained cells for Cytochrome C with a purpose to observe its launch from mitochondria (Fig 3D). In untreated cells, Cytochrome C was detected in punctate buildings all through the cytoplasm, in settlement with its mitochondrial localization. Upon Carboplatin publicity, Cytochrome C staining remained much like untreated cells, whereas we noticed lifeless cell remnants characterised by condensed DNA (Fig 3D, white arrows and 3E). Nonetheless, if launch of Cytochrome C occurred, it might result in fast apoptosis initiation and detachment of the cells, precluding the remark of cells through immunofluorescence. We ctherefore used the pan-caspase inhibitor Q-VD-Oph to inhibit the effector steps of apoptosis downstream of Cytochrome C launch, and noticed a inhabitants of cells the place Cytochrome C is diffuse within the cytoplasm and nucleus (Fig 3D, pink arrows and 3E). MOMP, Cytochrome C launch and caspase activation due to this fact happens in response to Carboplatin, suggesting that mitochondrial apoptosis is the principle cell dying mechanism at play.
Fig 3. Centrosome amplification enhances mitochondrial outer membrane permeabilization in response to Carboplatin.
(A) Consultant western blot detecting Caspase-3 cleavage. (B) Common and SEM of cleaved caspase 3 protein ranges from 2 unbiased experiments, normalized to ranges measured in Carboplatin handled PLK4Ctl cells at 72 h. (C) Bar graphs displaying the common and SEM of the share of cells in specified Annexin V-APC/PI gates analyzed by move cytometry. 4 replicates obtained from 2 unbiased experiments, with a minimal of 60,000 cells analyzed per situation and replicate. Statistical check: comparability of the share of Annexin V optimistic cells, utilizing ANOVA with Sidak’s a number of comparability check. Consultant cytometry profiles could be discovered within the S1_Appendix. (D) Consultant pictures of cells labeled with DAPI (grey) and antibodies in opposition to Cytochrome C (cyan) and centrosomes in pink. White arrows point out lifeless cell particles, pink arrows point out cells which have launched Cytochrome C within the cytoplasm, M signifies mitotic cells. Consultant insets are proven for particular person Q-VD-Oph handled cells. (E) Bar graphs displaying the common and SEM of the odds of indicated cell populations. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of cells releasing Cytochrome C. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (F) Left: Bar graphs displaying the common and SEM of the odds of cells with indicated nuclear phenotypes, throughout the populations releasing Cytochrome C. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of cells with fragmented nuclei. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. Proper: Consultant pictures of DAPI-stained nuclei with indicated phenotypes. Information for Fig 3 could be present in S3 Data.
Importantly, we noticed 27% of cells releasing Cytochrome C in PLK4OE cells in comparison with solely 12% in PLK4Ctl cells, suggesting that Carboplatin induces a stronger activation of apoptosis within the presence of centrosome amplification (Fig 3D and 3E). In settlement with completely different stresses resulting in apoptosis between PLK4Ctl and PLK4OE, we noticed that 36% of the cells that launched Cytochrome C introduced fragmented nuclei in PLK4Ctl in comparison with 5% in PLK4OE (Fig 3D and 3F). These fragmented nuclei are symptomatic of excessive chromosome mis-segregation throughout mitosis, supported by PLK4Ctl cells being killed by catastrophic mitosis, whereas centrosome amplification potentiates apoptosis independently of catastrophic mitosis in PLK4OE.
The canonical intrinsic apoptosis pathway linking the DNA injury response to MOMP happens through p53 stabilization which then drives the transcription of pro-apoptotic BCL2 household genes [38]. Centrosome amplification has been linked to p53 stabilization through PIDDosome activation, which relies on centriole distal appendage grouping. This results in Caspase-2 cleavage and activation and cleavage of MDM2—a serious p53 regulator—[34,35]. OVCAR8 cells have a mutant TP53 gene which results in different splicing of exon5, and a 6 amino-acid deletion in p53’s DNA-binding area [39]. We noticed that p53 protein is current, phosphorylated, and accumulates in response to Carboplatin (S3G Fig, quantified in S4B Fig). The deletion in p53’s DNA injury binding area is usually recommended to preclude its transcriptional actions [39], and in settlement with this, we noticed that its transcriptional targets p21 and PUMA don’t present a putting enhance upon Carboplatin publicity (S4A and S4B Fig). We nonetheless assessed p53’s involvement in Carboplatin-induced cell dying utilizing shRNA, and confirmed that p53 was dispensable in each PLK4Ctl and PLK4OE in response to Carboplatin (S4C and S4D Fig). p21 is finest characterised for its features in cell cycle arrest and apoptosis inhibition, however has additionally been proven to up-regulate apoptosis [40]. As we observed p21 to be up-regulated in PLK4OE cells in comparison with PLK4Ctl (S4A and S4B Fig), we examined its contribution to cell dying, however noticed no impact of pulling down the p21 coding gene CDKN1a (S4C and S4D Fig). Regardless of apoptosis being p53 unbiased in Carboplatin handled OVCAR8, we examined whether or not PIDDosome activation might contribute to enhancing intrinsic apoptosis in PLK4OE. Certainly, upon PLK4OE, we noticed cleavage of Caspase-2 and MDM2 reflecting PIDDosome activation (S4E and S4F Fig). We knocked-down the distal appendage protein required for PIDDosome activation ANKRD26 and noticed a robust discount of Caspase-2 and MDM2 cleavage in PLK4OE, reflecting environment friendly PIDDosome silencing (S4G Fig). Nonetheless, this had no impact on the improved cell dying noticed upon Carboplatin publicity in PLK4OE, suggesting that centrosome amplification potentiates apoptosis independently of the PIDDosome (S4H Fig). Altogether, our outcomes present that centrosome amplification results in enhanced apoptosis in response to Carboplatin unbiased of beforehand described centrosome signaling nodes.
