Quotation: Lee L, Rosin LF (2024) Uncharted territories: Fixing the mysteries of male meiosis in flies. PLoS Genet 20(3):
e1011185.
https://doi.org/10.1371/journal.pgen.1011185
Editor: Jean-René Huynh, School de France CNRS, FRANCE
Revealed: March 15, 2024
That is an open entry article, freed from all copyright, and could also be freely reproduced, distributed, transmitted, modified, constructed upon, or in any other case utilized by anybody for any lawful goal. The work is made out there beneath the Creative Commons CC0 public area dedication.
Funding: Work within the Rosin lab is funded by the Intramural Program of the Eunice Kennedy Shriver Nationwide Institute of Youngster Well being and Human Improvement, Nationwide Institutes of Well being (NICHD; HD009019-01 to L.F.R). The funders had no position in examine design, knowledge assortment and evaluation, choice to publish, or preparation of the manuscript.
Competing pursuits: The authors have declared that no competing pursuits exist.
Introduction
Meiosis is the specialised cell division that generates haploid gametes in eukaryotic species. Throughout the first meiotic division, homologous chromosomes (1 maternal and 1 paternal copy of the identical chromosome) should segregate into separate daughter cells. Correct chromosome segregation throughout meiosis I is facilitated by the juxtaposition of homologous companions, the place the two chromosome copies discover one another in 3D area and kind associations to maintain them linked till anaphase I. In most species, these associations kind throughout a chronic prophase I, which consists of chromatin decompaction and linearization to facilitate homolog recognition, alignment, and pairing/synapsis. Right here, we distinguish between homolog alignment, which we outline as the two homologous copies coming into shut bodily proximity within the nucleus and aligning from end-to-end, and pairing, which happens after alignment and entails intimate inter-chromosomal associations which will or might not be depending on the formation of the synaptonemal advanced (SC). In standard meiosis, the SC varieties in a structurally conserved tripartite method, with lateral components alongside every homolog linked to central components by transverse filaments (reviewed in [1,2]). These pairing interactions are stabilized by crossovers (COs) that kind throughout genetic recombination, creating bodily hyperlinks between the homologs that stay even after dissociation of the SC often known as chiasma (singular) or chiasmata (plural). The ensuing bivalent construction is critical for biorientation after nuclear envelope breakdown (NEBD) at metaphase I and correct chromosome segregation at anaphase I (reviewed in [3]).
Nonetheless, male Drosophila segregate their chromosomes in meiosis utilizing a singular mechanism, with out tripartite SC formation [4,5] and with out recombination/chiasmata to hyperlink homologs collectively on the finish of prophase I/metaphase I [6]. As an alternative, bivalents stay linked throughout spermatocyte maturation (Fig 1) through another pathway, referred to as different homolog conjunction (AHC; reviewed in [7]). 4 AHC proteins have been recognized up to now: SNM (stromalin 2), MNM (modifier of mdg4), UNO (univalents solely), and TEF (teflon) [8–10], however how and the place the AHC proteins operate throughout Drosophila male meiosis has remained elusive, and what prevents AHC proteins from linking non-homologous chromosomes collectively is unknown.
Fig 1. Cartoon schematic of germline development in a Drosophila testis.
