A analysis workforce from Pohang College of Science and Expertise unveiled the mechanism behind crossover interference throughout meiosis, fixing a long-standing thriller in genetics. This breakthrough may revolutionize agricultural breeding by enabling exact management over crop traits, paving the best way for improved illness resistance and productiveness in vegetation.
Films equivalent to ‘X-Males,’ ‘Incredible 4,’ and ‘The Guardians,’ which showcase vibrant mutant heroes, have captivated world audiences. Not too long ago, a high-throughput genetic screening of meiotic crossover price mutants in Arabidopsis thaliana garnered the curiosity of the tutorial neighborhood by unraveling a century-old thriller within the life sciences.
A analysis workforce, consisting of Professor Kyuha Choi, Dr. Jaeil Kim, and PhD candidate Heejin Kim from the Division of Life Sciences at Pohang University of Science and Technology (POSTECH), has achieved a exceptional feat by unveiling the molecular mechanism answerable for crossover interference throughout meiosis, a organic sample on the chromosome stage. The findings of this analysis have been revealed on February 20 in Nature Vegetation, a world journal within the discipline of life sciences.
The Position of Meiosis in Genetic Range
In sexually reproducing organisms, people resemble their dad and mom or siblings. Regardless of the putting similarities, it’s essential to acknowledge that absolute identicalness is unattainable. This variation is attributed to the method of meiosis, which generates reproductive cells like sperm and eggs in animals or pollen and ovules in vegetation. Not like somatic cell division, which duplicates and divides the genome identically, meiosis creates genetically various reproductive cells by a mechanism often known as crossover.
Meiosis and crossover play pivotal roles in biodiversity and have vital implications in breeding the place the choice and cultivation of superior traits in crops happen. Sometimes, most animal and plant species exhibit a minimal of 1 and a most of three crossovers per a pair of homologous chromosomes.
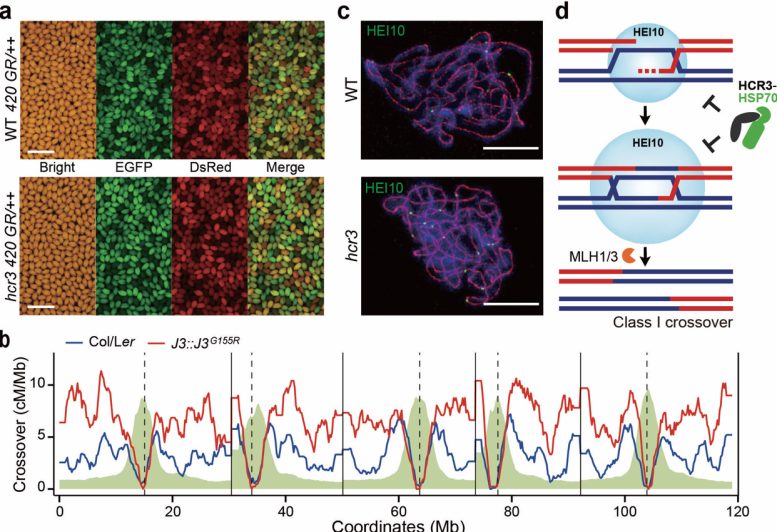
a. Genetic isolation of hcr3 mutants utilizing a fluorescent seed crossover measurement system. b. Genomic crossover maps displaying a 2-fold improve in crossover in J3G155R transgenic vegetation expressing hcr3 allele (highlighted in pink) in comparison with the wild kind (depicted in blue). c. hcr3 confirmed an elevated variety of HEI10 foci and diminished distance between HEI10 foci per bivalent. d. Mannequin illustrating management of HEI10 degradation-mediated crossover interference by the HCR3-HSP70 chaperone community. Credit score: POSTECH
The power to regulate the variety of these crossovers may result in cultivating crops with particular desired traits. Nevertheless, attaining such management has been difficult because of the ‘phenomenon of crossover interference.’ Crossover interference, the place one crossover inhibits the formation of one other crossover close by alongside the identical chromosome, was initially recognized by fruit fly geneticist Hermann J. Muller in 1916. Regardless of researchers’ persistent efforts over the previous century since its discovery, it is just not too long ago that the mechanisms underlying crossover interference have began to unveil their secrets and techniques.
Breakthrough in Understanding Crossover Interference
On this analysis, the workforce utilized a high-throughput fluorescent seed scoring technique to immediately measure crossover frequency in Arabidopsis vegetation. Via a genetic display screen, they recognized a mutant named hcr3 (excessive crossover rate3) that exhibited an elevated crossover price on the genomic stage. Additional evaluation revealed that the elevated crossovers in hcr3 was attributed to some extent mutation within the J3 gene, which encodes a co-chaperone associated to HSP40 protein.
This analysis demonstrated {that a} community involving HCR3/J3/HSP40 co-chaperone and the chaperone HSP70 controls crossover interference and localization by facilitating the degradation of the pro-crossover protein, HEI10 ubiquitin E3 ligase. The applying of genetic display screen approaches to uncover the crossover interference and inhibition pathway efficiently addressed a century-old puzzle within the life sciences.
POSTECH Professor Kyuha Choi said, “Making use of this analysis to agriculture will allow us to quickly accumulate useful traits, thereby decreasing breeding time.” He expressed optimism by saying, “We hope this analysis will contribute to the breeding of latest varieties and identification of helpful pure variations answerable for fascinating traits equivalent to illness and environmental stress resistance, improved productiveness, and high-value manufacturing.”
Reference: “Management of meiotic crossover interference by a proteolytic chaperone community” by Heejin Kim, Jaeil Kim, Namil Son, Pallas Kuo, Chris Morgan, Aurélie Chambon, Dohwan Byun, Jihye Park, Youngkyung Lee, Yeong Mi Park, John A. Fozard, Julie Guérin, Aurélie Hurel, Christophe Lambing, Martin Howard, Ildoo Hwang, Raphael Mercier, Mathilde Grelon, Ian R. Henderson and Kyuha Choi, 20 February 2024, Nature Vegetation.
DOI: 10.1038/s41477-024-01633-y
The analysis was carried out with help from the Primary Analysis Program in Science and Engineering and the Mid-Profession Researcher Program of the Nationwide Analysis Basis of Korea, the Subsequent-Era BioGreen 21 Program of the Rural Improvement Administration, the Suh Kyungbae Basis, and the Samsung Science & Expertise Basis.