Centrosome amplification primes for MOMP and sensitizes cells to a range of chemotherapies
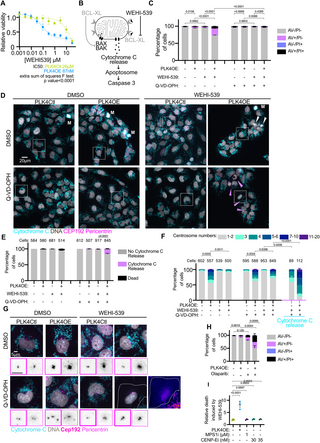
To date now we have noticed that PLK4OE cells execute apoptosis sooner and to the next proportion, in response to a degree of stress (mitotic behaviors, DNA injury ranges and DNA injury response) which isn’t any completely different than in PLK4Ctl. This due to this fact prompt that these cells could also be primed for MOMP, which means that the steadiness between pro-apoptotic and anti-apoptotic BCL2 household proteins that decide the exercise of the mitochondrial pore forming proteins BAX and BAK is tilted in direction of their activation in PLK4OE [41]. To check this chance, we carried out MTT dose-response assays throughout 72 h to medicine which mimic the exercise of particular pro-apoptotic BH3-only BCL2 household proteins. Strikingly, we noticed that PLK4OE induces a robust sensitization to WEHI-539—a selected BCL-XL inhibitor—(Fig 4A and 4B), however to not Venetoclax or A1210477—particular inhibitors of BCL-2 and MCL-1, respectively—(S5A and S5B Fig). We confirmed that 72 h 300 nM WEHI-539 publicity selectively induced apoptosis in PLK4OE cells in a Caspase dependent method, through Annexin V and PI cytometry (Fig 4C). We additionally noticed launch of Cytochrome C from mitochondria in WEHI-539-treated PLK4OE cells upon pan-caspase inhibition, confirming that BCL-XL inhibition induces MOMP particularly in PLK4OE (Fig 4D and 4E). Counting centrosomes in PLK4OE cells revealed that the 72 h 300 nM WEHI-539 remedy lowered the proportion of cells with further centrosomes to the extent noticed in PLK4Ctl cells (Fig 4F and 4G). These outcomes recommend environment friendly killing of cells with centrosome amplification, which is suppressed upon pan-caspase inhibition. We had been additionally capable of rely centrosomes within the inhabitants of cells that launch Cytochrome C from mitochondria and are due to this fact poised to die, revealing that almost all of those cells have centrosome amplification in WEHI-539 handled PLK4OE cells (Fig 4F). Apparently, an especially small proportion (1%) of PLK4Ctl cells additionally launch Cytochrome C in response to WEHI-539 (Fig 4E) and counting centrosomes in these cells revealed the next degree of centrosome amplification than the untreated PLK4Ctl inhabitants (Fig 4F). This implies that independently of induced PLK4 overexpression, centrosome amplification primes for MOMP in OVCAR8 cells. Sadly, SAS-6 overexpression as a substitute imply to induce centrosome amplification solely result in a weak enhance within the variety of cells with greater than 2 centrosomes and we due to this fact didn’t check if apoptotic priming happens on this system (S5C Fig). Nonetheless, we noticed that in parental OVCAR8 cells devoid of the inducible PLK4OE transgene, cells with supernumerary centrosomes are preferentially killed by WEHI-539 suggesting this apoptotic sensitization happens in cells with “naturally” occurring centrosome amplification (S5D Fig).
Fig 4. Centrosome amplification primes for mitochondria outer membrane permeabilization.
(A) Dose-response of PLK4Ctl and PLK4OE cells to WEHI-539 normalized to their respective untreated circumstances, obtained from MTT viability assays. Imply and SEM of three unbiased experiments every obtained from averaging 3 technical replicates. (B) Schematic of the induction of apoptosis by WEHI-539. When BAX and BAK channel formation are inhibited by BCL-XL, Cytochrome C is current within the mitochondria intermembrane house (left). WEHI-539 inhibits BCL-XL which relieves the inhibition of channel formation by BAX and BAK, resulting in Cytochrome C launch, Apoptosome activation, and cleavage of Caspase 3 (proper). (C) Bar graphs displaying the common and SEM of the share of cells in specified Annexin V-APC/PI gates analyzed by move cytometry. 4 replicates obtained from 2 unbiased experiments with a minimal of 15,000 cells analyzed per situation and replicate. Statistical check: comparability of the share of Annexin V optimistic cells, utilizing ANOVA with Sidak’s a number of comparability check. Consultant cytometry profiles could be discovered within the S1 Appendix. (D) Consultant pictures of cells labeled with DAPI (grey) and antibodies in opposition to Cytochrome C (cyan), CEP192 (magenta), and Pericentrin (magenta). White arrows point out lifeless cell particles, pink arrows point out cells which have launched Cytochrome C in to the cytoplasm, M signifies mitotic cells. Consultant insets are proven in panel (G). (E) Bar graphs displaying the common and SEM of the odds of indicated cell populations. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of cells releasing Cytochrome C. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (F) Bar graphs displaying the common and SEM of the share of cells with the indicated variety of centrosomes (decided by the co-localization of CEP192 and Pericentrin). Two unbiased experiments, statistical check: comparability of the share of cells with greater than 2 centrosomes, utilizing ANOVA with Sidak’s a number of comparability check. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (G) Insets from panel (D) with inverted grayscale insets zooming on the centrosomes displaying CEP192 (tick magenta border) and Pericentrin (mild magenta border). For PLK4OE, WEHI-539, Q-VD-OPH a “Hearth-lut” inset is proven. (H) Bar graphs displaying the common and SEM of the share of cells in specified Annexin V-APC/PI gates analyzed by move cytometry. 4 replicates obtained from 2 unbiased experiments with a minimal of 10,000 cells analyzed per situation and replicate. Statistical check: comparability of the share of Annexin V optimistic cells, utilizing ANOVA with Sidak’s a number of comparability check. Consultant cytometry profiles could be discovered within the S1 Appendix. (I) Scatter dot plots displaying the ratio between the odds of Annexin V optimistic cells noticed in presence and absence of WEHI-539 300 nM. Common and SEM of three replicates from 2 unbiased experiments. Statistical check: ANOVA with Dunnett’s a number of comparability check. Information for Fig 4 could be present in S4 Data.
We subsequent examined if related results is likely to be noticed in different cell strains and established inducible centrosome amplification through PLK4OE in ovarian most cancers cell strains COV504 and SKOV3 (S5E and S5F Fig). We examined if MOMP priming recognized in OVCAR8 was additionally noticed in these cell strains (S5G Fig). We used the much less particular BH3-mimetic Navitoclax (inhibitor of BCL2, BCL-XL, and BCL-W), because the dependency on BCL-XL in PLK4OE OVCAR8 is likely to be reflecting OVCAR8 apoptotic wiring somewhat than a selected impact of centrosome amplification on BCL-XL. We noticed that Navitoclax lowered the viability of PLK4OE cells preferentially in comparison with PLK4Ctl cells in COV504 (EC80 of 400 nM and three,6 μm, respectively) though to a lesser extent than in OVCAR8 (EC80 of 40 nM and a couple of,2 μm, respectively). Priming was nevertheless not noticed in SKOV3. We then established the IC50s for Carboplatin and Paclitaxel, and decided mixture concentrations in PLK4Ctl cells (S5H Fig). We used Trypan blue assays to find out viability in response to chemotherapy and confirmed this strategy in OVCAR8 by displaying that viability decreases extra in PLK4OE in comparison with PLK4Ctl (S5I Fig, high). Apparently, we noticed a gradation within the enhanced cell dying induced by centrosome amplification within the completely different cell strains, with the strongest impact noticed in OVCAR8 (S5I Fig, high), the weakest in SKOV3 (S5I Fig, backside), and an intermediate impact in COV504 (S5I Fig, center), which could be put in perspective with the noticed gradation in MOMP priming. For responses to Paclitaxel nevertheless now we have to keep in mind the truth that multipolar divisions are additionally elevated in COV504 PLK4OE cells (S5J Fig).
Our identification of apoptotic priming in cells with centrosome amplification means that it is likely to be related to enhanced cell dying in response to a bigger panel of medication. We due to this fact examined whether or not centrosome amplification sensitizes ovarian most cancers cells to PARP inhibitors which are actually included in commonplace of care protocols in epithelial ovarian most cancers [42], specializing in Olaparib (IC50s decided and introduced in S5H Fig). Trypan Blue viability assays in OVCAR8 and COV504 confirmed that PLK4OE results in lowered viability in comparison with PLK4Ctl in response to Olaparib (S5K Fig). This impact was not noticed in SKOV3 wherein now we have not noticed MOMP priming linked to centrosome amplification. Utilizing move cytometry to detect Annexin V optimistic cells, we confirmed that in OVCAR8 Olaparib certainly induces extra apoptosis in PLK4OE than PLK4Ctl (Fig 4H). Collectively, our outcomes recommend that centrosome amplification enhances cell dying in response to chemotherapy differentially relying on the cell line and that centrosome amplification related apoptotic priming can sensitize to a range of chemotherapies.
