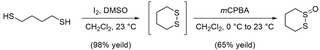Quotation: Hossain MA, Sarin R, Donnelly DP, Miller BC, Weiss A, McAlary L, et al. (2024) Evaluating protein cross-linking as a therapeutic technique to stabilize SOD1 variants in a mouse mannequin of familial ALS. PLoS Biol 22(1):
e3002462.
https://doi.org/10.1371/journal.pbio.3002462
Educational Editor: Josh Dubnau, Stony Brook College Medical Heart: Stony Brook College Hospital, UNITED STATES
Obtained: Could 17, 2022; Accepted: December 5, 2023; Revealed: January 30, 2024
Copyright: © 2024 Hossain et al. That is an open entry article distributed below the phrases of the Creative Commons Attribution License, which allows unrestricted use, distribution, and copy in any medium, offered the unique writer and supply are credited.
Knowledge Availability: All related information are inside the paper and its Supporting Information recordsdata. As well as, the mass spectrometry proteomics information have been deposited to the ProteomeXchange Consortium by way of the PRIDE accomplice repository with the dataset identifier (accession) PXD047030. Structural information was deposited to PDB and could be accessed with the next IDs: 8CCX and 8Q6M.
Funding: This work was supported by the Nationwide Institute of Neurological Issues and Stroke of Nationwide Institute of Well being (R01NS065263 to J.N.A.), the ALS Affiliation (18-IIA-420 to J.N.A., M.J.O., and R.M.), Johnston Rducational Ventures (#685162 to J.N.A.), and the Nationwide Science Basis (#MCB-1517290, CHE-1905214 to M.J.O.). The funders had no function in examine design, information assortment and evaluation, choice to publish, or preparation of the manuscript.
Competing pursuits: The authors have declared that no competing pursuits exist.
Abbreviations:
ALS,
amyotrophic lateral sclerosis; DSF,
differential scanning fluorimetry; fALS,
familial amyotrophic lateral sclerosis; FBS,
fetal bovine serum; FT-ICR-MS,
Fourier-transform ion cyclotron resonance mass spectrometry; H/D-X MS,
hydrogen deuterium alternate mass spectrometry; IV,
intravenous; LC-MS,
liquid chromatography-mass spectrometry; LC-MS/MS,
liquid chromatography-tandem mass spectrometry; MALDI,
matrix-assisted laser desorption ionization; MS,
mass spectrometry; NBT,
nitroblue tetrazolium; NCI,
Nationwide Most cancers Institute; PD,
pharmacodynamic; PTM,
posttranslational modification; RBC,
purple blood cell; sALS,
sporadic amyotrophic lateral sclerosis; SARS-CoV-2,
Extreme Acute Respiratory Syndrome Coronavirus 2; SAXS,
small-angle X-ray scattering; SC,
subcutaneous; SEC,
measurement exclusion chromatography; SOD1,
superoxide dismutase 1
Introduction
Amyotrophic lateral sclerosis (ALS) is a deadly neurodegenerative illness that usually results in demise inside 2 to five years of prognosis [1,2], and 5% to 10% of ALS instances are familial (fALS) whereas the remainder are sporadic [3,4]. Over 180 mutations within the gene encoding superoxide dismutase-1 (SOD1) contribute to roughly 20% of fALS instances [5,6]. These mutations are typically dominantly inherited, extremely penetrant, and related to various levels of illness severity and lack of enzymatic exercise [7,8]. For instance, sufferers harboring the SOD1 variant SOD1A4V don’t typically survive greater than 1 yr after prognosis, whereas these with SOD1H46R survive a mean of 18 years [9]. Many research have additionally implicated wild-type SOD1 in idiopathic, sporadic ALS (sALS). For instance, posttranslational modifications (PTMs) reminiscent of oxidation of Cys111 [10] are poisonous and autoantibodies [11] to SOD1 are extra prevalent in sALS instances. Nevertheless, the function of wild-type SOD1 in ALS stays a topic of debate [12]. The cyclic thiosulfinates used on this examine block oxidation of Cys111 and might be a helpful chemical device for addressing the function of wild-type SOD1 in sALS.
Wild-type SOD1 is a homodimer. A typical property of ALS-associated SOD1 mutations is that they enhance the propensity of the dissociation of the SOD1 dimer, which promotes misfolding and aggregation of the ensuing monomers into cytotoxic species [13]. Small molecules are due to this fact being developed to stabilize the quaternary construction of fALS SOD1 variants [14,15]. A similar technique was employed efficiently to create tafamidis, a drug for transthyretin amyloidosis and transthyretin cardiomyopathy, which stabilizes the native transthyretin tetramer [16]. Sadly, molecules supposed to stabilize SOD1 by way of non-covalent interactions on the dimer interface [17] as a substitute certain to the β-barrel area of the SOD1 aggregates [14] and sometimes exhibited plasma protein binding [15].
Right here, we pursue an alternate method to stabilize the SOD1 dimer. We postulate that 2 free cysteines located on adjoining monomers, Cys111a and Cys111b (separate by roughly 9 Å), might be cross-linked to forestall dimer dissociation. Cysteine residues could be focused with excessive selectivity because of the distinctive reactivity (nucleophilicity and polarizability) of the thiolate purposeful group. That is evidenced by quite a few medication that type covalent bonds to cysteine residues, together with the irreversible aldehyde dehydrogenase (ALDH1A1) inhibitor disulfiram (Antabuse) [18]; the blockbuster proton pump inhibitors, e.g., omeprazole (Prilosec) and its single enantiomer esomeprazole (Nexium) [19]; inhibition of Extreme Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) important protease by ebselen [20] and Paxlovid, and second-generation kinase inhibitors, e.g., afatinib (Gilotrif) [21]. Formation of covalent bonds to cysteines has additionally been proven to offer entry to “undruggable” targets, as evidenced by Amgen’s drug for KRAS G12C mutations, sotorasib. In proof-of-concept experiments, we demonstrated that cross-linking the Cys111a,b pair on the illness variants SOD1G93A and SOD1G85R may stabilize the SOD1 dimer and rescue superoxide dismutase exercise [22]. The bifunctional maleimide cross-linkers utilized in these research, nonetheless, are poisonous [23] as a result of they aim any uncovered free cysteine, lots of which serve important catalytic and redox roles for different proteins. Slightly than cross-linking, ebselen and cisplatin goal SOD1 Cys111 [24,25] and stabilize the SOD1 dimer by non-covalent interactions. Ebselen has been proven to delay onset and enhance survival within the G93A mouse mannequin [26]. These promising preclinical leads might share maleimide’s propensity for off-target binding, as evidenced by ebselen’s off-target binding to Cys6 [27] and the hepatotoxicity of cisplatin. A therapeutically viable cross-linking mechanism for SOD1-dependent fALS ought to bind the Cys111 pair whereas avoiding different cysteines.
We not too long ago launched cyclic thiosulfinates, which selectively cross-link pairs of free cysteines, whereas avoiding “dead-end” modifications of lone cysteines [28]. The cyclic thiosulfinate 1,2-dithiane-1-oxide (Sulfur cross-linking 6-membered ring, hereafter S-XL6) and β-lipoic acid (the S-oxo spinoff of α-lipoic acid) [28], goal the Cys111a and Cys111b residues on adjoining SOD1 monomers, forming a disulfide bond with every cysteine thiolate. Cyclic thiosulfinates are one oxygen (S-oxo) derivatives of cyclic disulfides. Nicely-known and well-tolerated cyclic disulfides embrace the pure product asparagusic acid and the multi-enzyme cofactor and dietary complement α-lipoic acid. On this examine, we check the next hypotheses: (1) cross-linking improves the biochemical properties of fALS SOD1 variants; and (2) S-XL6 engages SOD1 in vivo.
Outcomes
A computational search was carried out in PubChem to establish all compounds containing a cyclic disulfide motif. These compounds, which consisted of 186 cyclic disulfide derivatives, have been acquired from the Nationwide Most cancers Institute (NCI) and screened for the flexibility to cross-link SOD1. Of those, 69 compounds cross-linked SOD1 with a big selection of response charges. Solely 2 compounds may cross-link 100% of SOD1 in the course of the assay, together with one five-membered cyclic disulfide (4-Amino-1,2-Dithiolane-4-Carboxylic Acid) and one six-membered cyclic thiosulfinate (1,2-dithiane 1-oxide). Extra cyclic thiosulfinates have been synthesized to create sequence and fill gaps in chemical house, together with 5- (1,2-dithiolane 1-oxide), 6- (1,2-dithiane 1-oxide), 7- (1,2-dithiepane 1-oxide), and eight-membered rings (1,2-dithiocane 1-oxide) and their properties have been decided (e.g., EC50 for cross-linking SOD1, cytotoxicity, solubility, clearance, cross-linking price, LD50 in mice). The outcomes of those research—particularly, the distinction between EC50 and LD50—recommended that solely the 5- and six-membered rings had potential as fALS therapeutics. The six-membered cyclic thiosulfinate, 1,2-dithiane 1-oxide (S-XL6, Fig 1), by advantage of its scalable synthesis and decrease cytotoxicity and clearance, was chosen for preliminary characterization.
Fig 1. Cyclic thiosulfinate S-XL6 cross-links SOD1 variants by way of Cys111 residues on adjoining monomers.