Prime, from left to proper: somatic hub cells (vibrant inexperienced) are positioned on the most apical tip of the testis. These cells kind a distinct segment for the GSCs (cyan). GSCs endure uneven mitotic divisions, producing a brand new daughter GSC and a GB (darkish inexperienced). GBs endure 4 rounds of mitosis with incomplete cytokinesis, ensuing within the formation of 2-, 4-, 8-, and 16-cell interconnected spermatogonial cysts. The germ cells then endure an prolonged G2/prophase I for cell progress, which might be divided into 7 substages: S1, S2a, S2b, and S3–S6. Throughout this time, homologs are partitioned into separate areas within the nucleus often known as CTs. The formation of CTs might be noticed by solely visualizing DNA. By the S2a stage, nuclei present a tri-lobular morphology, and by S2b, nuclei harbor 3 distinct chromatin lots. These 3 lots are the CTs containing the bivalents of the three main fly chromosomes (2, 3, and X/Y). The small fourth chromosome tends to share a territory with the intercourse chromosomes. By S5, the CTs within the mature spermatocyte nuclei are unfold out, creating an roughly equilateral triangle. Whereas just one cell with CTs is proven, all 16 cells from the cyst progress to this stage and have this chromosome morphology. Backside: The pairing of homologs happens progressively throughout the mitotic divisions. Two consultant homologous centromeres are proven in purple. C(3)G foci are proven in orange. Within the GSCs, solely 20%–30% of homologs are paired, however by the 16-cell cyst stage, greater than 80% of homologs are paired. This pairing happens concurrently with the formation of C(3)G foci, a transverse filament protein concerned in Drosophila SC formation. C(3)G mutants fail to correctly pair homologs by the 16-cell cyst stage, implicating the SC in correct homolog pairing in male Drosophila for the primary time. CT, chromosome territory; GB, gonioblast; GSC, germline stem cell; SC, synaptonemal advanced.
Regardless of the dearth of SC and chiasmata formation in fly spermatogenesis, homologs fully align and are sequestered into separate areas within the nucleus often known as chromosome territories (CTs) [11]. The formation of CTs might be noticed by solely visualizing DNA. By mid-prophase I, nuclei present a tri-lobular morphology and harbor 3 distinct chromatin lots. These 3 lots are the CTs containing the bivalents of the three main fly chromosomes (2, 3, and X/Y; Figs 1 and 2A). The small fourth chromosome tends to share a territory with the intercourse chromosomes, though the mechanism behind that is unclear to at the present time. The CTs within the mature spermatocyte nuclei are unfold out, creating an roughly equilateral triangle. But, how homologs are partitioned into their respective CTs and what prevents heterologous chromosomes from forming CTs collectively is unknown.
Fig 2. Karyotypic adjustments affect CT formation throughout spermatogenesis.
Prime: Karyotype and ensuing CTs throughout fly spermatogenesis. Drosophila wild-type karyotype and CTs are proven in A. (B–E) Present numerous mutant karyotypes and the ensuing CTs. G1 chromosomes (unreplicated chromatids) are proven for simplicity. Centromeres are proven in grey. A genome with 3 main chromosomes results in the formation of three CTs (A and C). Karyotypes with 4 main chromosomes result in the formation of 4 CTs (B and E). When inter-chromosomal translocations happen (for instance, between chromosomes 2 and three; C), this results in the formation of quadrivalents and ends in 2 main CTs. Backside: Karyotype and ensuing CTs shaped in Kc (somatic) cells (F and G). On this context, homology virtually fully dictates CT formation, whatever the underlying karyotypic adjustments. CT, chromosome territory.
Right here, we evaluate current research elucidating the foundations governing CT formation and chromosome pairing throughout Drosophila spermatogenesis.
Pairing initiation in Drosophila male meiosis
How homologs discover one another in Drosophila male meiosis is unclear, as is the case in most species. What has grow to be clear in current many years is that in most species, together with fruit flies, nematodes, budding yeast, and mice, homolog alignment is unbiased of double-strand break (DSB) formation (reviewed in [12]). Nonetheless, in most species, DSBs are required later in prophase I to stabilize pairing interactions. This isn’t the case for male Drosophila, which by no means kind DSBs and by no means kind COs, and but homologs nonetheless affiliate [6]. Intriguingly, current work from Rubin and colleagues investigating chromosome dynamics within the fly male germline means that meiotic pairing in male meiosis is just not merely a continuation of pre-meiotic pairing in flies, as was beforehand believed to be the case. As an alternative, Rubin and colleagues present that homologs are unpaired in germline stem cells within the testes and that pairing of homologs is re-initiated throughout the mitotic division previous meiosis (Fig 1) [13].