Centrosome amplification primes for MOMP independently of chromosome instability or lengthened mitosis
Centrosome amplification induces a rise in chromosome instability and a spindle-assembly checkpoint-dependent extension of mitosis period [7,8,11], each of which we noticed in PLK4OE OVCAR8 cells (S1E and S1G Fig). Apoptotic priming and specifically sensitization to BCL-XL inhibition has beforehand been linked to mitotic defects. Particularly chromosome instability, micronuclei formation and cGAS/STING signaling can drive a transcriptional response that drives apoptosis or priming [43,44]. Alternatively, prolonged mitotic period can result in the proteosomal degradation of anti-apoptotic BCL2 household proteins, resulting in BCL-XL sensitization [45–48]. We had been due to this fact keen on figuring out if apoptotic priming noticed in response to centrosome amplification is attributable to cumulated mitotic stress in these cells.
First, we aimed to higher characterize the mitotic stress induced by centrosome amplification within the already chromosomally instable OVCAR8 cell line (S1G Fig). We used single-cell DNA-sequencing to evaluate karyotype heterogeneity and noticed scores of 0,119 in PLK4Ctl, 0,137 in PLK4OE cells, and 0,283 in PLK4Ctl cells handled with 1 μm of the MPS1 inhibitor AZ3146 as a optimistic management of chromosome mis-segregation (S6A Fig). PLK4OE due to this fact solely mildly elevated aneuploidy, in step with ranges of chromosome-mis-segregation noticed by time-lapse imaging (S6B Fig). We additionally assessed the extent of mitotic lengthening induced in PLK4OE cells (S6C Fig) and noticed it was delicate (median = 60 min in PLK4OE and median = 35 min in PLK4Ctl) in comparison with that induced by low doses of the CENP-E inhibitor GSK923295 (median = 100 min at 30 nM and 175 min at 35 nM), regardless of ranges of chromosome mis-segregation being related (S6B Fig).
Mitotic stress is due to this fact delicate in PLK4OE in comparison with the opposite perturbations we examined, however we had been nonetheless keen on figuring out if it contributes to apoptotic priming. We weren’t capable of cut back mitotic period in PLK4OE cells through spindle meeting checkpoint inhibition with out inducing a robust enhance in multipolar divisions, so we used MPS1 and CENP-E inhibition to imitate mitotic stress noticed in PLK4OE. We pretreated PLK4Ctl cells with inhibitors throughout 72 h earlier than including WEHI-539 for a further 24 h. In PLK4OE cells, this result in 33% Annexin V optimistic cells, whereas it solely induces 9% in response to MPS1 inhibition (S6D Fig), making it unlikely that priming happens in response to chromosome instability in PLK4OE. In CENP-E inhibition pretreated cells at 30 nM and 35 nM, WEHI-539 induces 12% and 25% Annexin V cells, respectively, in step with mitotic lengthening inducing priming [48]. Importantly nevertheless, 35 nM CENP-E inhibition pretreatment already induces 12% Annexin V optimistic cells which is appreciable in comparison with the 4% noticed in PLK4OE (S6D Fig), and probably is defined by the in depth mitotic lengthening noticed in response to 35 nM CENP-E inhibition (S6C Fig). This ends in the ratio of cell dying induced by WEHI-539 relative to the basal noticed degree of cell dying to be comparable in PLK4Ctl cells and CENP-E inhibition pretreated cells (round 2-fold). In distinction, the ratio of cell dying was a lot increased and near 8-fold enhance upon PLK4OE (Fig 4I). Due to this fact, the priming induced in PLK4OE stands out from that induced by different sources of mitotic stress in that PLK4OE cells are viable however strongly depending on BCL-XL. These outcomes additionally recommend that the mixture of chromosome instability and mitotic lengthening just isn’t the main contributor to MOMP priming upon centrosome amplification in OVCAR8 cells.
To determine transcriptomic signatures which will affect cell dying responses in cells with further centrosomes, we used bulk RNAseq evaluating PLK4OE and PLK4Ctl OVCAR8 cells. A powerful inflammatory signature in PLK4OE (S6E Fig) was recognized, and we additionally noticed STING phosphorylation (S6F Fig), suggesting that the cGAS/STING pathway might form the transcriptional response to centrosome amplification. We had been due to this fact keen on immediately testing if cGAS/STING signaling would possibly contribute to priming, though this appeared unlikely as CENP-E and MPS1 inhibition additionally activate STING (S6F Fig) however usually are not related to priming. We used a bulk LentiCRISPR knock-out strategy of STING, however noticed no affect on PLK4OE cells sensitivity to WEHI-539 (S6G and S6H Fig).
We due to this fact determine that centrosome amplification in OVCAR8 results in MOMP priming which is revealed by a selective sensitization to the BCL-XL inhibitor WEHI-539. Evaluating with different mitotic perturbations, we conclude that the centrosome amplification related priming is unbiased of mitotic lengthening and chromosome instability.
Excessive centrosome numbers are related to a greater response to chemotherapy in a excessive grade serous ovarian most cancers affected person cohort
To evaluate if centrosome amplification is related to chemotherapy responses in sufferers, we turned to the characterization of centrosomes we beforehand carried out in situ in treatment-naive epithelial ovarian tumors (Cohort description in S1 Table). Right here, centrosomes had been detected because the colocalization of Pericentrin and CDK5RAP2 in confocal pictures of methanol-fixed affected person tissue sections [5]. To evaluate centrosome numbers in samples we outlined the centrosome to nucleus ratio (CNR) because the variety of centrosomes detected in a subject by the variety of nuclei which we averaged over 10 fields per affected person (S7A Fig). In wholesome tissues obtained from prophylactic oophorectomy or hysterectomy, the CNR was 1.02 ± 0.02 suggesting cells have on common 1 centrosome per cell which is anticipated for a non-proliferative tissue. On common, the CNR in tumor tissues was 1.43 ± 0.04, with the minimal at 0.61 and most at 2.55. Whereas solely 9% of tumors had a CNR above 2, suggesting that pervasive centrosome amplification—when outlined by the presence of greater than 2 centrosomes per cell—is rare, 89% of the tumors introduced a CNR superior to the common CNR present in wholesome tissues. Contemplating that the CNR didn’t correlate with proliferation as established by the mitotic index (S7B Fig) and Ki67 staining (S7C Fig), centrosome amplification might contribute to this enhance in CNR in tumors in comparison with wholesome samples. Certainly, we did observe nuclei related to greater than 2 centrosomes, though the tumor-wide frequency of such occasions reached a most of three,2% in our earlier research [5].