(High) Crystal construction of wild-type SOD1 (PDB ID: 1SPD [5], cartoon illustration generated with Maestro 11.8) highlighting opposing Cys111 residues on each monomer A (blue) and monomer B (inexperienced) with a illustration of the S-XL6 cross-linked SOD1 dimer. Cu and Zn molecules are represented by orange and grey spheres, respectively. (A) Trypsin digest and MALDI-ToF-MS evaluation of SOD1A4V gave a mixed sequence protection of 94%, together with peaks similar to the Cys57-Cys146 disulfide linked peptides (blue) and the Cys111- containing peptide (purple) linked by S-XL6. N-terminal acetylation is proven in blue textual content, cysteines are proven in purple textual content, and the SOD1A4V mutation is underlined. (B) Inspecting the upper mass vary revealed multiply charged types of SOD1A4V together with the S-XL6 linked dimer, in addition to 2 totally different peaks similar to Cys111-linked peptides. (C) MALDI-FTICR-MS evaluation of pepsin digested SOD1H46R. The highest panel reveals SOD1H46R management pattern and center panel reveals peaks similar to Cys111 linked peptides by way of S-XL6. The cross-linked peptides are proven in black textual content with cysteines highlighted in purple. The underside panel reveals affirmation of cross-linked peptides utilizing deuterated S-XL6 (mass shift of 8 Da). MALDI, matrix-assisted laser desorption ionization; MS, mass spectrometry; SOD1, superoxide dismutase 1.
To check the efficacy of cross-linker-mediated SOD1 stabilization, we selected fALS variants starting from “wild-type-like” (metalated and enzymatically energetic SOD1A4V and SOD1G93A), to “metallic poor” (impaired metallic binding and inactive SOD1H46R and SOD1G85R). These variants are aggregation-prone, span a variety of scientific prognoses (1 yr to 18 years survival) and have a variety of properties that’s believed to signify most fALS variants. Utilizing these, we examined the speculation that cross-linking improves the biochemical/biophysical properties of fALS SOD1 variants, e.g., that S-XL6 will increase thermal stability and restores construction and enzymatic exercise. We then examined the speculation that cross-linking can stabilize a fALS SOD1 variant in vivo and enhance survival.
S-XL6 cross-links and stabilizes fALS SOD1 variants with numerous physicochemical properties
We employed a mass spectrometry (MS) assay for intact protein cross-linking [22,29]. On this assay, any non-covalent (i.e., native) dimer is dissociated by a mixture of acidic, natural media and MS supply situations. Utilizing this assay, we noticed S-XL6-dependent formation of cross-linked dimers for wild-type SOD1, SOD1A4V, SOD1G93A, SOD1H46R, and SOD1G85R (Fig 2, left). This response was environment friendly (full conversion of monomer to dimer) and stoichiometric (occurring at ratios as little as 1:1 cross-linker to dimer). Useless-end modifications of SOD1 weren’t noticed on this assay. We confirmed the mechanism of motion (MoA, e.g., lack of oxygen from the cross-linker concomitant with dimer formation, Fig 1) and decided the location of cross-linking utilizing endoproteinase digestion and MALDI (matrix-assisted laser desorption ionization) peptide mass fingerprinting evaluation of cross-linked samples (Fig 1).
Fig 2. S-XL6-mediated cross-linking and stabilization of wild-type and fALS SOD1 variants.
High: chemical mechanism of motion of S-XL6. The cross-linking response proceeds by an preliminary thiolate-disulfide interchange between a cysteine thiolate and the cyclic thiosulfinate, producing a sulfenic acid intermediate upon opening of the ring construction. This sulfenic acid intermediate kinds a cross-link by fast condensation with a second cysteine thiolate. Mass spectra of untreated (left column, common mass [± 1 Da]) and S-XL6 cross-linked (center column, common mass [± 2 Da]) proteins are in keeping with cross-linking. DSF outcomes (proper column) point out that cross-linking elevated the thermal stability of SOD1 and its variants. We quantify ΔTu because the distinction in unfolding temperature measured between the most important inflections of the untreated and cross-linked samples. The unfolding temperature of the untreated and S-XL6 cross-linked SOD1 proteins are as follows: wild sort (WT) from 75.9°C to 90.2°C (ΔTu ~14°C); SOD1A4V from 62.9°C to 79.3°C (ΔTu ~16°C); SOD1G93A from 70.6°C to 85.0°C (ΔTu ~14°C); SOD1H46R from 46.3°C to 69.3°C (ΔTu ~23°C); SOD1G85R from 40.8°C to 65.1°C (ΔTu ~24°C). Be aware: two inflections could be noticed when there’s a combination of partially (decrease ΔTu) and absolutely metalated (greater ΔTu) SOD1 proteoforms, which applies to the next proteins: untreated wild-type SOD1 unfolds at 75.9°C and 87.6°C; untreated SOD1G93A unfolds at 70.6°C and 84.4°C. All samples have been analyzed in technical triplicate (customary errors <0.3°C). fALS, familial amyotrophic lateral sclerosis; SOD1, superoxide dismutase 1.
SOD1 mutations destabilize the native dimer to various levels. Lack of stability of fALS variants correlates with illness severity, particularly with fast development [30]. To grasp how cross-linking results SOD1 stability, we quantified modifications in unfolding temperature (ΔTu, hereafter thermal stability). Purified wild-type, SOD1A4V, SOD1G93A, SOD1H46R, and SOD1G85R proteins have been handled with S-XL6. Thermally induced unfolding of SOD1 variants was measured utilizing a beforehand revealed methodology, differential scanning fluorimetry (DSF) (Fig 2, proper) [31]. The will increase in thermal stability attained with S-XL6 (14 to 24°C) evaluate favorably with one of the best preclinical candidate for stabilizing SOD1 (e.g., 11°C for ebselen analogues, [26]) and with the one accepted drug with a kinetic stabilizing mechanism, tafamidis (6°C) [32].
Structural evaluation of S-XL6 cross-linked fALS SOD1 variants
To find out the affect of cross-linking on the upper order construction of fALS SOD1 variants, we utilized measurement exclusion chromatography (SEC), hydrogen deuterium alternate mass spectrometry (H/D-X MS), X-ray crystallography, and small-angle X-ray scattering (SAXS). Whereas the MS assay above was carried out below denaturing situations, SEC was carried out below native situations. SEC research confirmed earlier analysis exhibiting {that a} substantial proportion of fALS variants are monomeric close to physiological focus and would profit from kinetic stabilization (Figs 3 and S9). The SEC outcomes additionally confirmed the conversion of fALS SOD1 monomers into dimer following S-XL6 remedy and supply a complementary measurement of cross-linking effectivity. Whereas MS-based detection is closely biased in the direction of low molecular mass species (i.e., typically can’t detect massive aggregates) SEC can detect massive oligomers [14,33–37]. The SEC outcomes present that giant (>1,000 kDa) aggregates are current in all preparations of untreated fALS variants. Notably, these aggregates have been diminished in all of the fALS variants we examined following remedy with S-XL6. The power of S-XL6 to decrease aggregates is anticipated based mostly upon S-XL6-mediated stabilization of SOD1 variants, on condition that stabilization will increase the time required for the nucleation part of aggregation. We’re at the moment unable to measure the charges of aggregation as a result of current assays require reductants that may additionally take away S-XL6 [38,39].
Fig 3. SEC demonstrates that the monomeric inhabitants of fALS variants is decreased by S-XL6 remedy and that that is related to decreased aggregation.
High: SEC outcomes for the molecular weight calibration requirements, Thyroglobulin (660 kDa), IgG (150 kDa), BSA (66 kDa), and myoglobin (17 kDa). Backside: SEC outcomes for a similar variants as Figs 2 and 4, with and with out S-XL6 remedy. The height labeled “aggregates” elute close to the void quantity of the SEC column and are due to this fact too massive (>1,000 kDa) for MW dedication. fALS, familial amyotrophic lateral sclerosis; SEC, measurement exclusion chromatography.
H/D-X MS assesses construction and dynamics by measuring variations in deuterium uptake [40]. Following cross-linking, we noticed that the buildings of fALS SOD1 variants bear a more in-depth resemblance to, however don’t absolutely recapitulate, the construction of wild-type SOD1 construction (Fig 4). For instance, decreased uptake on the N- and C-termini of SOD1 (the dimer interface) was noticed in all cross-linked variants. In all however one variant, SOD1G93A, the uptake round residues 37–43 extra intently resembled wild sort. Notably, structural modifications on this area are chargeable for the “achieve of interplay” with the disordered electrostatic loop, which has been proposed to result in the aggregation of fALS variants [41]. Earlier research have categorized the enzymatically energetic SOD1A4V and SOD1G93A proteins [41,42] as wild-type-like variants and the inactive, less-folded SOD1G85R and SOD1H46R proteins [43,44] as metal-deficient variants. In line with this, bigger variations in uptake by way of H/D-X MS have been noticed for SOD1G85R and SOD1H46R. Refined (<5%) variations in deuteration ranges have been additionally noticed in different areas of the SOD1 variants. In abstract, cross-linking makes the construction and dynamics of fALS SOD1 variants extra like that of wild-type SOD1 and in a way that’s in keeping with lowering their aggregation propensity.
Fig 4. Cross-linking enhances the construction of fALS SOD1 variants.
Variations in deuterium uptake (ΔU, legend proven under half b) of untreated and cross-linked variants for the 4-h time level in comparison with the wild-type SOD1 (WT) protein are reported right here. (A) Black arrows point out areas the place distinguished shifts in ΔU have been noticed (e.g., residues 2–7 for SOD1A4V untreated 13.8% to 1.2% cross-linked, for full outcomes see S1 Fig). (B) ΔU for untreated and cross-linked SOD1A4V mapped onto the cartoon illustration of wild-type SOD1 construction (PDB ID: 1SPD) [5], generated with Maestro 11.8. ΔU forever factors (15 s, 50 s, 500 s, 1 h, 4 h) are reported in S1 Fig. fALS, familial amyotrophic lateral sclerosis; SOD1, superoxide dismutase 1.
Enzymatic exercise of cross-linked SOD1 variants is rescued to that of wild-type SOD1
Using each plate-based (biochemical) and gel-based assays, we characterised the affect of cross-linking on the enzymatic exercise of SOD1 variants. Cross-linking elevated the exercise of fALS SOD1A4V, SOD1G93A, and SOD1G85R variants to ranges corresponding to that of wild-type SOD1 (Fig 5A and 5B and S1 Table). No change in exercise was noticed for H46R, which was as anticipated as a result of His46 is required for Cu binding and enzymatic exercise. This outcome, mixed with metallic content material in S4 Table indicating that variant have much less Cu than WT-SOD1, and the requirement of Cu for enzymatic exercise, indicate that S-XL6 allows extra Cu incorporation into SOD1.