How do the homologs discover one another? Research from many species recommend that fast chromosome actions facilitate the preliminary seek for homology and homolog alignment. These actions are proposed to convey homologs nearer in 3D area and likewise to assist resolve entanglements between heterologous chromosomes (reviewed in [14]). Such actions have beforehand been proven to play a job in pairing in Drosophila females, the place nuclear rotations and centromere actions are required to provoke pairing [15]. Of their 2022 examine, Rubin and colleagues show that motion additionally performs a job in Drosophila male meiosis, the place sluggish chromosome actions happen throughout pairing initiation [13]. These actions are pushed by microtubule motors linked to the Drosophila SUN/KASH protein advanced, much like meiotic chromosome actions in different species. Evaluation of pre-meiotic pairing in mutants of the SUN-domain protein, Klaroid, and KASH-domain protein, Klarsicht, additional demonstrated that these elements are required for pairing initiation in Drosophila males. Due to this fact, though pairing in male flies happens throughout the pre-meiotic mitotic divisions, comparable mechanisms appear to use to the seek for homology as in additional canonical meiosis.
Regardless of earlier beliefs that no SC varieties in Drosophila male meiosis for linking homologs collectively [4,5,7], Rubin and staff level out that in response to each FlyAtlas2 and ModENCODE, the Drosophila SC proteins C(3)G and Corona are expressed in testes. Along with validating this expression utilizing RT-qPCR, Rubin and colleagues surprisingly discovered that C(3)G foci kind alongside the chromosome arms by each IF and GFP-tagging throughout these early pairing occasions within the mitotic zone (Fig 1). This foci phenotype is very harking back to insulator “buttons” that facilitate homolog pairing in fly somatic cells [16–18]. Moreover, research utilizing C(3)G and Corona mutants revealed that each these elements are required for the well timed re-initiation of pairing within the mitotic area of the male germline, implicating SC proteins in pairing in Drosophila males for the primary time. Nonetheless, regardless of widespread pairing defects within the mitotic zone of C(3)G and Corona mutants, flies remained fertile, suggesting that this SC-mediated pairing is probably going redundant with different mechanisms regulating meiotic chromosome separation in Drosophila spermatogenesis.
Proof for non-SC-mediated pairing in fly spermatogenesis comes from classical research taking a look at X-Y pairing. Whereas primarily no euchromatic homology exists between the X and Y intercourse chromosomes in flies, a mix of cytological and genetic research by Kenneth Cooper and others as early because the Sixties revealed {that a} area of the shared rDNA repeats connects these chromosomes throughout spermatogenesis [19–21]. This thread-like rDNA connection, often known as the collochore, retains the intercourse chromosomes linked all through prophase I and ensures their correct separation at metaphase I. No proof for such a genetic-based pairing mechanism has ever been validated for the autosomes in flies, and autosomal homolog recognition in flies stays largely elusive. Nonetheless, pericentric heterochromatin and centromeres have been implicated in early pairing occasions in each female and male fly meiosis for all chromosomes [19–27], opening the door to the likelihood that heterochromatin-associated elements might play a job in pairing initiation on this species.
CT formation in Drosophila spermatocytes
Homolog pairing and CT partitioning are intimately linked in fly male meiosis. It’s subsequently unsurprising that the mechanism by which paired homologs are separated into their respective CTs was as elusive as pairing initiation itself. In considered one of their current research, Vernizzi and Lehner got down to decide how homologs, as soon as paired, are sequestered to their separate CTs [28]. Earlier work from Vernizzi and Lehner revealed that, like its position within the soma [29,30] and feminine germline [31], the Condensin II advanced antagonizes inter-chromosomal interactions within the male germline and facilitates the individualization of chromosomal bivalents [32,33]. As soon as individualized, homologous associations are stabilized by the AHC advanced [8–10].