We subsequent examined if the CNR was related to chemotherapy responses limiting our evaluation to the high-grade serous ovarian cancers (HGSOCs) in our cohort. We dichotomized our inhabitants into 2 teams utilizing the Classification and Regression Timber (CART) methodology. Making an allowance for the binary final result “relapse inside 6 months or no relapse inside 6 months,” this methodology resulted within the categorization of the cohort into 55 low CNR (≤1.45) and 33 excessive CNR (>1.45). Importantly, we noticed no affiliation between CNR and FIGO stage. Most sufferers on this cohort—59%—are stage III sufferers and comprise each excessive and low CNRs (S7D Fig). We subsequent plotted HGSOC affected person survival curves based on the CNR standing. We discovered that prime CNR was related to higher general survival (S7E Fig). These outcomes recommend that regardless of its oncogenic potential [7,17–19], centrosome amplification would possibly enhance affected person prognosis at the very least in ovarian most cancers. This puzzling remark may very well be defined if excessive CNR promotes chemotherapy responses as general survival knowledge displays sufferers full scientific course which incorporates Carboplatin and/or Paclitaxel remedy for 84% of sufferers on this cohort. To immediately assess a hyperlink between CNR and chemotherapy responses, we plotted affected person time to relapse and located that prime CNR was related to an extended time to relapse (S7F Fig). Collectively, this work means that centrosome standing in ovarian most cancers can affect affected person final result, specifically with excessive CNR potentiating the response to chemotherapy. The general low ranges of centrosome amplification noticed within the sufferers nonetheless ask if and the way the preferential concentrating on of those cells can drive an improved therapeutic response, prompting the necessity for additional in vivo experimentation.
Dialogue
Centrosome amplification as a therapeutic goal has been primarily explored from the prism of multipolar division and mitotic medicine similar to HSET or Aurora A inhibitors have the potential to induce multipolar divisions and so to effectively goal most cancers cells in vitro [22,25], even when the potential of mitotic inhibitors have thus far been unsuccessful in scientific trials [49]. Our outcomes determine apoptotic priming as a novel cell dying susceptibility conferred by centrosome amplification. Particularly, we present that centrosome amplification sensitizes cells to BH3-mimetic medicine.
The apoptotic priming appears to be particular to centrosome amplification, somewhat than a consequence of the related mitotic stress. Potential causes of this priming may very well be disruption of mitochondrial networks throughout mitosis, or in interphase. This can be in hyperlink with current observations of subcellular reorganization in response to centrosome amplification in RPE-1 cells [50]. Centrosomes are additionally concerned in a number of signaling pathways [32,51] and given the pleiotropic results of centrosome amplification which additionally induce ROS and irritation [15]; we take into account that one of the best methodology to determine the supply of the priming can be whole-genome screening approaches.
From a scientific perspective, our evaluation of a affected person cohort reveals that prime centrosome numbers restrict relapse in response to chemotherapy, indicating that centrosome amplification have to be thought-about past its malignant potential. Given the toxicity of cytotoxic therapies, the angle of higher affected person stratification and response prediction, contemplating centrosome amplification as a sensitizing issue gives promising views. Our remark that centrosome amplification enhances cell dying independently of multipolar mitosis broadens the therapeutic significance of this most cancers cell function, past therapies that focus on spindle meeting and mitosis.
Our identification of apoptotic priming in cells with centrosome amplification additionally has scientific relevance, relative to the usage of BH3-mimetics within the clinic. A number of scientific trials contain the usage of these medicine alone or together with standard chemotherapy, with essentially the most promising instance being BCL-2 inhibitor Venetoclax together with Azacitidine, which is accepted for the remedy of acute myeloid lymphoma. Nonetheless, BH3 mimetic use within the clinic is hampered by lack of excellent prognostic markers and on-target toxicity of BCL-XL inhibitors WEHI-539 and Navitoclax, resulting in thrombocytopenia [52]. Our findings are fascinating for each of those challenges, first by inserting centrosome amplification as a possible biomarker for sensitivity to BH3-mimetics. Moreover, we ask if the apoptotic priming we describe may very well be associated to BCL-XL inhibitor toxicity, through sensitization of platelet-producing megakaryocytes that current centrosome amplification [53].
There are a number of limitations to our research. We should emphasize that the degrees of centrosome amplification within the cohort is low [5] and that centrosome loss may additionally contribute to modulating chemotherapy responses. It nonetheless stays fascinating to contemplate that concentrating on low ranges of centrosome amplification might have an observable scientific impact, and to clarify these outcomes we suggest that elimination of cells with centrosome amplification is likely to be advantageous given the malignant potential of those cells [7,13–15,19]. We’re additionally wanting to know if centrosome numbers affect responses to chemotherapy in extra epithelial ovarian most cancers cohorts and in numerous most cancers sorts. An essential step to facilitate broader research is the automatization of centrosome detection and counting in affected person tissues. Moreover our identification of apoptotic priming in response to centrosome amplification and the related scientific views justify the necessity for a greater understanding of the priming mechanism induced by centrosome amplification. This may additionally assist determine the contexts wherein this priming emerges as now we have not noticed it in all of the most cancers cell strains studied.
Materials and strategies
Examine design
This work is a research of the affect of centrosome amplification on the response to chemotherapy in epithelial ovarian most cancers. The targets of the cell biology work had been to determine if and the way centrosome amplification potentiates cell dying in response to chemotherapy, utilizing a mixture of single-cell reside imaging, and classical cell biology experiments similar to cytometry and western blot. All of the introduced knowledge has been replicated in 2 to five organic replicates. The target of the scientific work was to determine if centrosome numbers affect scientific parameters in a affected person cohort. All samples had been taken earlier than chemotherapy administration and obtained from the Organic Useful resource Heart (BRC) of Institut Curie (certification quantity: 2009/33837.4; AFNOR NF S 96 900). Sufferers offered oral consent for his or her samples for use for analysis functions. In compliance with the French regulation, sufferers had been knowledgeable of the research carried out on tissue specimens and didn’t specific opposition. The Nationwide Fee for Information Processing and Liberties (N° approval: 1487390) accepted all evaluation, in addition to The Institutional Overview Board and Ethics committee of the Institut Curie. Centrosomes had been beforehand stained and detected in an HGSOC affected person cohort of 100 sufferers [5]. Right here, we decided an index permitting the quantitative evaluation of centrosome numbers in affected person tissues, which we then correlated with affected person scientific parameters. Information assortment for every experiment is detailed within the respective determine legend.
Cell strains and cell tradition
All cell strains had been cultured at 37°C with 5% CO2 in DMEM/F12 media (Thermo Fisher Scientific #31331028) supplemented with 10% Tetracyclin-free Fetal Bovine Serum (Dutscher #500101L), 100 μg/ml streptomycin, and 100 U/ml penicillin (Thermo Fisher Scientific #15140122). OVCAR8 and COV504 had been obtained from the laboratory of F. Mechta-Grigoriou, and SKOV3 had been bought from ATCC (#HTB-77). Cell cultures underwent authentication by brief tandem repeat evaluation (powerplex16 HS equipment, Promega #DC2101) and had been routinely checked for mycoplasma (PlasmoTest Mycoplasma detection equipment, InvivoGen, #rep-pt1).
Cell line technology
Inducible PLK4 overexpression, H2B-RFP expression, FUCCI expression, GFP-tubulin expression, shRNA expression, and bulk CRISPR-Cas9 knock-out of STING had been stably established by lentiviral an infection. Viruses had been produced in HEK cells utilizing Lipofectamine 2000 (Thermo Fisher Scientific #11668019) to co-transfect lentiviral constructs with pMD2.G and psPAX2 plasmids. Viral particles had been collected within the supernatant 48 h after transfection, filtered, and used to contaminate the cell strains throughout 24 h. Cells had been then FACS sorted (inducible PLK4 overexpression, H2B-RFP, Tubulin-GFP) or chosen utilizing Puromycin at 5 μg/ml (shRNA strains and CRISPR-Cas9 knock-out of STING) or utilizing Blasticidin at 5 μg/ml (when the plasmid Lenti Tet-ON Myc-hPLK4 Blasticidin is used to induce PLK4 overexpression). Effectivity of knock-down and knock-out was assessed by western blot. The checklist of plasmids used is offered in S2 Table.