Fig 5. Enzymatic exercise of SOD1 variants after S-XL6 remedy.
Plate-based biochemical (A) and gel-based (B) assays have been used to evaluate the affect of cross-linking on the enzymatic exercise of SOD1 variants. Upon remedy with cross-linker the exercise of SOD1A4V, SOD1G93A, and SOD1G85R elevated to that of wild-type SOD1. Therapy with crosslinker didn’t have an effect on the enzymatically inactive SOD1H46R. “+” and “–” denote samples handled with S-XL6 and untreated samples, respectively. The info underlying this determine could be present in S1 Data. SOD1, superoxide dismutase 1.
A cross-linker with promising toxicological properties promotes SOD1 dimer formation in cellulo
We characterised the effectiveness of our crosslinker utilizing Hep G2 cells. SOD1 cross-linking was monitored utilizing SDS-PAGE and western blotting. S-XL6 cross-linked wild-type SOD1 in cells in PBS buffer with the half maximal efficient concentrations (EC50) at circa (ca.) 5 μm (Fig 6A). These outcomes are additionally in keeping with earlier analysis indicating that cyclic thiosulfinates are actively transported throughout the mobile membrane [28] the place they continue to be intact [45]. Plasma protein binding was immediately assessed utilizing human plasma and no binding of S-XL6 was noticed (Fig 6C). The minimal plasma protein binding is promising and is enabled by cyclic thiosulfinates’ means to keep away from lone cysteines, together with the extremely considerable and reactive lone cysteine (Cys34) [46] of serum albumin.
Fig 6. Cross-linking at sub-toxic concentrations inside cells.
(A) Western blot evaluation utilizing a SOD1-selective antibody signifies that cross-linking proceeds in HEP G2 cells with an EC50 of roughly 5 μm (in PBS buffer). (B) Low cytotoxicity of S-XL6 (LC50 roughly 446 μm) was noticed utilizing the MTT (3-(4,5-Dimethylthiazol 2-yl)-2,5-diphenyltetrazolium bromide) cytotoxicity assay (triplicate pattern evaluation). Outcomes are proven as proportion of viable cells in comparison with a car management. Be aware: cell viability <70% (dotted line) is mostly thought of as the edge for cytotoxicity, and viabilities >100% point out a trophic impact. Controls included EMEM with 0.1% DMSO (dimethyl sulfoxide, detrimental management) and 500 μm of chlorpromazine (optimistic management). (C) Plasma protein binding was assayed in human platelet wealthy (hPRP) and platelet poor plasma (hPPP) utilizing an LC-MS/MS assay at 3 concentrations (10, 50, and 250 μm) and no binding of S-XL6 was noticed. Likewise, S-XL6 was not certain to purified alpha 1-acid glycoprotein and human serum albumin. The info underlying this determine could be present in S1 Data. LC-MS/MS, liquid chromatography-tandem mass spectrometry; MS, mass spectrometry; SOD1, superoxide dismutase 1.
The cytotoxicity of S-XL6 was measured in Hep G2 utilizing a regular MTT assay (LC50 roughly 446 μm, Fig 6B). Provided that the LC50 is 90× the EC50, S-XL6 can promote dimer formation in cellulo with minimal toxicity. S-XL6 didn’t have an effect on the survival or aggregation of EGFP-labeled G93A in NSC-34 cells (S3 Fig). S-XL6 remedy of EGFP-labeled wild-type SOD1 elevated mobile aggregation and a high-molecular weight species noticed in western blots in NSC-34 cells (S3 Fig). We word that the identical high-molecular weight species weren’t noticed in HEPG2 (Fig 6) or HELA cells [28], which natively specific wild-type SOD1. Additional research will decide if S-XL6-dependent inclusion formation of wild-type SOD1 is neuron-related or EGFP-labeling-related. Outcomes obtained in NSC-34 cells, particularly the shortage of cross-linking within the Cys111Ser variant, point out that the Cys111 residue is focused by S-XL6 and is required for cross-linking. Outcomes obtained in NSC-34, particularly the shortage of binding to a constitutively monomeric variant, additionally point out that the dimeric subpopulation of variants is focused. The binding of S-XL6 to a wider array of fALS variants (EGFP-labeled) was additionally examined in NSC-34 cells and was in comparison with that of our greatest third-generation compounds. In comparison with our main compounds, S-XL6 persistently resulted within the highest cross-linking yield and was due to this fact chosen for in vivo research (Fig 7).
Fig 7. S-XL6 outperforms main cyclic thiosulfinates, binds a wide range of fALS variants, and requires the Cys111 residue in cellulo.
C-terminally EGFP-labeled SOD1 (wild-type SOD1, 8 fALS variants, a Cys111Ser variant, and a constitutively monomeric F50E/G51E variant) have been expressed in NSC-34 cells. S-XL6 was the simplest compound of the top-five six-membered cyclic thiosulfinates recognized by mass spectrometry screening of NCI compounds and our personal medicinal chemistry efforts (labeled “Comp 2–5”). Compounds 2 and 3 are epimers of the 4,5-hydroxyl derivates of S-XL6; the identification of compounds 4 and 5 are unpatented and can’t be disclosed right here, and 20 μm S-XL6 engaged all fALS variants aside from G85R and wild-type SOD1, however not Cys111Ser or F50E/G51E. EGFP-wild-type and EGFP-D90A SOD1 fashioned an off-product (unintended) high-molecular weight species in keeping with trimeric SOD1. S-XL6 remedy didn’t have an effect on the mobile viability or aggregation of G93A SOD1. S-XL6 remedy didn’t have an effect on the viability however did speed up the aggregation of wild-type SOD1 (S3 Fig). fALS, familial amyotrophic lateral sclerosis; SOD1, superoxide dismutase 1.
S-XL6 binds to Cys111 and cross-links the monomers growing dimer stability
Co-crystallographic construction of SOD1 soaked with S-XL6 was decided to 1.67 Å decision utilizing the X-ray facility on the Barkla X-ray laboratory of Biophysics, Liverpool. As a management, as-isolated SOD1 construction was additionally decided below an analogous low-dose situations to 1.77 Å decision. Decision of the as-isolated construction is rather less as a smaller crystal was used for this function. The high-resolution construction of the complicated offers clear proof for the binding of the compound at Cys111 whereas the as-isolated construction reveals various water molecules (Fig 8). Particularly, the construction of SOD1 with S-XL6 reveals that the linker is hooked up by S-S bridges to SD atoms of Cys111 and connects 2 monomers (Fig 8). The electron density for carbon atoms of the compound is far weaker than for sulphur atoms, in all probability because of the flexibility of carbon chain and/or partial opening of the ring construction resulting from a water-mediated hydrolysis [20].
Fig 8. Cyclic thiosulfinate S-XL6 cross-links wild-type SOD1 by way of Cys111 residues on adjoining monomers.
On the left crystal construction of as-isolated wild-type SOD1 at 1.77 Å decision (cartoon illustration generated with Pymol) highlighting opposing Cys111 residues on each monomer A (darkish inexperienced) and monomer B (darkish blue) with a number of water molecule current at dimer interface (PDB ID 8Q6M). Proper hand panels present the crystal construction of S-XL6 cross-linked wild-type SOD1 at 1.67Å decision (PDB ID 8CCX) clearly displaying extra density for S atoms from S-XL6 on each monomers A (mild inexperienced) and B (cyan). 2FoFc electron density maps for the buildings are proven as grey mesh, contoured at 1 σ stage round Cys111, water molecules on the SOD1 dimer interface and cross-linker. Waters are indicated as small purple spheres, Cu and Zn molecules are represented by cyan and orange spheres, respectively. These water molecules are largely excluded upon the incorporation of S-XL6. The info underlying this determine could be discovered at rcsb.org (PDB IDs 8Q6M and 8CCX). SOD1, superoxide dismutase 1.
Earlier research have proven that SAXS can monitor modifications to protein tertiary and quaternary construction in resolution. Due to this fact, SAXS (Fig 9E and 9F) was carried out to find out the consequences of cross-linking upon the construction of fALS variant SOD1G93A, the identical variant expressed by the mice that have been dosed within the current examine. Whereas earlier SAXS research of SOD1G93A preparations confirmed a radius of gyration (Rg) of 20 to 24 Å [14,47], our SAXS information for as-isolated and untreated SOD1G93A couldn’t be match to find out an Rg, in all probability because of the aggregation of SOD1. The Rg of 20 Å estimated from scattering information was in keeping with S-XL6 handled SOD1G93A being a local dimer.
Fig 9. S-XL6 cross-links SOD1 in an ALS mouse mannequin and reduces aggregation in vitro.
(A) Schematic of the pharmacodynamic workflow. (B) Pharmacodynamic profiling of RBC proteins by LC-MS evaluation signifies the formation of an S-XL6 cross-linked dimer in handled SOD1G93A mice. Hemizygous fALS SOD1G93A mice have been dosed as soon as at 10 mg/kg with S-XL6 by way of tail vein injection, blood was collected periodically over a 7-day interval, and LC-MS evaluation (C and D) was carried out to evaluate proportion of cross-linked SOD1 (particulars in S1 Table). Spectra signify a 6-s common at peak apex. (E and F) In vitro SAXS experiments for SOD1G93A: Semi-log plot of the scattering depth (I, log scale) as a perform of momentum switch, q; and Kratky plot [q2•I(q) versus q]. The plot form for untreated SOD1G93A is in keeping with a heterogenous combination of unfolded proteins together with amorphous aggregates, whereas plot form for cross-linked SOD1G93A is in keeping with a folded, globular construction. The estimated radius of gyration is roughly 20 Å. (G) A 3-D reconstruction of cross-linked SOD1G93A is superimposed upon the SOD1G93A crystal construction (PDB: 3GZO). (H) Goal engagement by way of LC-MS evaluation in transgenic (Tg) SOD1G93A mouse mind dosed as soon as at 30 mg/kg with S-XL6 by way of subcutaneous injection. The info underlying this determine could be present in S1 Data. ALS, amyotrophic lateral sclerosis; fALS, familial amyotrophic lateral sclerosis; LC-MS, liquid chromatography-mass spectrometry; RBC, purple blood cell; SOD1, superoxide dismutase 1.
