Most cancers is characterised by irregular metabolic processes completely different from these of regular cells. Due to this fact, most cancers metabolism has been extensively studied to develop efficient analysis and remedy methods. Notable achievements of most cancers metabolism research embrace the invention of oncometabolites* and the approval of anticancer medication by the U.S. Meals and Drug Administration (FDA) that focus on enzymes related to oncometabolites. Permitted anticancer medication comparable to ‘Tibsovo (lively ingredient: ivosidenib)’ and ‘Idhifa (lively ingredient: enasidenib)’ are each used for the remedy of acute myeloid leukemia. Regardless of such achievements, learning most cancers metabolism, particularly oncometabolites, stays difficult resulting from time-consuming and costly methodologies comparable to metabolomics. Thus, the variety of confirmed oncometabolites could be very small though a comparatively massive variety of cancer-associated gene mutations have been nicely studied.
Most cancers is characterised by irregular metabolic processes completely different from these of regular cells. Due to this fact, most cancers metabolism has been extensively studied to develop efficient analysis and remedy methods. Notable achievements of most cancers metabolism research embrace the invention of oncometabolites* and the approval of anticancer medication by the U.S. Meals and Drug Administration (FDA) that focus on enzymes related to oncometabolites. Permitted anticancer medication comparable to ‘Tibsovo (lively ingredient: ivosidenib)’ and ‘Idhifa (lively ingredient: enasidenib)’ are each used for the remedy of acute myeloid leukemia. Regardless of such achievements, learning most cancers metabolism, particularly oncometabolites, stays difficult resulting from time-consuming and costly methodologies comparable to metabolomics. Thus, the variety of confirmed oncometabolites could be very small though a comparatively massive variety of cancer-associated gene mutations have been nicely studied.
*Oncometabolite: A metabolite that exhibits pro-oncogenic perform when abnormally amassed in most cancers cells. An oncometabolite is commonly generated on account of gene mutations, and this accumulation promotes the expansion and survival of most cancers cells. Consultant oncometabolites embrace 2-hydroxyglutarate, succinate, and fumarate.
On March 18th, a KAIST analysis crew led by Professor Hyun Uk Kim from the Division of Chemical and Biomolecular Engineering developed a computational workflow that systematically predicts metabolites and metabolic pathways related to somatic mutations in most cancers by means of collaboration with analysis groups beneath Prof Youngil Koh, Prof. Hongseok Yun, and Prof. Chang Wook Jeong from Seoul Nationwide College Hospital.
The analysis groups have efficiently reconstructed patient-specific genome-scale metabolic fashions (GEMs)* for 1,043 most cancers sufferers throughout 24 most cancers sorts by integrating publicly obtainable most cancers sufferers’ transcriptome knowledge (i.e., from worldwide most cancers genome consortiums comparable to PCAWG and TCGA) right into a generic human GEM. The ensuing patient-specific GEMs make it doable to foretell every affected person’s metabolic phenotypes.
*Genome-scale metabolic mannequin (GEM): A computational mannequin that mathematically describes all the biochemical reactions that happen inside a cell. It permits for the prediction of the cell’s metabolic phenotypes beneath varied genetic and/or environmental situations.
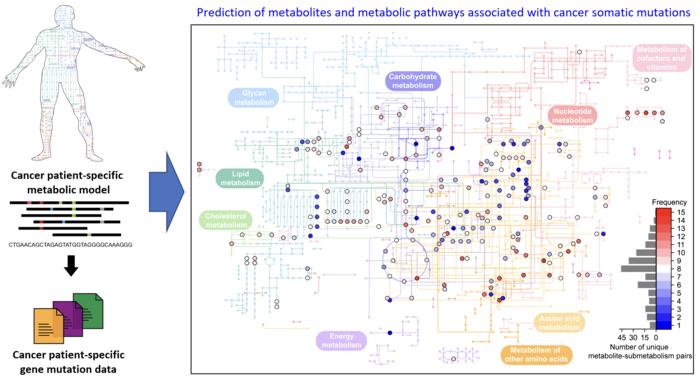
< Determine 1. Schematic diagram of a computational methodology for predicting metabolites and metabolic pathways related to most cancers somatic mutations. of a computational methodology for predicting metabolites and metabolic pathways related to most cancers somatic mutations. >
The crew developed a four-step computational workflow utilizing the patient-specific GEMs from 1,043 most cancers sufferers and somatic mutation knowledge obtained from the corresponding most cancers sufferers. This workflow begins with the calculation of the flux-sum worth of every metabolite by simulating the patient-specific GEMs. The flux-sum worth quantifies the intracellular significance of a metabolite. Subsequent, the workflow identifies metabolites that seem like considerably related to particular gene mutations by means of a statistical evaluation of the anticipated flux-sum knowledge and the mutation knowledge. Lastly, the workflow selects altered metabolic pathways that considerably contribute to the biosynthesis of the anticipated oncometabolite candidates, finally producing metabolite-gene-pathway units as an output.
The 2 co-first authors, Dr. GaRyoung Lee (presently a postdoctoral fellow on the Dana-Farber Most cancers Institute and Harvard Medical College) and Dr. Sang Mi Lee (presently a postdoctoral fellow at Harvard Medical College) stated, “The computational workflow developed can systematically predict how genetic mutations have an effect on mobile metabolism by means of metabolic pathways. Importantly, it may well simply be utilized to various kinds of most cancers based mostly on the mutation and transcriptome knowledge of most cancers affected person cohorts.”
Prof. Kim stated, “The computational workflow and its ensuing prediction outcomes will function the groundwork for figuring out novel oncometabolites and for facilitating the event of varied remedy and analysis methods”.
This research, which was supported by the Nationwide Analysis Basis of Korea, has been revealed on-line in Genome Biology, a consultant journal within the area of biotechnology and genetics, beneath the title “Prediction of metabolites related to somatic mutations in cancers by utilizing genome‑scale metabolic fashions and mutation knowledge”.
DOI
10.1186/s13059-024-03208-8
Methodology of Analysis
Computational simulation/modeling
Topic of Analysis
Cells
Article Title
Prediction of metabolites related to somatic mutations in cancers by utilizing genome‑scale metabolic fashions and mutation knowledge
Article Publication Date
11-Mar-2024