Drug therapies
All chemical compounds used are listed in S3 Table. To induce centrosome amplification, cells had been uncovered to doxycycline (1 μg/ml) or DMSO (diluent management, 1/10,000) for 72 h. If cells had been subsequently handled with one other drug, cells had been indifferent and replated with out addition of doxycycline to the PLK4OE inhabitants, and left to connect for 8 h. Drug therapies had been then carried out for 72 h on the indicated concentrations. For the experiments evaluating centrosome amplification to different mitotic stresses (CENP-E and MPS1 inhibition), cells had been uncovered to doxycycline (1 μg/ml for centrosome amplification), AZ3146 (1 μm, for MPS1 inhibition), GSK923295 (30 to 35 nM, for CENP-E inhibition), or DMSO (diluent management) for 72 h. Subsequent therapies (WEHI-539) or evaluation (live-imaging of mitotic phenotypes) had been then carried out in presence of the identical preliminary concentrations of drug for twenty-four h.
Dwell-imaging and evaluation
For live-imaging of chemotherapy responses, cells had been plated on Ibidi μ-Slide 8 Properly slides (Clinisciences, #80806-G500). Chemotherapy handled and untreated cells from each PLK4Ctl and PLK4OE populations had been imaged throughout the identical experiment. Imaging was carried out with a 20× dry goal (CFI Plan Apo LBDA 20× 0,75N.A) through an EMCCD digicam (Evolve, Photometrics) on an inverted microscope (Inverted Ti-E Nikon) outfitted with a spinning disk (CSU-X1 Yokogawa), a stage-top temperature and CO2 incubator (Tokai Hit) and built-in in Metamorph software program. For every effectively, 4 to six positions had been acquired each 10 min throughout 72 h, with a single slice within the brightfield channel and 10 3 μm slices per z-stack within the H2B-RFP channel or within the mKO2-Cdt1(30–120) and mAzami-Inexperienced-Gem1(1–110) channels for the FUCCI cells. For increased decision imaging, cells had been plated in 4 compartment Cellview dishes (Greiner Bio-One 627870) and three positions had been acquired in every effectively each 2,5 min throughout 24 h, utilizing a 40× oil goal (CFI Plan Fluor 40× 1,3N.A), buying a single slice within the brightfield channel and 20 1 μm slices per z-stack within the H2B-RFP channel. Time-lapse films had been then analyzed manually utilizing a customized Fiji macro to document an inventory of occasions, and a customized Python script to generate excel knowledge information and single-cell profiles.
For live-imaging of mitotic phenotypes induced by centrosome amplification, MPS1 inhibition and CENP-E inhibition, the identical strategy was used, buying every place each 5 min throughout 24 h. Imaging was carried out utilizing the tools described above.
Immunofluorescence
Cells had been plated on 18-mm glass coverslips in 12-well plates. Cells had been mounted for five min in ice-cold methanol (for S1A and S5E Figs), for 10 min in 4% PFA in PBS at room temperature (for Figs 3D, 4D and 4G), for 10 min in 4% PFA in PBS at 4°C (for S3I Fig), or for 10 min in 4% PFA + 0,025% Glutaraldehyde in 80 mM Pipes + 5 mM EGTA + 1 mM MgCl2 + 0,1% Triton X-100, earlier than 10 min quenching in 0,1% NaBH4 (for S1I, S2A and S2D Figs). Cells had been washed 3 instances in PBST (PBS + 0,1% Triton X-100) and incubated in PBST + BSA 0,5% for 30 min at room temperature. Cells had been then incubated for 1 h in main antibodies diluted in PBST + BSA 0,5%, washed 3 instances in PBST, incubated for 30 min in secondary antibodies diluted in PBST + BSA 0,5%, and washed 3 instances in PBST. Cells had been then stained for DNA utilizing 3 μg/ml DAPI diluted in PBST + BSA 0,5%, washed 3 instances in PBS, and mounted with mounting medium (1.25% n-propyl gallate, 75% glycerol, in H2O). Antibodies used are listed in S4 Table.
Immunofluorescence imaging and quantifications
Immunofluorescence pictures had been acquired with a sCMOS digicam (Flash 4.0 V2, Hamamatsu) on a widefield microscope (DM6B, Leica programs), with a 63× goal (63× HCX PL APO 1.40 to 0.60 Oil from Leica) or a 100× goal (100× HCX PL APO 1.40 to 0.70 Oil from Leica, for S1I, S2A and S2D Figs), utilizing Metamorph software program. Z-stacks had been acquired at 0,3 μm intervals.
Centrosome numbers, cytochrome C launch, and mitotic spindles had been scored manually. DNA injury marker depth or foci quantity had been decided on z-projections of pictures, utilizing a customized Python script to run the h_maxima perform from the skimage.morphology.extrema module.
Western blotting
Cells had been lysed in RIPA (150 mM sodium chloride, 1% NP-40, 0.5% sodium deoxycholate, 0.1% sodium dodecyl sulfate, 50 mM Tris, pH 8.0) complemented with protease (Sigma-Aldrich #11697498001) and phosphatase (Sigma-Aldrich #4906845001) inhibitors. Samples had been dosed utilizing a BiCinchoninic acid Assay (Pierce BCA protein assay, Thermo Fisher Scientific #23227). Samples had been diluted in RIPA with 4X NuPage LDS sampling buffer (Thermo Fisher Scientific #NP0007) and heated at 80°C for 10 min. Roughly 20 μg of protein was loaded in Bolt 4–12% Bis-Tris precast gels (Thermo Fisher Scientific #NW04125BOX) and subjected to electrophoresis in Bolt MOPS SDS operating buffer (Thermo Fisher Scientific #B0001). The gels had been transferred to nitrocellulose membranes (Dutscher #10600001) utilizing switch buffer (25 mM Tris, 192 mM Glycine, 20% Methanol) for 90 min at 4°C. Membranes had been stained in main or horseradish peroxidase coupled secondary antibodies diluted in PBS or TBS + 0,5% Tween 20 + 0,5% BSA or non-fat milk based on suppliers directions. Membranes had been first stained utilizing Ponceau, earlier than saturating for 1 h at room temperature in 5% non-fat dry milk or 5% BSA in PBS or TBS + 0,5% Tween20. Membranes had been then incubated in a single day in main antibodies, washed 5 instances in PBS or TBS + 0,5% Tween 20, then incubated for 1 h at room temperature in secondary antibodies. Membranes had been then washed once more 5 instances in PBS or TBS +0,5% Tween 20. Horseradish peroxidase response was developed utilizing SuperSignal Plus Chemiluminescent substrates (Thermo Fisher Scientific #34580 and #34094) and imaged (BioRad ChemiDoc MP). The Picture Lab software program (BioRad model 6.0.1) was used to measure background-adjusted quantity depth, which was normalized utilizing GAPDH sign. Antibodies used are listed in S4 Table.
Transfection
HT-DNA was transfected as a optimistic management for cGAS/Sting activation. Transfection of 1 μg/ml HTDNA was carried out utilizing Lipofectamine 2000 (Thermo Fisher Scientific #11668019) for twenty-four h.