Pharmacodynamic profiling, stabilization of SOD1G93A in vivo, and blood–mind barrier penetration
To evaluate the time course of S-XL6 goal engagement in vivo, pharmacodynamic (PD) profiling was carried out utilizing B6 G93A SOD1 mice [13]. Mice have been dosed with S-XL6 by way of intravenous (IV) injection and blood was collected at totally different time factors, publish dose, from the tail vein. To detect the intact SOD1G93A protein-cross-linker complicated, we utilized a facile MS assay, particularly a mixture of solvent-extraction, hemoglobin precipitation, and liquid chromatography coupled with mass spectrometry (LC-MS) [48]. We noticed {that a} single IV dose of S-XL6 at 10 mg/kg transformed 63% of the SOD1G93A (Fig 9D) right into a cross-linked dimer in blood at 1-h post-dose by the supposed MOA (Fig 1), whereas untreated mice confirmed solely SOD1 monomer (Fig 9C). A half-life enchancment was noticed for cross-linked SOD1G93A (Fig 9B, 26 h 1-compartment mannequin, 11 and 68 h utilizing a 2-compartment mannequin) in comparison with the 10-h half-life reported for untreated SOD1G93A in vivo [49]. Subsequent, we assessed S-XL6’s goal engagement in SOD1G93A mouse mind (Fig 9H). Dosed (30 mg/kg by way of SC) mice confirmed 86% conversion of SOD1 to cross-linked dimer at 1 h.
Survival research
Two fALS traces have been used to evaluate survival in genetic backgrounds with totally different illness severity. The hybrid “B6SJL G93A mice” are generally used for fALS SOD1 survival research, exhibit earlier onset (roughly 125 days), have a blended B6/SJL background, and are typically much less inclined to environmental stress. A second and congenic line was obtained by crossing (>10 technology) fALS “B6 G93A mice” with “YFP-16 mice” [50], expresses YFP in motor and sensory neurons to allow additional strategies growth for goal engagement profiling at cellular-resolution [51], exhibit later onset (roughly 155 days), and has a pure C57BL/6J genetic background. Dose and dose routine have been estimated from the blood PK/PD profile and BBB penetration described above. A subcutaneous (SC) route of administration was chosen as a result of in our fingers it ends in extra exact supply. After an preliminary (SC) dose of 20 mg/kg B6SJL G93A mice, the PK/PD profile demonstrated decrease goal engagement than anticipated (Figs 10 and S8) and the dose was elevated to 50 mg/kg (dosed as soon as a day, Monday to Friday) from day 70 onward. This dose offered no survival profit in B6SJL G93A mice. The identical dose, ranging from day 108, offered a modest survival profit in B6 G93A mice (153 +/− 15 days, management; 169 +/− 11 (STD) days dosed). In each research, mice have been gender and littermate matched and non ALS-related deaths have been excluded. We’re at the moment creating the strategies obligatory for quantifying copy quantity and can’t rule out decreased SOD1 expression as having contributed to the survival profit in handled B6 G93A mice. Given this and that the B6SJL G93A outcomes have been obtained in a regular genetic background by an skilled laboratory (Brown) and the B6 G93A outcomes have been obtained in a novel line (with YFP expressed in motor neurons) by a much less expertise laboratory (Agar), we interpret these survival outcomes conservatively as there having been modest to no total profit (or toxicity) from S-XL6 administration when roughly 40% of SOD1 is engaged (on common) over time. As detailed under, we consider that there’s room for significantly improved goal engagement in mice, however this may require improved bioanalytical strategies for covalent drug PK/PD evaluation.
Fig 10. Survival research of fALS mice handled with subcutaneously administered S-XL6.
(a) Therapy with S-XL6 in B6SJL G93A (hybrid line) mice started at age 60 days in cohorts of 40 mice (18 untreated and 18 handled till end-stage; 2 mice from every group dosed for ca. 50 days earlier than being killed for goal engagement evaluation in mind). After 1 week at a 20 mg/kg dose goal engagement in blood was monitored as proven in Fig 9. Based mostly upon these outcomes, to enhance goal engagement to >50% peak impact, the dose was elevated to 50 mg/kg. No survival enchancment was noticed in B6SJL mice. Physique weight, illness rating, and all paws’ grip rating evaluation for B6SJL mice are given in S8 Fig. (b) B6 G93A/YFP (congenic line) mice have been dosed on weekdays ranging from day 108 to endpoint and demonstrated a modest survival enchancment. Instantly after dosing mice exhibited lethargy and shaking that subsided after 20 min. The info underlying this determine could be present in S1 Data. fALS, familial amyotrophic lateral sclerosis.
Dialogue
We’ve got proven {that a} cyclic thiosulfinate cross-linker, S-XL6, binds to Cys 111, cross-links monomers, and stabilizes the dimer of SOD1 fALS variants with numerous physicochemical properties and illness severities. Notably, S-XL6 has promising preclinical properties, together with: oral bioavailability, avoiding plasma proteins, low toxicity (c.a. 1 g/kg/day LD50 by way of PO), conforming to Lipinski’s guidelines, and exhibiting BBB penetration. Notably, remedy with S-XL6 extends the organic half-life of SOD1G93A [49]. The big diploma of stabilization of fALS variants by S-XL6 is promising. First, lack of thermodynamic stability is the biggest epidemiological danger issue for fALS. Second, lack of thermodynamic stability (hazard ratio = 24) and achieve of aggregation propensity (hazard ratio = 13) act synergistically (mixed hazard = 333), indicating that one has an impact upon the opposite [9,52]. That is anticipated on condition that protein misfolding and decreased thermodynamic stability typically enhance the speed for the nucleation part of aggregation. S-XL6 promoted the aggregation of EGFP-conjugated wild-type SOD1, however not SOD1G93A, within the NSC-34 cell line, and had no impact on the survival. Our outcomes point out that S-XL6 remedy offers modest (congenic G93A line) to no (hybrid G93A line) survival profit in fALS SOD1 mouse fashions. We consider that additional research are obligatory to completely assess the therapeutic potential of cyclic thiosulfinate-mediated fALS SOD1 stabilization (and kinetic stabilization of SOD1). First, the pharmacological burden of stabilizing SOD1 in a mouse mannequin is far greater than in people, as a result of mice specific 28 copies of G93A SOD1 and presumably require 14 instances greater dose than people. Second, fALS SOD1 sufferers (aside from D90A homozygotes) specific a fALS SOD1 variant and wild-type SOD1, that are believed to work together. Lastly, the bioanalytical strategies and mathematical fashions required to evaluate and optimize goal engagement of covalent medication (and S-XL6) in vivo are at the moment inadequate. That is the results of the established uncoupling of the pharmacokinetics and pharmacodynamics (e.g., lack of correlation between free drug focus and impact) for medication that don’t reversibly bind their targets. We’re at the moment creating the required analytical strategies and mathematical fashions to fill this want and are addressing the protection and selectivity profile of S-XL6. Regardless of its limitations, this examine offers necessary benchmarks for additional compound growth, and the primary chemical device able to testing the therapeutic speculation that kinetic stabilization of the fALS SOD1 dimer stabilization will enhance survival. The simplest preclinical therapeutic technique for SOD1-related fALS has been the stabilization of SOD1 by way of the CuATSM (Cu (II)-diacetyl-bis(N(4)-methylthiosemicarbazone))-mediated incorporation of metals [53]. Provided that S-XL6 improves metallic binding by stabilization, and CuATSM delivers metals, we anticipate a profit to combining these approaches.
Supplies and strategies
Ethics assertion
Work with vertebrate animals have been carried out at Northeastern College and UMass Chan Medical College and have been carried out in accordance with the Information for the Care and Use of Laboratory Animals (Nationwide Institutes of Well being, Bethesda, Maryland, United States of America). All strategies and animal manipulations carried out by the Agar laboratory have been accepted by the Northeastern College Institutional Animal Care and Use Committee (IACUC) below protocol #16-0303-R and manipulations carried out by the Brown laboratory have been accepted by the College of Massachusetts Chan Medical Institutional Animal Care and Use Committee below protocol #201900287. No human experiments have been carried out.