Nonetheless, many questions remained, together with how cells know which chromosomes to place into separate territories. Is every particular person chromosomal entity (made up of linked homologs) partitioned right into a separate CT? Or does chromosome identification issue into this mechanism? To check the mannequin that territory group is ruled by forces maximally separating distinct chromosomal entities, Vernizzi and Lehner used a mix of dwell imaging and classical squash approaches to visualise CTs in testes from flies carrying a wide range of chromosomal aberrations. In wild-type flies, 3 fundamental territories are usually noticed (for chr2, chr3, and chrX/Y/4). Apparently, in flies carrying chr2 compound chromosomes, the place every arm of chr2 varieties a separate chromosome, Vernizzi and Lehner noticed an extra, fourth territory (Fig 2B). Even with the addition of a fourth CT, all CTs had been nonetheless evenly and maximally spaced out within the nucleus. The identical phenomenon was noticed for flies carrying chr3 compound chromosomes. As some heterochromatic repeats are doubtless shared between the arms of chromosomes, the formation of separate CTs by chr2L and chr2R compounds means that homology in heterochromatic domains doesn’t forestall chromosomes from being partitioned into separate territories. Equally, when the authors checked out spermatocytes from flies carrying a posh chr2 fusion which additionally incorporates repetitive components usually discovered on the intercourse chromosomes, C(2)EN, they discovered that spermatocytes harbored 3 equally spaced CTs, much like WT flies (Fig 2C). This remark means that (1) a scarcity of euchromatic homology between chr2L and 2R is just not adequate to drive chromosomes aside throughout CT formation; and (2) the presence of heterochromatic homology between the intercourse chromosomes and C(2)EN is just not adequate to convey chromosomes collectively right into a single CT.
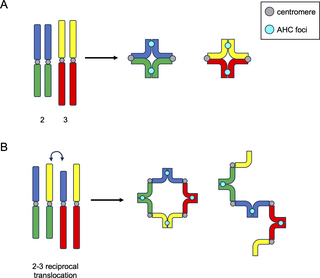
Intriguingly, when CT formation was examined in flies heterozygous for reciprocal translocations between the two massive fly chromosomes (chr2 and chr3), Vernizzi and Lehner noticed solely a single massive CT (Fig 2D). One attainable rationalization for the formation of this single massive CT is that every one linked chromosomal models, no matter identification, are segregated into the identical CT, and all unlinked models kind separate CTs. On this mannequin, such reciprocal translocations would end result within the formation of advanced linkages between chr2 and chr3, forming a quadrivalent (4 linked chromosomes) throughout meiosis (Fig 3), which might result in all chr2 and chr3 segments being partitioned into the identical CT. Certainly, utilizing dwell imaging, Vernizzi and Lehner confirmed that spermatocytes from flies with chr2-chr3 reciprocal translocations harbor both massive ring-, square-, or rod-shaped quadrivalents (Fig 3). Importantly, the ring-shaped quadrivalents noticed in spermatocytes with reciprocal translocations between chr2 and chr3 extremely resemble quadrivalent rings that kind in canonical meiosis with reciprocal translocations, for instance, in mice [34] and crops [35]. The truth that quadrivalents between heterologous chromosomes kind a single CT additional helps the concept that chromosomal identification doesn’t play a job in CT formation within the male germline, and as a substitute, linkages between chromosomes throughout CT formation dictate how chromosomes are partitioned into CTs throughout spermatogenesis in flies.
Fig 3. AHC foci hyperlink homologs in bivalents and quadrivalents in a mechanism akin to chiasmata formation.
Left: Drosophila wild-type (A) or mutant (B) cartoon karyotypes. Centromeres are proven in grey. AHC foci are proven in cyan. (B) Exhibits a big translocation between chromosomes 2 and three. G1 chromosomes are proven for simplicity. In wild-type cells, homologs kind bivalent constructions much like these noticed exterior of Drosophila. Nonetheless, on this occasion, AHC foci hyperlink homologs in lieu of chiasmata. When quadrivalents kind (B) both as rings (center) or rods (proper), AHC foci kind equally spaced alongside the quadrivalent construction, in the identical approach canonical chiasmata would. AHC, different homolog conjunction.