Cytometry
Cells had been indifferent, rinsed in PBS, rinsed in AnnexinV Binding Buffer (BioLegend #422201), and round 100,000 cells had been resuspended in 50 μl Annexin V Binding Buffer. Cells had been stained utilizing Annexin V APC and Propidium Iodide (Biolegend #640932) at 0,6 μg/ml and 10 mg/ml, respectively. Cells had been incubated for 15 min, after which diluted in 200 μl Annexin V Binding Buffer. Cells had been analyzed utilizing a Bio-Rad ZE5 analyzer, and knowledge was analyzed utilizing FlowJo 10.6.0 software program.
MTT viability assays
For dose-response to medicine, cell viability was assessed utilizing MTT viability assays. Cells had been plated in triplicates at 15,000 cells/effectively in 96-well plates and left for two h to stick previous to drug addition. Cells had been left to develop for 72 h, and MTT diluted in PBS was added at 5 μg/ml. After 4 h incubation, medium was eliminated and changed by 150 μl DMSO, and 570 nm absorbance was carried out on a BMG Labtech ClarioStar plate reader. Triplicates had been averaged and normalized by untreated controls.
Trypan blue proliferation and viability assays
For proliferation and viability assays, cells had been plates at 100,000 cells/effectively in 6-well plates. Cells had been then indifferent, resuspended in 500 μl medium, and reside/lifeless cells had been counted utilizing a Beckman Coulter Vi-Cell cell counter.
RNA sequencing
Following centrosome amplification with doxycycline for PLKOE versus DMSO for PLK4Ctl, whole RNA was extracted with RNeasy Mini equipment (Qiagen #74104) following producer’s directions. RNA integrity and high quality had been checked with Agilent RNA 6000 Nano Package (Agilent, #5067–1511) and corresponding gadgets. Samples had been processed at Institut Curie NGS platform from cDNA synthesis, amplification, high quality evaluation, and sequencing. Novaseq 6000 system (Illumina) was used for sequencing (learn size of 100 bp, paired finish). All of the bioinformatic evaluation had been completed by Genosplice (http://www.genosplice.com) together with high quality management of sequences generated, learn mapping, and gene differential evaluation (R software program, Deseq2). Organic interpretation of the recognized genes was completed utilizing GSEA software for pathway enrichment evaluation between distinct circumstances.
Single-cell complete genome sequencing
Cells had been handled with DMSO (1/10,000), Doxycycline (1 μg/ml), or AZ3146 (1 μm) for 72 h. Cells had been then frozen in freezing medium (10% DMSO, 40% FBS in DMEM-F12).
Nuclei preparation and sorting
Cells had been thawed, and single-cell sequencing was carried out on cell nuclei remoted from cell lysis, leaving the nucleus intact. Thawed cells had been ready by resuspending in PBS + 1% BSA, washing, and pelleting. To generate nuclei, cells had been resuspended and incubated (15 min on ice in darkish atmosphere) in cell lysis buffer (100 mM Tris-HCl (pH 7.4), 154 mM NaCl, 1 mM CaCl2, 500 μm MgCl2, 0.2% BSA, 0.1% NP-40, 10 μg/ml Hoechst 33358, 2 μg/ml propidium iodide in ultra-pure water). Ensuing cell nuclei had been gated for G1 section (as decided by Hoechst and propidium iodide staining) and sorted into wells of 96 wells plates on a MoFlo Astrios cell sorter (Beckman Coulter), depositing 1 cell per effectively, and 96 wells plates containing nuclei and freezing buffer had been saved at −80°C till additional processing. Automated library preparation was then carried out as beforehand described [54].
AneuFinder evaluation
Sequencing was carried out utilizing a NextSeq 2000 machine (Illumina; as much as 120 cycles; single finish or as much as 113 and seven cycles; paired finish). The generated knowledge had been subsequently demultiplexed utilizing sample-specific barcodes and turned into fastq information utilizing bcl2fastq (Illumina; model 1.8.4). Reads had been afterwards aligned to the human reference genome (GRCh38/hg38) utilizing Bowtie2 (model 2.2.4; [55]). Duplicate reads had been marked with BamUtil (model 1.0.3; [56]).
The aligned learn knowledge (bam information) had been analyzed with a replica quantity calling algorithm referred to as AneuFinder (model 1.14.0; [57]) utilizing an euploid reference [58]. Following GC correction (R package deal: BSgenome.Hsapiens.UCSC.hg38_1.4.1; The Bioconductor Dev Group 2015) and blacklisting of artefact-prone areas (excessive low or excessive protection in management samples), libraries had been analyzed utilizing the dnacopy and edivisive copy quantity calling algorithms with variable width bins (common binsize = 1 Mb; step dimension = 500 kb) and breakpoint refinement (refine.breakpoints = TRUE). Outcomes had been afterwards curated by requiring a minimal concordance of 90% between the outcomes of the two algorithms. Libraries with on common lower than 10 reads per bin and per chromosome copy (roughly 55,000 reads for a diploid genome) had been discarded.
Aneuploidy rating
The aneuploidy rating of every bin was calculated as absolutely the distinction between the noticed copy quantity and the anticipated copy quantity when euploid. The rating for every library was calculated because the weighted common of all of the bins (dimension of the bin as weight) and the pattern scores had been calculated as the common of the scores of all libraries.
Heterogeneity rating
The heterogeneity rating of every bin was calculated because the proportion of pairwise comparisons (cell 1 versus cell 2, cell 1 versus cell 3, and many others.) that confirmed a distinction in copy quantity (e.g., cell 1: 2-somy and cell 2: 3-somy). The heterogeneity rating of every pattern was calculated because the weighted common of all of the bin scores (dimension of the bin as weight).
Centrosome numbers in tumors
For every pattern, 10 randomly chosen fields had been thought-about. Utilizing ImageJ software program, we visually counted the variety of nuclei and the variety of centrosomes in a blind method with out bearing in mind tumor id. The centrosome to nuclei ratio (CNR) was obtained by dividing the overall variety of centrosomes by the overall variety of nuclei in every subject.
Proliferation and mitotic index
For Ki67 proliferation evaluation, we carried out immunochemistry assays utilizing mouse anti-human ki67 antibody (M7240, DAKO, 1/200 at pH 9) in a sequence of paraffin-embedded tissue blocks of HGSOC. Sections of three μm had been lower utilizing a microtome from the paraffin-embedded tissue blocks of regular tissue and invasive lesions. Tissue sections had been deparaffinized and rehydrated by way of a sequence of xylene and ethanol washes. Briefly, the important thing steps included: (i) antigen retrieval with ER2 pH9, (Leica: AR9640); (ii) blocking of endogenous peroxidase exercise with Bond polymer refine detection equipment (Leica: DS9800); (iii) incubation with main antibodies in opposition to the focused antigen; and (iv) immunodetection with Revelation and counter staining Bond polymer refine detection equipment (Leica: DS9800). Immunostaining was carried out utilizing a Leica Bond RX automated immunostaining machine. We carried out an immunohistochemical rating (frequency x depth) by way of evaluation of 10 high-power fields (HPFs, x 400). All quantifications had been carried out by 2 pathologists with blinding of affected person standing.
For mitotic index, paraffin-embedded tissue sections of tumors had been stained with hematoxylin and eosin. The mitotic rely was decided by the variety of mitotic figures present in 10 consecutive HPFs, in essentially the most mitotically energetic a part of the tumor (total part). Solely identifiable mitotic figures had been counted. Hyperchromatic, karyorrhectic, or apoptotic nuclei had been excluded.