Synthesis of cyclic thiosulfinate 1,2-dithiane-1-oxide (S-XL6)
Synthesis of 1,2-dithiane-1-oxide (S-XL6) was achieved in accordance with the beforehand revealed literature process [28]. To a round-bottom flask was added I2 (2.08 g, 8.18 mmol, 0.20 equiv.) and DMSO (2.91 mL, 40.90 mmol, 1.0 equiv.). The ensuing darkish brown resolution was stirred gently. 1,4-butanedithiol (4.76 mL, 40.90 mmol, 1.0 equiv.) was dissolved in CH2Cl2 (16.3 mL) and added dropwise to the stirring resolution of I2 in DMSO (Fig 11). The speed of stirring was elevated regularly with the addition of 1,4-butanedithiol in CH2Cl2. Upon completion of addition the ensuing resolution was stirred for 1 h at 23°C. The sunshine brown resolution was quenched with gradual addition of 10% aq. Na2S2O3 (25 mL). The layers have been separated, and the aqueous layer was extracted with CH2Cl2 (3 × 50 mL). The mixed natural layers have been concentrated below decreased strain. The concentrated light-yellow oil was taken up in 100 mL EtOAc and washed with sat. aq. NaCl (50 mL). The layers have been separated, and the aqueous layer was extracted with EtOAc (3 × 75 mL). The mixed natural layers have been dried over Na2SO4 and concentrated below decreased strain to offer pure 1,2-dithiane (4.82 g, 98% yield) as a faint yellow stable. Rf = 0.80 (5:1 hexanes:EtOAc). 1,2-dithiane was then used with out additional purification to type 1,2-dithiane-1-oxide (S-XL6). In a round-bottom flask, 1,2-dithiane (4.82 g, 40.09 mmol, 1.0 equiv.) was dissolved in CH2Cl2 (40.0 mL) and cooled to 0°C. An answer of mCPBA (9.48 g, 73% weight, 40.09 mmol, 1.0 equiv) in CH2Cl2 (100.0 mL) was added dropwise by way of addition funnel. The ensuing colorless, cloudy resolution was allowed to stir at 0°C for 1 h whereas warming to 23°C and allowed to stir 1 h at 23°C. The response was cooled once more to 0°C and quenched with stable Na2CO3 (40.0 g, 377 mmol, 1 g per mmol mCPBA). The ensuing dense slurry was filtered by a brief plug of celite and concentrated below decreased strain. The ensuing cloudy oil was purified by flash column chromatography on silica gel with 2% MeOH/CH2Cl2 to offer S-XL6 (3.53 g, 64% yield) as a colorless stable. Rf = 0.33 (2% MeOH / CH2Cl2). 1H NMR (500 MHz, CDCl3, δ): 1.82–1.91 (m, 1H), 1.95–2.08 (dtt, J = 14.0, 12.8, 3.0 Hz, 1H), 2.09–2.17 (m, 1H), 2.60–2.73 (m, 2H), 3.03–3.13 (dt, J = 13.2, 3.0 Hz, 1H), 3.17–3.24 (td, J = 13.4, 3.7 Hz, 1H), 3.61–3.71 (ddd, J = 14.0, 12.0, 2.5 Hz, 1H) (S4 Fig); 13C NMR (100 MHz, CDCl3, δ): 15.29, 23.47, 25.74, 51.92 (S5 Fig). [M+H]+ was noticed for C4H8OS2 and located 137.00914 Da (S6 Fig) by way of Bruker 9.4T SolariX XR Mass Spectrometer (Bruker, Billerica, Massachusetts, USA). Melting level discovered to be 83–86°C (lit 83–86°C).
Expression and purification of wild-type SOD1, SOD1A4V, SOD1G93A, SOD1H46R, and SOD1G85R
Expression and purification of SOD1 have been carried out as beforehand revealed [22]. Briefly, EGy118ΔSOD1 yeast have been remodeled with a wild-type SOD1, SOD1A4V, SOD1G93A, SOD1H46R, or SOD1G85R YEp351 expression vector and grown at 30°C for 44 h. Cultures have been centrifuged, lysed in a blender utilizing 0.5 mm glass beads, and subjected to a 60% ammonium sulfate precipitation. Then, the pattern was centrifuged, and the ensuing supernatant was diluted to 2.0 M ammonium sulfate. The diluted pattern was handed by a phenyl-sepharose 6 quick circulation (excessive sub) hydrophobic interplay chromatography column (Cytiva Life Sciences, Marlborough, Massachusetts, USA) utilizing a linearly reducing salt gradient from excessive salt buffer (2.0 M ammonium sulfate, 50 mM potassium phosphate dibasic, 150 mM sodium chloride, 0.1 mM EDTA, 0.25 mM DTT (pH 7.0)) to low salt buffer (50 mM potassium phosphate dibasic, 150 mM sodium chloride, 0.1 mM EDTA, 0.25 mM DTT (pH 7.0)) over 300 mL. Fractions containing SOD1 eluted between 1.6 and 1.1 M ammonium sulfate and have been confirmed with SDS-PAGE. These fractions have been pooled and exchanged into low salt buffer (10 mM Tris (pH 8.0)). Pooled fractions have been then handed by a Mono Q 10/100 anion alternate column (Cytiva Life Sciences, Marlborough, Massachusetts, USA) utilizing a linearly growing salt gradient from low salt buffer to excessive salt buffer (10 mM Tris (pH 8.0), 1 M sodium chloride) from 0% to 30%. SOD1 fractions have been collected between 5% and 12% excessive salt buffer and have been confirmed with SDS-PAGE, western blot, and Fourier-transform ion cyclotron resonance mass spectrometry (FT-ICR-MS).
Affirmation of cross-link formation in vitro
Wild-type SOD1, SOD1A4V, SOD1G93A, SOD1H46R, and SOD1G85R inventory options have been diluted to 40 μm in 10 mM ammonium acetate (pH 7.4). DMSO inventory of S-XL6 was used to arrange 400 μm (10×) in 10 mM ammonium acetate (0.5% DMSO). Protein and cross-linker samples have been mixed in equal volumes (last focus 20 μm SOD1, 200 μm cross-linkers, 0.25% DMSO) and incubated at 37°C for 4 h at 350 rpm. Full cross-linking was confirmed by mass spectrometry on a 9.4T Bruker SolariX (Bruker Company, Billerica, Massachusetts) by way of direct infusion as beforehand described [54,55]. Previous to infusion, samples have been diluted to 1 μm in 50:50 acetonitrile:water, with 0.1% formic acid. Throughout evaluation, 32 scans have been acquired in optimistic mode and averaged. Funnel 1 and skimmer 1 have been stored round 150 V and 20 V, respectively, and funnel RF amplitude was held at 60.0 Vpp.
Differential scanning fluorimetry
The impact of cross-link formation on the thermal stability of SOD1 was decided by differential scanning fluorimetry as beforehand described [22]. Wild-type SOD1, SOD1A4V, SOD1G93A, SOD1H46R, and SOD1G85R (20 μm in 10 mM ammonium acetate (pH 7.4)) have been incubated with 10-fold molar extra S-XL6 for 4 h at 37°C in protein low bind Eppendorf tubes utilizing Eppendorf Thermomixer at 350 rpm (Eppendorf North America, Enfield, Connecticut, USA). After incubation, SYPRO Orange (Invitrogen Company, Carlsbad, California, USA), an environmentally delicate fluorescent dye which is quenched in an aqueous setting however turns into unquenched as soon as it binds hydrophobic residues, was added to the response combination to a last focus of 200×. The samples have been then transferred to a quick optical 96-well response plate (Utilized Biosystem, Life Applied sciences Company, Carlsbad, California, USA) and loaded on to the real-time PCR machine (Utilized Biosystem, Life Applied sciences Company, Carlsbad, California, USA) after plate was spun all the way down to remove bubbles. The Soften curve template in StepOne Plus software program was used to arrange a way. The SYBR reporter and ROX quencher information have been collected for 35 cycles with a temperature gradient starting from 25 to 99.9°C at a ramp price of 0.5°C/min. The ultimate focus of DMSO within the response combination was checked because it contributes to the denaturation of proteins. Dilutions have been carried out accordingly to take care of the ultimate focus of DMSO lower than 1% (vol/vol). All samples have been run in technical triplicates. The background information was subtracted from the common relative fluorescence and common derivate for every temperature level, and the information have been normalized earlier than Tus have been quantified by averaging the detrimental first spinoff of relative fluorescence of all 3 runs and figuring out the native minima.
Crystallographic construction dedication
Crystals of SOD1 grown by mixture of hanging drop methodology with micro seeding. For crystallization 2 μl of 10 mg/ml of recombinant wild-type SOD1 have been blended with 2 μl of crystallization resolution containing 2.5 M ammonium sulfate 150 mM NaCl, 50mM acetate buffer (pH 4.75) and equilibrated over 400 μl properly containing the identical crystallization resolution. To make S-XL6 complicated, crystals that seem as plates have been soaked for two h in stabilizing resolution (3 M ammonium sulfate 150 mM NaCl Acetate buffer (pH 4.75)) blended with DMSO inventory of fifty mM S-XL6 in 1:16 ratio, to last focus of S-XL6 3 mM. For the as-isolated wild-type SOD1 Paratone-N oil was used as a cryo protectant. X-ray information was collected utilizing the FRE+ X-ray supply coupled to an EIGER R 4M photon counting detector at 1.54 Å wavelength to 1.67 Å decision for S-XL6 complicated and to 1.77 Å decision for as-isolated wild-type SOD1. Knowledge have been processed with Mosflm [56]. Constructions have been solved by molecular alternative with Molrep [57] utilizing beginning mannequin 2C9V with following isotropic refinement in CCP4i2 with refmac5 [58]. The cycles of refinement have been iterated with mannequin rebuilding in COOT [59]. Closing buildings have been validated with Molprobity [60] and submitted to PDB ID with 8CCX for S-XL6 complicated and 8Q6M for as-isolated wild-type SOD1, used as management. Knowledge assortment and refinement statistics are summarized in S2 Table.
Hep G2 cell tradition
The Hep G2 human hepatocarcinoma cell line (American Kind Tradition Assortment, ATCC, Manassas, Virginia, USA) was cultured utilizing EMEM (Eagle’s minimal important medium, ATCC) supplemented with 10% FBS (fetal bovine serum, ATCC) and 1% penicillin streptomycin as beforehand described [45] (10,000 items/mL penicillin and 10,000 μg/mL streptomycin, Fisher scientific, Hampton, New Hampshire, USA). The cells have been grown and subcultured round 72 h at 37°C, 5% CO2 below managed humidity. The cells development and morphology have been inspected utilizing an inverted Carl Zeiss microscope (Carl Zeiss Microscopy LLC, White Plains, New York, USA).