In an much more placing experiment to assist this notion, Vernizzi and Lehner then used CRISPR know-how to chop chr3 in half by focusing on Cas9 to the dodeca pericentric satellite tv for pc repeat on chr3. This separation of the left and proper halves of chr3 led to 4 CTs in spermatocytes (Fig 2E). Most curiously, considered one of these 2 chr3 sub-CTs virtually at all times lacked a centromere (as anticipated for the place of dodeca), and but each sub-CTs had been virtually at all times in a position to retain the maximized spacing within the nucleus. This end result corroborates the mannequin that individualized, unlinked chromosomal entities kind their very own CTs and additional means that centromere-associations with the nuclear envelope usually are not concerned within the spatial separation of CTs.
This paradigm the place individualized entities kind separate CTs no matter chromosomal identification is in distinction to findings of CT formation in fly somatic cells. In 2018, Rosin and colleagues confirmed that chromosome identification virtually fully dictates CT partitioning within the soma [29]. Drosophila cultured cell traces harboring heterologous translocations nonetheless kind regular interphase CTs by Oligopaints FISH. For instance, a translocation between chr2 and chr3, much like C(2)EN, doesn’t disrupt the chr2 or chr3 territories, and each bit of DNA joins its endogenous CT (Fig 2F and 2G). Nonetheless, a majority of these translocations do alter the place of interphase CTs, bringing the translocated chromosomes nearer collectively in interphase [29].
Collectively, these outcomes recommend that chromosomal identification (i.e., being a part of chr2) is just not adequate to group chromosomes collectively into the identical territory and doesn’t forestall chromosomes from shifting aside and forming separate CTs in spermatocytes. As an alternative, these knowledge all assist a mannequin the place individualized chromosomes are all separated into distinct CTs that are then maximally spaced out within the nucleus.
Hyperlinks between homologs and the AHC
In some species, hyperlinks between homologs throughout meiosis are achieved in a crossover-independent method. For instance, females of the silkworm moth Bombyx mori, males of Drosophila, and the nematode C. elegans all kind inter-chromosomal linkages unbiased of crossovers [36–40]. Drosophila males, particularly, make use of a very different mechanism to pair homologs in meiosis referred to as “different homolog conjunction” (AHC). Whereas it has been recommended that AHC meeting happens on the similar time or barely after CT formation [41], which might forestall non-homologous associations, how the 4 AHC proteins (TEF, SNM, MNM, and UNO) work together with one another and chromosomes to hyperlink homologs collectively was not clear.
In 2 current research, Kabakci and colleagues investigated how AHC proteins assemble and bind to DNA [42,43]. These research had a number of key findings. First, TEF binds on to chromosomes and recruits MNM in early meiosis, when AHC is first established. Nonetheless, TEF is now not current in late spermatocytes [9,43], suggesting that TEF doesn’t play a serious position in sustaining homolog linkages later in meiosis. Second, SNM, UNO, and MNM (collectively often known as “SUM”) stay localized to bivalents all through early meiosis, initially forming weak foci alongside bivalents and within the sub-nucleolar area of the nucleus [8–10]. Apparently, these weak SUM foci start to coalesce into 1 single, vibrant focus randomly positioned alongside chromosomal bivalents across the similar time as NEBD and chromosome condensation throughout meiosis I (Fig 3; [28,43]).