Supporting data
S1 Fig.
(A) Consultant pictures of OVCAR8 cells stained with DAPI (grey) and antibodies in opposition to CEP192 (Cyan) and Pericentrin (Magenta). (B) Bar graphs displaying the averages and SEM of the share of cells with the indicated variety of centrosomes (CEP192 dots colocalizing with Pericentrin). Three unbiased experiments, statistical check: Fisher’s actual check evaluating the variety of cells with greater than 2 centrosomes. (C) Dose-response of PLK4Ctl and PLK4OE cells to Paclitaxel (left) and Carboplatin (proper), normalized to their respective management circumstances, obtained from MTT viability assays. Imply and SEM of two unbiased experiments every obtained from averaging 3 technical replicates. (D) Mixture matrixes for Carboplatin and Paclitaxel mixed remedy, representing proportion of viability inhibition in comparison with management cells. Chosen working concentrations are highlighted in pink. (E) Scatter dot plots of Interphase size (high) and First mitosis size (backside), with Median and interquartile vary. Information from 2 unbiased experiments are pooled for Mixed remedy and Carboplatin remedy, knowledge from the 4 corresponding management experiments are pooled for Untreated. For interphase size a minimal of 26 cells was analyzed, and for mitosis size a minimal of 133 cells was analyzed. Statistical assessments: Kruskal–Wallis with Dunn’s a number of comparisons assessments. (F) Single-cell profiles of PLK4Ctl (left) and PLK4OE (proper) Untreated cells. Shade coding of mitosis and fates refers to classes outlined in Fig 1A. (G) Averages and SEM of the odds of mitotic phenotypes (legends in Fig 1A and 1F). Two unbiased experiments, statistical check: Fisher’s actual check on the variety of Slight Mis-segregation occasions. (H) Proportion of multipolar divisions noticed in presence or absence of 5 nM Paclitaxel. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of multipolar divisions. (I) Consultant pictures of OVCAR8 cells in prometaphase/metaphase stained with DAPI (blue), and antibodies in opposition to α-Tubulin (inexperienced), CEP192 (magenta), and Centrin3 (cyan) Numbered insets to point out centrosomes are introduced beneath the spindle picture. Scale bars are 10 μm for spindle pictures, and 1 μm for centrosome insets. Spindles of the “undefined” class are solely seen in cells handled with paclitaxel. (J) Averages and SEM of the odds of mitotic phenotypes recognized in panel I. Two unbiased experiments, statistical check: Fisher’s actual check on the overall variety of bipolar and pseudo-bipolar spindles. Information for S1 Fig could be present in S5 Data.
https://doi.org/10.1371/journal.pbio.3002759.s001
(TIFF)
S2 Fig.
(A) Consultant pictures of OVCAR8 cells in anaphase stained with DAPI (grey), and antibodies in opposition to α-Tubulin (inexperienced), CEP192 (magenta) and Centrin3 (cyan). Numbered insets to point out centrosomes and spindle poles are introduced beneath the spindle picture. Scale bars are 10 μm for spindle pictures and 1 μm for centrosome insets. (B) Averages and SEM of the odds of spindle sorts proven in panel C. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of bipolar spindles. (C) Averages and SEM of the odds of Carboplatin handled PLK4Ctl cells presenting extra MTOCs with the indicated traits. (D) Consultant picture of a Carboplatin handled PLK4OE OVCAR8 cell in anaphase stained with DAPI (grey), and antibodies in opposition to α-Tubulin (inexperienced), CEP192 (magenta) and CREST (cyan). Numbered insets to point out chromosomes with (1) or with out (2) centromeres. Scale bars are 10 μm for spindle picture and 1 μm for chromosome insets. (E) Averages and SEM of the odds of mitotic figures relying on the presence of acentric chromosomes. Two unbiased experiments, statistical check: Fisher’s actual check on the overall variety of bipolar and pseudo-bipolar spindles. (F) Averages and SEM of the odds of mitotic behaviours. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of Excessive mis-segregation occasions. (G) Averages and SEM of the odds of the indicated mis-segregation occasions inside bipolar divisions. Two unbiased experiments, statistical check: ANOVA with Sidak’s a number of comparability check. (H) Time lapse stills of iOVCAR8 cells expressing GFP-Tubulin (inexperienced) and H2B-RFP (pink) to visualise the mitotic spindle and chromosomes handled with DOX and carboplatin. Time is proven in minutes. White arrows level to centrosomes that don’t present a clustering habits, whereas yellow arrows level to clustered centrosomes. Within the high row, a cell enters mitosis with at the very least 5 centrosomes, it goes by way of a multipolarity standing earlier than reaching bipolarity and dividing in a bipolar method. The second row reveals a cell with at the very least 4 centrosomes that cluster to type a bipolar spindle. The white arrow at time level 70.00 min reveals a centrosome that emerges from the bipolar spindle however doesn’t end in multipolarity. The third row reveals a cell with extreme chromosome mis-alignment and mis-segregation defects, the place the spindle goes by way of cycles of collapsing and bipolarization. This cell takes over 10 h to divide. The fourth row reveals a cell with at the very least 6 centrosomes at mitotic entry. These are clustered in 3 foremost poles and the cell divides in a multipolar method. The fifth cell reveals a cell dying throughout mitosis. Word that mitotic time just isn’t prolonged as in different examples. (I) Averages and SEM of the odds of mitotic behaviors. Shade scheme is outlined in S2F Fig. Two unbiased experiments, statistical check: Fisher’s actual check on the variety of Excessive mis-segregation occasions. Information for S2 Fig could be present in S6 Data.
https://doi.org/10.1371/journal.pbio.3002759.s002
(TIFF)
S3 Fig.
(A, B) Single-cell profiles of FUCCI PLK4Ctl (A) and PLK4OE (B) untreated cells. See panel C for color-coded legends of cell cycle section and cell destiny. (C) Bar graphs displaying the averages and SEM of the odds of cell dying occasions occurring within the indicated cell-cycle phases. Two unbiased experiments, statistical check: Fisher’s actual check on variety of dying occasions occurring in S/G2. (D, E) Scatter dot plot graph of S/G2 (D) and G1 (E) section lengths within the second technology, with median and interquartile vary. Two unbiased experiments with a minimal of 18 instances analyzed per class. Statistical assessments: Kruskal–Wallis with Dunn’s a number of comparisons assessments. (F) Scatter dot plot graph of G1 section size within the second technology, with median and interquartile vary, relying on the destiny of the cells within the second technology. Two unbiased experiments with a minimal of 70 instances analyzed per class. Statistical assessments: Kruskal–Wallis with Dunn’s a number of comparisons assessments. (G) Consultant pictures of western blot evaluation of phosphorylated Chk1 and p53. (H) Graph displaying the common and SEM of phosphorylated protein relative to whole protein ranges, normalized to the degrees detected in PLK4OE cells handled with Carboplatin for twenty-four h, from 2 unbiased experiments. (I) Consultant pictures of cells stained with DAPI an antibodies in opposition to FANCD2 (cyan), 53BP1 (grey), and CEP192 (magenta). Grayscale pictures of RAD51 and γH2AX are proven. (J) Dot-plot representing built-in nuclear γH2AX fluorescence depth per cell or numbers of Rad51, FANCD2, or 53BP1 foci per cells. Common and SEM of the averages obtained from 3 unbiased experiments, every quantifying a minimal of 94 cells per situation. Values are normalized to the common of untreated PLK4Ctl cells at 24 h. Statistical check: ANOVA with Sidak’s a number of comparability assessments. Information for S3 Fig could be present in S7 Data.
https://doi.org/10.1371/journal.pbio.3002759.s003
(TIFF)
S4 Fig.