Cell viability assay/cell cytotoxicity assay
Utilizing producer’s directions, Hep G2 cells have been cultured for twenty-four h with a density of two × 105 cells/mL in a 96-well tradition plate at 37°C, 5% CO2 below managed humidity. The cells have been handled with S-XL6 ranging with 10, 100, 150, 250, 400, 600, 800, and 1,000 μm for twenty-four h (0.1% DMSO). Cell cytotoxicity was carried out by way of the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay at 570 nm absorbance utilizing BioTek synergy H1 (Vermont, USA) plate reader in accordance with beforehand revealed paper [45]. The MTT inventory was made at 5 mg/mL utilizing 0.9% sodium chloride resolution which was bought from Sigma Aldrich (St. Louis, Missouri, USA). Twenty-four hours after dosing the plate, the aesthetic medium was aspirated and changed with the equal quantity of MTT resolution (10× dilution utilizing EMEM full media). The plate was incubated for 4 h (37°C, 5% CO2). The MTT resolution was changed by the equal quantity of acidified isopropanol (0.3% hydrochloric acid (v/v)) after which fashioned formazan was dissolved by shaking the plate gently for 30 min. The absorbance was measured at 570 nm for each the samples and acidified isopropanol for background calculation. Chlorpromazine and cells with 0.1% DMSO have been used as optimistic management and car, respectively. The cell viability of every focus was measured by normalizing towards imply worth of cells with 0.1% DMSO (car).
SDS-PAGE and western blot
The Hep G2 human hepatocarcinoma cell line (American Kind Tradition Assortment, ATCC, Manassas, Virginia, USA) was cultured utilizing full EMEM (Eagle’s minimal important medium, ATCC) media together with 10% FBS (fetal bovine serum, ATCC) and 1% penicillin streptomycin (10,000 items/mL penicillin and 10,000 μg/mL streptomycin, Fisher Scientific, Hampton, New Hampshire, USA) [28]. Cells have been counted and checked for morphology utilizing an inverted Carl Zeiss microscope (Carl Zeiss Microscopy LLC, White Plains, New York, USA). A 96-well plate (Corning Life Sciences, Tewksbury, Massachusetts, USA) was used for culturing cells at 0.2 mill/mL focus for twenty-four h at 37°C, 5% CO2 below managed humidity for western blot experiment. Cells have been dosed with S-XL6 in triplicate (wells) at totally different concentrations (0.001 μm by 200 μm) and incubated for 30 min by changing the prevailing EMEM with PBS buffer. Wild-type SOD1 purified from human erythrocyte (Sigma Aldrich, St. Louis, Missouri, USA) was used as optimistic management (customary) within the experiment. The expansion buffer was aspirated and changed with 30 μL of lysis buffer (150 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate, 0.1% SDS, and 50 mM Tris (pH 8.0), all chemical compounds have been bought from Sigma Aldrich (St. Louis, Missouri, USA)) and incubated for 10 min adopted by addition of non-reducing pattern buffer. Samples have been loaded right into a Bio-Rad mini TGX gel (Bio-Rad Life Sciences, Hercules, California, USA) together with PageRuler prestained protein ladder (Thermo Fisher Scientific, Waltham, Massachusetts, USA) and ran utilizing a Bio-Rad electrophoresis cell. The gel was eliminated rigorously and transferred right into a beaker stuffed with switch buffer (25 mM Tris, 192 mM glycine, 0.1% SDS, and 10 mM β-mercaptoethanol, all chemical compounds have been bought from Sigma Aldrich (St. Louis, Missouri, USA)) and incubated briefly at 90°C after which transferred to a Trans-Blot Turbo mini nitrocellulose membrane utilizing Trans-Blot Turbo Switch system (Bio-Rad, Hercules, California, USA). The membrane was incubated in a single day in antibody buffer (50 mM Tris-HCl, 150 mM sodium chloride, 0.1% tween-20, 2.5% dry milk, all chemical compounds have been bought from Sigma Aldrich (St. Louis, Missouri, USA)) with Cu/Zn SOD polyclonal antibody (1:1,000 dilution, Enzo Life Sciences, Farmingdale, New York, USA). The membrane was then incubated with HRP-linked antibody buffer (1:1,000 dilution, Cell Signaling Know-how, Danvers, Massachusetts, USA). The membrane was incubated briefly in the dead of night with enhanced chemiluminescent (ECL, Thermo Fisher Scientific, Waltham, Massachusetts, USA), adopted by imaging utilizing BioRad Chemidoc MP Imaging System (Bio-Rad, Hercules, Massachusetts, USA).
Gel-based SOD1 enzymatic exercise assay
Enzymatic exercise of SOD1 variants have been assessed utilizing a gel-based assay with riboflavin and nitroblue tetrazolium (NBT) as beforehand described [61,62]. Wild-type SOD1, SOD1A4V, SOD1G93A, SOD1H46R, and SOD1G85R have been incubated with or with out 10-fold molar extra S-XL6 at 37°C for 4 h at 350 rpm in 10 mM ammonium acetate (pH 7.4) (DMSO content material <1%). The samples have been blended with native pattern buffer (Bio-Rad) in a 1:1 ratio and have been loaded into every properly to a 12% polyacrylamide gel (Bio-Rad). The gel was stained in darkish utilizing 50 mM potassium phosphate (pH 7.8), 0.13 mg/mL of NBT, and 0.1 mg/mL of riboflavin for 45 min. The gel was uncovered to mild for with 1 μL/mL of N,N,N′,N′-tetramethylethylenediamine (TEMED) for two min. SOD1 exercise could be seen as colorless bands because the SOD1 scavenges the superoxide that inhibits insoluble blue coloration formazan formation.
Chemical-based SOD1 enzymatic exercise assay
All samples have been ready as described in gel-based enzymatic exercise assay part and have been (10 μL) blended with 160 μL of resolution A (6,000 items/mL catalase, 10 mM hypoxanthine, and 5 mM XTT) and 160 μL of resolution B (xanthine oxidase 16 mg/mL) [61] in a flat-bottom 96-well plate and was monitored for absorbance at 490 nm for 30 min. All samples have been analyzed in technical triplicate utilizing slope perform in Microsoft Excel.
Measurement exclusion chromatography (SEC)
We’ve got analyzed samples with and with out cross-linker by way of SEC with excessive efficiency liquid chromatography (SEC-HPLC) to evaluate the SOD1 protein composition in its native state [63,64]. Wild-type SOD1, SOD1G93A, SOD1A4V, SOD1H46R, and SOD1G85R (20 μm inventory) have been incubated with or with out 10-fold molar extra S-XL6 in 200 mM ammonium acetate (pH 7.4) or 4 h at 350 rpm, and every cross-linked dimer have been examined utilizing a Bruker 9.4T FT-ICRMS mass spectrometry. For inducing aggregation, all samples (last conc. 1 μm) have been incubated at 37°C for 1 h prior the SEC evaluation. An Agilent HPLC 1290 Infinity II LC system was used for duplicate evaluation of every pattern sort by injecting 5 μL pattern by way of Waters Acquity protein BEH SEC column (125 Å, 1.7 μm, 4.6 mm × 150 mm, Waters Company, Milford, Massachusetts, USA). An isocratic cellular part consisting of 100 mM sodium phosphate, 200 mM sodium chloride (pH 7.4) was used for six min to investigate every pattern at wavelength of 280 nm. Waters BEH 200 customary was injected firstly, center, and on the finish to verify the retention time was not shifting all through the complete experiment.
NSC-34 cell culturing and transfection
Mouse neuroblastoma/motor neuron hybrid cells (NSC-34) [65] have been cultured and maintained as beforehand described [42]. Briefly, NSC-34 cells have been cultured in Dulbecco’s Modified Eagle’s Medium with F12 complement (DMEM/F12, Invitrogen Australia) containing 10% (v/v) heat-inactivated fetal bovine serum (FBS, Bovogen Biologicals, Australia) and a pair of mM L-Glutamine (Invitrogen, Australia). Cells have been passaged utilizing 0.25% trypsin, 0.02% EDTA (Invitrogen, Australia). Cells have been maintained inside humidified incubators at 37°C with 5% CO2.
To transfect NSC-34 cells, they have been first plated out to 40% confluency in both 6-well or 12-well tissue culture-treated plates and cultured for twenty-four h. Following this, TransIT-X2 reagent (Mirus Bio, USA) was used in accordance with the producer’s directions to transfect NSC-34’s (1 μg DNA per properly for 12-well plates, 2.5 μg per properly for a 6-well plate). Cells have been then incubated for 8 h previous to replating for experiments into different plates or chambers.
Immunoblotting for SOD1-EGFP
Transfected NSC-34 cells have been plated out into 12-well plates and left in a single day. The next day, cells have been handled with S-XL6 (20 μm) or car (0.04% v/v DMSO) and incubated for one more 24 h. Cells have been then harvested by way of aspiration and pelleted (300 × g). Lysis and immunoblots have been carried out as beforehand described [33]. Briefly, cell pellets have been lysed in ice-cold lysis buffer containing protease inhibitors (Thermo Fisher, USA) and N-ethylmaleimide (NEM) at a last focus of 1 mg/mL. Protein focus was measured by DC assay (BioRad, Australia) and samples have been denatured in SDS-sample buffer containing 40 mM NEM at 90°C for five min. Lysates have been loaded into 26-lane stain-free gels (BioRad, Australia) and electrophoresed. Proteins have been transferred to 0.2 μm PVDF membranes for 1 h at 4°C. Membranes have been blocked in 5% skim milk in TBST for 1 h previous to addition to main antibody which as a rabbit anti-GFP antibody (ab290, 1:5,000, Abcam USA) and incubated in a single day at 4°C. Membranes have been washed 3× after which incubated for 1 h at room temperature in secondary antibody (goat anti-rabbit HRP conjugate at 1:5,000 (catalog no.: P0448; Dako). Membranes have been then washed 3× previous to creating with SuperSignal West Pico Plus substrate (Thermo Fisher Scientific).
Willpower of cells containing inclusions
Transfected NSC-34 cells have been plated out into 96-well plates and left in a single day. The next day, cells have been handled with varied concentrations of S-XL6 (1.25–20 μm) or car (0.0025% to 0.04% v/v DMSO) and incubated for one more 48 h. Cells have been then mounted utilizing 4% (w/v) paraformaldehyde in 1×PBS for 20 min. Plates for imaging have been then processed as beforehand described [33] with some modifications. Cells have been imaged utilizing a DMi8 epifluorescent microscope (Leica, Germany) fitted with a Mercury Arc Lamp, a ten× 0.25NA (CAT NO) air goal and an automatic stage [33]. A software-based autofocus technique was utilized to concentrate on the GFP sign inside cells, with a subsequent picture being taken within the DAPI channel to view Hoescht-stained nuclei. Relative focus correction was utilized between channels to offset emission wavelength results to the focal airplane. Following picture acquisition, photos have been corrected for illumination variation by way of a background methodology and processed utilizing a mixture of FIJI [66], StarDist, CellProfiler [66], and CellProfiler Analyst [67] as beforehand described [33].