This alteration within the quantity and construction of AHC foci is remarkably much like protein diffusion and “coarsening” noticed just lately for pro-CO elements in crops and nematodes [44–48]. Correct homolog segregation in meiosis I in these and plenty of different species is ensured by the formation of COs that hyperlink homologs collectively till anaphase I. CO formation is tightly regulated: there must be no less than 1 CO per chromosome pair, however not quite a lot of COs, and the COs that do kind must be spatially separated alongside the chromosome axis (CO assurance and CO interference, respectively; reviewed in [49,50]). Professional-CO elements initially localize all through the SC, forming a number of small foci termed early recombination nodules (RNs), however later grow to be restricted to a single, or at most a number of, massive foci (late RNs, [51]). Current research in each crops and nematodes have recommended that this variation in RN quantity and construction happens via a phase-separation-based mechanism the place pro-CO elements act as biomolecular condensates that “coarsen” and mixture all through prophase I [45–48]. The adjustments within the construction and variety of RNs that happen because of “coarsening” are extremely much like the adjustments within the construction and variety of AHC foci noticed by Lehner and colleagues, supporting a mannequin the place AHC foci are functionally changing crossovers in fly male meiosis.
Additional assist for this mannequin comes from the ring-shaped quadrivalents noticed in the identical research. Alongside the ring-shaped quadrivalents that shaped in spermatocytes with reciprocal translocations between chr2 and chr3, “mature” AHC foci are evenly spaced inside the quadrivalent, much like CO place in quadrivalents in canonical meiosis because of CO interference (Fig 3; [28]). Intriguingly, such a “coarsening” of homolog-connecting elements is probably going not restricted to RNs and AHC foci. A current examine in feminine Bombyx mori, which additionally make use of an achiasmatic meiosis, demonstrated that SC elements stay between homologs all through prophase I however grow to be dramatically restructured, steadily changing into dense proteinaceous globs between homologs by metaphase I [52,53]. This might additionally presumably happen by a mechanism akin to the coarsening of biomolecular condensates, main us to take a position that “coarsening” could possibly be a common facet of inter-homolog associations.
Lastly, Kabakci and colleagues used a spectrum of computational and biochemical assays to indicate that SUM proteins SNM and UNO are Cohesin-related proteins that facilitate homolog linkages. UNO is an α-kleisin-like protein analogous to Rad21 with a maintained Separase cleavage website [10]. Apparently, the conjunction made by SUM proteins doesn’t appear to contain DNA entrapment, because it does with canonical Cohesin, however slightly direct DNA binding of SUM multimers in a sequence-nonspecific method [42].
In abstract, these outcomes reveal the similarities between AHC-based homolog linkages and chiasmata in canonical meiosis.
Conclusions and abstract
How homologs discover one another in 3D area and pair from end-to-end stays one of many greatest unsolved mysteries in chromosome biology. On this evaluate, we recap current findings concerning the DSB- and CO-independent pairing occasions that happen in Drosophila spermatogenesis. Whereas homolog pairing is sort of ubiquitous in fly somatic cells [54], germline pairing in each females [15] and males [13] is just not merely a continuation of somatic pairing. In males particularly, pairing is re-initiated throughout the pre-meiotic mitotic divisions in germ cells. This pairing entails chromosome actions directed by microtubules, in addition to the SC proteins C(3)G and Corona [13]. As soon as paired, homologs are partitioned into spatially separated CTs. This course of is no less than partially pushed by the motion of Condensin II, which antagonized inter-chromosomal interactions to assist separate bivalents [30–33]. The current research mentioned right here additional reveal that this course of sequesters any linked chromosomal models right into a single CT, no matter chromosome identification [28]. After CT formation, homolog–homolog interactions are stabilized by the AHC pathway [8–10]. These current research show that AHC SUM proteins are Cohesin-related proteins that multimerize, bind on to DNA in a sequence-independent method, and are cleaved by Separase for homolog separation [42,43]. Importantly, SUM proteins clearly hyperlink homologs in a mechanism much like canonical COs. SUM proteins coalesce into distinct foci alongside chromosomes which keep away from pericentromeric areas and kind round one focus per chromosome, and these foci are equidistant on advanced chromosomal constructions like quadrivalents [28]. These new discoveries have elucidated most of the perplexities of the recombination-independent mechanisms occurring in fly spermatogenesis. Additional research will likely be required to validate the similarities between canonical CO formation and the fly AHC pathway and to make clear precisely how AHC proteins are eliminated after metaphase I.