(A) Consultant pictures of western blot evaluation of p21 and PUMA. (B) Graph displaying the common and SEM of protein ranges from 4 (p53 and p21) or 3 (PUMA) unbiased experiments, normalized to ranges measured in untreated PLK4Ctl cells at 48 h. (C and G) Consultant pictures of western blot evaluation of indicated shRNA cell strains. (D and H) Bar graphs displaying the common and SEM of the share of cells in specified Annexin V-APC/PI gates analyzed by move cytometry. (D) Six replicates obtained from 4 unbiased experiments, with a minimal of 10,000 cells analyzed per situation and replicate. (H) 4 replicates obtained from 2 unbiased experiments, with a minimal of 10,000 cells analyzed per situation and replicate. Statistical check: comparability of the share of Annexin V optimistic cells, utilizing ANOVA with Sidak’s a number of comparability check. Consultant cytometry profiles could be discovered within the S1 Appendix. (E) Consultant pictures of western blot evaluation of Caspase2 and MDM2 cleavage. (F) Common and SEM of protein ranges from 3 (p19 Caspase2) or 2 (p55 MDM2) unbiased experiments, normalized to ranges measured in untreated PLK4OE cells at 24 h. Information for S4 Fig could be present in S8 Data.
https://doi.org/10.1371/journal.pbio.3002759.s004
(TIFF)
S5 Fig.
(A, B) Dose-response of PLK4Ctl and PLK4OE cells to A1210477 (A) and Venetoclax (B), normalized to their respective untreated circumstances, obtained from MTT viability assays. Imply and SEM of two unbiased experiments every obtained from averaging 3 technical replicates. (C) Bar graphs displaying the common and SEM of the share of cells with the indicated variety of centrosomes (CEP192 dots colocalizing with Pericentrin). Two unbiased experiments, statistical check: Fisher’s actual check evaluating the variety of cells with greater than 2 centrosomes. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (D) Bar graphs displaying the common and SEM of the share of cells with the indicated variety of centrosomes (CEP192 dots colocalizing with Pericentrin). Two unbiased experiments, statistical check: Fisher’s actual check evaluating the variety of cells with greater than 2 centrosomes. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (E) Consultant pictures of COV504 and SKOV3 cells stained with DAPI (grey) and antibodies in opposition to CEP192 (cyan) and Pericentrin (magenta). (F) Bar graphs displaying the common and SEM of the odds of cells with the indicated variety of centrosomes (CEP192 dots colocalizing with Pericentrin). Three unbiased experiments, statistical assessments: Fisher’s actual assessments evaluating the variety of cells with greater than 2 centrosomes. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (G) Dose-response of PLK4Ctl and PLK4OE OVCAR8 (high), COV504 (center), and SKOV3 (backside) to Navitoclax, normalized to their respective untreated circumstances, obtained from MTT viability assays. Imply and SEM of two to three unbiased experiments every obtained from averaging 3 technical replicates. (H) Desk summarizing the IC50s and drug concentrations utilized in combos, decided through MTT dose-response viability assays, for PLK4Ctl OVCAR8, COV504, and SKOV3. (I) Viability (% of Trypan Blue damaging cells) counted for PLK4Ctl and PLK4OE OVCAR8 (high), COV504 (center), and SKOV3 (backside), in response to indicated chemotherapies utilizing concentrations indicated in panel G for 72 h. Common and SEM of the variety of unbiased experiments indicated, every obtained from averaging 3 technical replicates. Statistical check: two-way ANOVA with Sidak’s a number of comparability check. (J) Proportion of multipolar divisions noticed in COV504 in response to five nM Paclitaxel publicity. (Ok) Viability (% of Trypan Blue damaging cells) counted for PLK4Ctl and PLK4OE OVCAR8 (high), COV504 (center), and SKOV3 (backside), in response to Olaparib utilizing concentrations indicated in panel G for 72 h. Common and SEM of 4 unbiased experiments, every obtained from averaging 3 technical replicates. Statistical check: two-way ANOVA with Sidak’s a number of comparability check. Information for S5 Fig could be present in S9 Data.
https://doi.org/10.1371/journal.pbio.3002759.s005
(TIFF)
S6 Fig.
(A) Genome-wide copy-number plots for G1 OVCAR8 cells. Every row represents a cell. Indicated aneuploidy and heterogeneity scores are calculated as described in Materials and strategies. (B) Bar graphs displaying the common and SEM of the share of mitotic phenotypes as outlined in Fig 1A. Two unbiased experiments, statistical check: comparability of the share of cells with no mis-segregation utilizing ANOVA with Dunnett’s a number of comparability check. Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. (C) Scatter dot plot graph of mitosis size with median and interquartile vary. No less than 96 mitosis analyzed from 2 unbiased experiments, statistical check: Kruskall–Wallis with Dunn’s a number of comparability check. (D) Bar graphs displaying the common and SEM of the share of cells in specified Annexin V-APC/PI gates analyzed by move cytometry. Three replicates obtained from 2 unbiased experiments, with a minimal of 20,000 cells analyzed per situation and replicate. Statistical check: comparability of the share of Annexin V optimistic cells, utilizing ANOVA with Sidak’s a number of comparability check. Consultant cytometry profiles could be discovered within the S1_Appendix. (E) GSEA Hallmarks with |normalized enrichment rating >1,5 and p-value <0,05, from differential RNA expression evaluation of PLK4OE cells in comparison with PLK4Ctl. (F) Western blot evaluation of STING phosphorylation after 72 h of indicated drug therapies with quantification of pSTING relative to whole STING. Common and SEM of two unbiased experiments, normalized to untreated PLK4Ctl. HT-DNA transfection was carried out 24 h earlier than cell assortment. (G) Western blot evaluation of indicated bulk LentiCRISPR cell strains. (H) Bar graphs displaying the common and SEM of the share of cells in specified Annexin V-APC/PI gates analyzed by move cytometry. 4 replicates obtained from 2 unbiased experiments with a minimal of 15,000 cells analyzed per situation and replicate. Statistical check: comparability of the share of Annexin V optimistic cells utilizing ANOVA with Sidak’s a number of comparability check. Consultant cytometry profiles could be discovered within the S1_Appendix Numbers on the highest of every graph symbolize the variety of cells analyzed per situation. Information for S6 Fig could be present in S10 Data.
https://doi.org/10.1371/journal.pbio.3002759.s006
(TIFF)
S7 Fig.
(A) Common and SEM of the CNR established in 10 fields per tumor or wholesome tissue pattern. Pink dotted line signifies CNR = 1,45, the cut-off between Excessive-CNR and Low-CNR sufferers. (B, C) Distribution of the Mitotic Index (B) and of the share of Ki67 optimistic cells (C) as a perform of CNR. Statistical check: Spearman correlation. (D) Common and SEM of CNR per affected person labeled relying on FIGO stage. (E, F) Kaplan–Meier curves for general survival (B) and relapse-free time after the primary line of chemotherapy (C) based on CNR standing. Information for S7 Fig could be present in S11 Data.
https://doi.org/10.1371/journal.pbio.3002759.s007
(TIFF)