Willpower of relative cell viability and relative SOD1-EGFP ranges
Picture set information taken from the earlier inclusion formation assay have been used to look at each relative cell viability and relative ranges of SOD1-EGFP. To find out relative viability, we extracted the entire variety of transfected cells in every remedy group and normalized the values to the untreated group to find out the relative variety of cells at assay endpoint.
For EGFP stage evaluation, a part of the information evaluation used to find out which cells have or should not have inclusions is the measurement of imply cell fluorescence depth within the GFP channel. This worth was extracted from the imaging information and plotted as a imply per drug remedy.
Mouse dosing and SOD1 isolation
Pharmacodynamic profiling was carried out utilizing hemizygous mice expressing human SOD1G93A (“fast-line” Jackson Laboratory; B6SJL-Tg(SOD1*G93A)1Gur/J, often known as SOD1-G93A stock- 002726) [13]. Mice have been bred to precise YFP in neurons in anticipation of BBB-penetration assays and Matrix-Assisted Laser Desorption Ionization Mass Spectrometry Imaging (MALDI MSI) evaluation in accordance with the revealed protocol [68,69]. Extraction of SOD1G93A from purple blood cell (RBC) protein was carried out as beforehand described [48]. S-XL6 was ready in 1× PBS and mice have been dosed at 10 mg/kg by way of IV injection of their lateral tail vein slowly and gently. Mice have been trapped in restrainer and round 40 μL of blood was collected in Greiner Bio-one Okay3EDTA tubes (Greiner Bio-one, North Carolina, USA) publish injection at 30 min, 1 h, 2 h, 4 h, 8 h, 12 h, 24 h, 48 h, 72 h, and 168 h. Tubes have been centrifuged at 2,000 rpm at 4°C for five min instantly after accumulating blood. Fastidiously plasma was discarded and acid citrate dextrose resolution (0.48% citric acid, 1.32% sodium citrate, 1.47% glucose, all chemical compounds have been bought from Sigma Aldrich (St. Louis, Missouri, USA)) was used to clean the pattern by centrifuging at 2,000 rpm at 4°C for five min. Following wash, the supernatant was eliminated, and RBC have been lysed by the addition of 8× equal of 10 mM ammonium acetate. To the hemolysate, 0.15 equivalents of chilly chloroform and 0.25 equivalents of chilly ethanol was added. Samples have been vortexed at 1,800 rpm at 4°C for 15 min and centrifuged at 12,000 rpm for 10 min. The supernatant was collected and saved at −80°C after flash freezing for LC-MS evaluation. Previous to LC-MS evaluation, samples have been acidified to 10% formic acid.
Affirmation of S-XL6 cross-linked SOD1G93A dimer formation in vivo
In vivo cross-linking from purified RBC was confirmed utilizing an H-Class Acquity UPLC (Extremely Efficiency Liquid Chromatography) system coupled to a Xevo G2-S Q-ToF (Quadrupole Time of Flight) mass spectrometer (Waters Corp, Milford, Massachusetts, USA) as beforehand described [55,70]. The LC system was outfitted with reversed part Acquity UPLC Protein BEH C4 (300 Å pore measurement, 1.7 μm particle measurement, 100 mm mattress size, 2.1 mm ID × 100 mm) column at 60°C with a circulation price of 0.2 mL/min. The cellular part consisted of a combination of 0.1% formic acid in water (solvent A) and 0.1% formic acid in acetonitrile (solvent B). The pattern was launched in 10% formic acid and 5 μL was injected for evaluation. UNIFI software program (Waters Corp, Milford, Massachusetts, USA) was used for system management and information processing. Solvents A and B have been mixed in a gradient: 0 to 2 min: 95% A; 2 to 70 min: 30% A; 72 to 75 min: 5% A; 78 to 80 min: return to preliminary situations. The MS was operated in optimistic mode and calibrated previous to evaluation (ca. every day). The chromatographic window containing the SOD1G93A monomer and dimer have been assigned utilizing EICs (Extracted Ion Chromatogram) m/z 1322.5 to 1323.5 (monomer) and m/z 1180.0 to 1180.5 (dimer) and mass spectra from this area have been summed. Uncooked MS information from this composite spectrum have been deconvoluted and common plenty have been calculated utilizing the MaxEnt1 algorithm. The ratio of dimer depth (31836 Da, SD 1.3 Da) to the sum of monomer (15858 Da, SD 0.8 Da) plus dimer depth have been then used to calculate the proportion of SOD1G93A dimer at totally different time factors, and 10 picomole of the SOD1G93A monomer and dimer have been analyzed (as above) in particular person experiments and their MS sign intensities have been inside 10% (i.e., the variations of their chromatographic retention and ionization effectivity have been inside experimental error). The half-life of the S-XL6 cross-linked complicated was calculated from obvious half-life of disappearance of the % dimer by linear regression of the terminal beta part (24 to 168 h time level).
Goal engagement in SOD1G93A transgenic mouse mind
Wild-type (non-transgenic) C57BL/6 and transgenic (Tg) SOD1G93A ALS mouse have been used to find out the SOD1 goal engagement in mind utilizing western blot assay. Non-transgenic mouse was used as a detrimental management, transgenic management mouse acquired 1× PBS solely by way of SC, and the transgenic-treated mouse acquired 30 mg/kg of S-XL6 by way of SC; 1 h post-dose the mind was collected by sacrificing the mice. The mind was then homogenized, and pattern was ready in an analogous approach as described within the earlier part of pharmacodynamic pattern preparation of this manuscript prior LC-MS evaluation.
Willpower of acute toxicity (LD50) in vivo
The acute toxicity examine was carried out utilizing C57BL/6 (Jackson Laboratories, Maine) by way of PO, IV, and SC to find out the LD50 (Table 1 and S7 Fig) of S-XL6 in accordance with the Bruce’s up/down methodology [71]. The management group (n = 3) was dosed with 1× PBS (PO/IV/SC) and 0.25 mg/kg of S-XL6 (PO/IV/SC) was the beginning dose for the remedy group. Any scientific abnormalities together with however not restricted to ruffled fur, unstable gait, extreme lethargy, and mortality was noticed after dosing. The post-dose washout interval was maintained for 48 h for each PO, IV, and SC route of administration. For greater PO dosing inventory, a suspension of 200 mg/mL of S-XL6 was made utilizing a car consisting of 1% (v/v) Tween 80 and 20% (v/v) PEG 400 in sodium citrate buffer (20 mM (pH 5.5)). The ultimate dosage type of S-XL6 was made just by vortexing for 1 min to get uniform micelles prior dosing.
In vivo examine approval
All research have been carried out in accordance with the Information for the Care and Use of Laboratory Animals (Nationwide Institutes of Well being, Bethesda, Maryland, USA). The protocols for working with mice have been accepted by the Northeastern College Institutional Animal Care and Use Committee (IACUC) and protocol for working with rats have been accepted by the Charles River Laboratories IACUC.
Small-angle x-ray resolution scattering (SAXS)
SAXS information have been collected on the G1 beamline at Cornell Excessive Vitality Synchrotron Supply (CHESS). For every pattern, 2 measurements have been taken, one of many protein with buffer and one with buffer by itself. Resolution scattering information have been captured each second for 10 frames. The ten frames of each buffer and protein have been then averaged and the buffer was subtracted out to get the scattering for the protein. Samples have been run in a 96-well plate and held at 4°C constantly. Knowledge assortment was within the scattering angle (q) vary of 0.008 to 0.71 Å-1 and processed utilizing the software program, RAW.
Preparation of SOD1 samples for SAXS evaluation
SOD1G93A was ready at 3 mg/mL (approx. 190 μm) in HEPES buffer (115 mM NaCl, 1.2 mM CaCl2, 1.2 mM MgCl2, 2.4 mM Okay2HPO4, 20 mM HEPES (pH 7.4)). Inventory options of S-XL6 was freshly made at 10 mM in HPLC grade methanol and diluted to 946 μm in HPLC grade water (5-fold focus of protein); 100 μL of three mg/mL protein was blended with 110 μL of 946 μm of S-XL6. Management pattern contained 5% MeOH (last conc. 2.5%). Samples have been then incubated at 37°C for six h to make sure full cross-linking. After incubation, extra compound was buffer exchanged out of every pattern utilizing a ten kDa MWCO ultrafiltration gadget, and 320 μL is diluted to fifteen mL into HEPES buffer, spun all the way down to roughly 500 μL, and resuspended in one other 15 mL of HEPES buffer. After the ultimate spin, samples have been eliminated and concentrated in a smaller ultrafiltration gadget and delivered to roughly 70 μL last quantity (4.7 μm). Samples have been flash frozen and saved at –80°C previous to evaluation.
SAXS information evaluation and reconstruction of molecular envelopes
Packages inside the ATSAS suite [72] have been used to find out the estimated radius of gyration and three-dimensional molecular envelopes for SOD1G93A with and with out S-XL6 utilizing the x-ray resolution scattering information. The GNOM program was used to guage the pair distribution plot utilizing an oblique Fourier rework. SOD1G93A with S-XL6 had a Dmax worth of roughly 79 Å, whereas untreated SOD1G93A had a a lot greater Dmax of over 200 Å resulting from protein unfolding. The GASBOR program was used to generate three-dimensional ab initio fashions of related beads to suit the GNOM information, with the variety of beads set roughly to the entire variety of amino acids within the SOD1 constructs. With the intention to assess the distinctiveness of those options, 10 bead fashions have been generated with none symmetry utilized, then in contrast and averaged. Figures have been produced utilizing CHIMERA [73] adopted by superimposition of envelopes.
Peptide-level hydrogen deuterium alternate mass spectrometry (H/D-X MS)
To five μL of 40 μm SOD1 pattern (in 10 mM ammonium acetate in H2O (pH 7.4)) was added 20 μL of 99% D2O pattern buffer (10 mM ammonium acetate in D2O (pH 7.3)), diluting the focus of the protein down to eight μm. For the reference pattern, HPLC grade water was added as a substitute of D2O buffer. Samples with D2O buffer have been incubated at 37°C for five publicity time factors: 15 s, 50 s, 500 s, 1 h, and 4 h. All alternate reactions occurred at 37°C, pH roughly 7.4 to imitate in vivo situations. The response combination was quenched by the addition of 25 μL of quench buffer (8 M Gunadinium Hydrochloride (GnHCL), 0.5 M tris(2-carboxyethyl) phosphine (TCEP), 0.2 M Citric Acid at pH 2.35), reducing the pH of the ultimate combination all the way down to pH 2.45, diluting the focus of protein within the pattern all the way down to 4 μm whereas concurrently denaturing the protein and lowering the disulfides. For the 15-s time level, to make sure well timed quenching of the alternate response, D2O buffer beforehand saved at 37°C was added and the incubation itself was carried out at room temperature. All samples preparations have been carried out in technical triplicate, flash frozen instantly after the quenching response and saved at −70°C till evaluation. Previous to evaluation, 50 μL of 0.1% formic acid in H2O was added to the pattern to scale back the GnHCL focus all the way down to 2 M, the really useful focus threshold for the pepsin column. This was instantly adopted by injecting the pattern onto a Waters UPLC system designed for H/D-X MS evaluation the place the samples have been digested, desalted, and separated, on-line. The digestion and trapping of peptides was carried out throughout a 3-min trapping step over an immobilized pepsin column with a circulation price of 100 μL/min in 0.1% formic acid and water at 10°C. The peptides have been trapped on an ACQUITY HSS T3 100 Å, 1.8 μm entice column (Waters Corp, Milford, Massachusetts, USA) maintained at 0°C. On the finish of the trapping step, inside the 0°C chamber, the circulation was directed to the ACQUITY HSS T3 100 Å, 1.8 μm analytical column (Waters Corp, Milford, Massachusetts, USA) at 75 μL/min (common again strain was round 7,500 psi). The analytical separation step was carried out over a 9-min gradient of 5% to 25% (0 to 7 min) of buffer B; 25% to 95% (7 to eight.5 min) of buffer B (buffer A, 0.1% formic acid in water, buffer B, 0.1% formic acid in acetonitrile). Eluate from the analytical column was directed right into a Waters QToF Synapt G2 HD mass spectrometer with electrospray ionization and lock-mass correction (utilizing the Glu-fibrinogen peptide). Blanks have been used between every pattern injection to make sure there isn’t a carryover of peptides between runs. Mass spectra have been acquired between 50 and a pair of,000 m/z, in optimistic polarity and backbone mode. Scan time was set to 0.5 s, cone voltage to 30 V, capillary was 3.5 kV, entice collision power was 6 V, and desolvation temperature of 175°C.
Previous to H/D-X MS experiments, pattern preparation and run situations have been optimized by various concentrations of GnHCL (2 M, 4 M, 8 M), incubation temperature with D2O buffer (4°C, 37°C), circulation price over the pepsin column and HPLC gradient. Situations listed above yielded one of the best sequence protection (larger than 98%) and backbone within the shortest run time. Moreover, to attenuate price of again alternate from D again to H, the pattern combination pH after the quench response was lowered to roughly 2.5 and majority of the run on the instrument was carried out at 0°C [74]. All H/D-X MS experiments have been carried out below an identical situations; due to this fact, deuterium ranges reported are relative and weren’t corrected for again alternate. Optimized pattern preparation and instrument situations to attenuate again alternate from deuterium to hydrogen and technical triplicate measurements for every pattern allowed for prime confidence comparability assessments between pattern sorts [75].
H/D-X MS information evaluation
Mass spectra and chromatography information have been acquired utilizing MassLynx (Waters Corp). The peptides have been recognized and confirmed by way of MS2 utilizing the PLGS software program. The PLGS generated peptide lists and MassLynx acquired mass spectra have been imported into the DynamX HDX information evaluation software program 3.0 (Waters Corp) for additional evaluation. Solely peptides that have been recognized throughout all 6 replicates of every SOD1 variant (instance: triplicates of untreated wild-type SOD1 and S-XL6 cross-linked wild-type SOD1) have been thought of. The utmost sequence size of peptides was set to 45, minimal peak depth set to 10,000, and most MH+ error set to 10 ppm. Sequence protection throughout all samples achieved was roughly 99% and the common redundancy for coated amino acids was 11. The mass spectra have been processed inside the DynamX software program by centroiding the isotopic distribution of assorted cost states for all of the peptides (usually +2, +3, +4). Deuterium uptake ranges have been measured by calculating the variations between the centroid of the deuterated peptide versus the undeuterated reference peptide. These mass shifts and variations are plotted towards the alternate time factors (S2 Fig for the SOD1A4V, SOD1H46R, SOD1G85R N and C terminal peptides). Closing proportion uptake was calculated by averaging the % uptake throughout technical triplicate measurements using overlapping peptides and recurring residues to elucidate residue stage uptake measurements, wherever potential. Three replicates of every pattern sort have been run on totally different days, and the usual deviation was roughly 3% or much less. A conservative threshold of 5% was set in the course of the comparative evaluation between pattern sorts and solely constant variations in % deuterium uptake above 5% have been emphasised. Peptic maps have been obtained from the DynamX software program. Maestro 11.8 (Schrödinger Maestro, New York, USA) was used to map the conformational modifications onto the crystal construction of wild-type SOD1 (PDB: 1SPD).
Willpower of Cu and Zn metallic content material
All samples have been despatched to Aspect Supplies Know-how (Santa Fe Springs, California, USA) for quantitation of Cu and Zn metallic content material by way of Inductively Coupled Plasma Mass Spectrometry [76]. Briefly, a pattern portion (0.05 g) was blended with inner requirements (In-Tb-Sc) after which diluted to a last mass of 5 g with an answer of 0.1% ammonium hydroxide, 0.05% EDTA, and 0.05% Triton X100. The pattern appeared to have utterly dissolved. Parts have been analyzed on an Agilent 7500 ICP-MS (Agilent Applied sciences) with an octopole collision cell. The usual working situations used have been RF energy: 1,550 W, Pattern depth: 8 mm, Provider fuel circulation: 1 L/min, Spray chamber temperature: 2°C, Nebulizer Pump: 0.1 rps, Collision fuel: 5.4 mL/min helium. For high quality management, spike restoration experiments have been carried out the place detection limits of Cu and Zn have been measured to be 0.01 ppm and 1 ppm, respectively. Metallic content material for untreated and S-XL6 handled samples are reported in S4 Table.
Proteolytic digestion and MALDI-TOF-MS peptide evaluation
Samples of SOD1A4V with or with out 10-fold molar extra of S-XL6 in 10 mM Tris HCl (pH 7.4) have been incubated for 4 h at 37°C [77,78]. After incubation, samples have been alkylated with iodoacetamide (100 mM for 30 min), heated to 75°C for 20 min, after which handled with 2 volumes of Poroszyme immobilized trypsin (Utilized Biosystems, Life Applied sciences Company, Carlsbad, California, USA) at 37°C for 15 min, mixing each couple of minutes to maintain beads suspended. Beads have been eliminated by centrifugation earlier than evaluation. SOD1H46R was incubated with or with out 10-fold molar extra of S-XL6 and deuterated S-XL6 adopted by pepsin digestion (1:20 w/w, pepsin:protein) for 120 min. Each the digested samples have been analyzed utilizing a microflex MALDI-TOF mass spectrometer (Bruker Daltonics, Billerica, Massachusetts, USA) in reflectron mode within the 2 to five kDa vary and linear mode within the 4 to twenty kDa vary. Spectra have been calibrated utilizing Peptide and Protein I Calibrant (Bruker Daltonics, Billerica, Massachusetts, USA). Matrix solely and trypsin digest response combination with out SOD1 spectra have been acquired as detrimental controls. Spectra have been analyzed in flexAnalysis and BioTools 3.2 (Bruker Daltonics, Billerica, Massachusetts, USA). Peptide mass fingerprinting was carried out utilizing MASCOT (Matrix Science, Boston, Massachusetts, USA) utilizing trypsin because the enzyme with as much as 5 missed cleavages, 100 ppm mass tolerance, and cysteine carbamidomethylation as a variable modification.
Survival evaluation
Two fALS traces have been used to evaluate survival in congenic (later onset at roughly 155 days) and hybrid (earlier onset at roughly 125 days) backgrounds. The primary line was derived by crossing (over >10 generations) congenic fALS “B6 G93A mice” (B6.Cg-Tg(SOD1*G93A)1Gur/J, Inventory No. 004435, The Jackson Laboratory, Bar Harbor, Maine, USA) with “YFP-16 mice” [50] (B6.Cg-Tg(Thy1-YFP)16Jrs/J, Inventory No. 003709, The Jackson Laboratory, Bar Harbor, Maine, USA). This line maintains the C57BL/6J genetic background and expresses YFP in motor and sensory neurons to allow cellular-level molecular profiling [51] in future research. The second line is the usual “Hybrid B6SJL G93A mice” (B6SJL-Tg(SOD1*G93A)1Gur/J Inventory No. 002726, The Jackson Laboratory, Bar Harbor, Maine, USA). Strategies for survival research have been described [79] and adopted the conventions for optimum trial design with fALS mice. Mice have been examined for distinction in: (1) illness onset (at the least a 20% lower in grip power and open-field motor efficiency, which we count on to correlate with change in weight); (2) illness end-stage (illness rating, e.g., lack of righting reflex); and (3) weight. The hybrid line was moreover examined for variations in forelimb and hindlimb grip power.