Summary
Just a few brief a long time have handed for the reason that sequencing of GFP, but the trendy repertoire of transgenically encoded optical instruments implies an exponential proliferation of ever enhancing constructions to interrogate the subcellular surroundings. A myriad of tags for labeling proteins, RNA, or DNA have arisen in the previous few a long time, facilitating unprecedented visualization of subcellular elements and processes. Improvement of a broad array of contemporary genetically encoded sensors permits real-time, in vivo detection of molecule ranges, pH, forces, enzyme exercise, and different subcellular and extracellular phenomena in ever increasing contexts. Optogenetic, genetically encoded optically managed manipulation methods have gained traction within the organic analysis neighborhood and facilitate single-cell, real-time modulation of protein operate in vivo in ever broadening, novel purposes. Whereas this discipline continues to explosively develop, references are wanted to help scientists in search of to make use of and enhance these transgenic gadgets in new and thrilling methods to interrogate improvement and illness. On this overview, we endeavor to focus on the state and trajectory of the sector of in vivo transgenic optical instruments.
Quotation: Fenelon KD, Krause J, Koromila T (2024) Opticool: Chopping-edge transgenic optical instruments. PLoS Genet 20(3):
e1011208.
https://doi.org/10.1371/journal.pgen.1011208
Editor: Gregory S. Barsh, HudsonAlpha Institute for Biotechnology, UNITED STATES
Revealed: March 22, 2024
Copyright: © 2024 Fenelon et al. That is an open entry article distributed beneath the phrases of the Creative Commons Attribution License, which allows unrestricted use, distribution, and copy in any medium, supplied the unique writer and supply are credited.
Funding: The research obtained assist from College of Texas at Arlington, UTA STARS, to Dr TK. The funders had no position within the research design, knowledge assortment and evaluation, determination to publish, or preparation of the manuscript.
Competing pursuits: The authors have declared that no competing pursuits exist.
Introduction
Organic inquiries using stay organisms supply superior relevancy to cell tradition and glued tissue research in unraveling the expansive mysteries which stay in cell, developmental, and illness biology as a result of the analysis is being carried out in essentially the most functionally related surroundings. A panoply of instruments has been developed to interrogate these perplexities of nature, however none as thrilling because the rising variety of optical instruments which have lately change into out there to be used in stay organism research.
As such, this overview will concentrate on these optical instruments that may be readily utilized in transgenesis and won’t cowl the instruments for interrogating organic processes by way of fastened samples, nonoptical means, or by utilizing instruments that require exterior biochemical therapies. Within the first occasion, there are a number of out there in depth critiques overlaying advances in chromatin architectural characterization strategies [1,2], fluorescent in situ hybridization (FISH)-related strategies [3,4], and transcriptomics [5,6] and proteomics [7,8]. Equally, enhancements in multiomics [9], mass spectrometry [10,11], DamID and BioID [12–14] have been lately chronicled. Thrilling new in vivo dye, drug, and different exterior therapy strategies, whereas past the scope of this inquiry, are exploding in utility in primary and translational molecular biology fields.
Transgenic fashions recapitulate the pure settings of organic inquiries whereas decreasing the frequency of required optimization steps and autonomously reproducing experimental toolkits. Right here, we endeavor to compile a reference for anybody in search of to grasp or develop new transgenic fashions expressing optical instruments for visualization, quantification, and/or manipulation of subcellular and tissue-level molecular processes.
1 Superior instruments for visualizing subcellular features in vivo
Fluorescent transgenic protein detection tags.
Whereas GFP has solely been used as a transgenic software for lower than 30 years [15], transgenic fluorescent markers for molecular biology have superior to the purpose that mere labeling of proteins with even among the most optimized fluorophores [16,17] is usually considered banal with rising criticisms of those fluorescent proteins’ (FP) limitations (Fig 1A). Maturation charges of FPs have been a very fashionable goal of current optimization research to ameliorate unavoidable delays in protein detection of target-FP fusions [18]. Nonetheless, such limitations at the moment are being averted altogether via an alternate strategy whereby FPs are ubiquitously expressed individually from the goal protein however directed to the goal protein via tag binding in vivo. On this manner, fluorescence might be detected instantly upon translation of the goal protein.
Fig 1. Instruments for visualizing subcellular elements in vivo.
(A) Sequencing of GFP in 1992 allowed for creation of fluorescently labeled, transgenic labels and set in movement the progress of the sector. (B) SunTag:FP fucion proteins bind to the SunTag scaffolding to brightly label POIs. (C) LlamaTag binds on to particular FPs to quickly label POIs. (D) Stem loop binding proteins are used to label RNA sequences. (E) elF4a is a two-part coat protein dimer that binds its particular stem loop construction and permits for cut up FP background elimination. (F) Pumilio permits for labeling of RNA sequencing via a genetically encodable 8 bp-binding area which might be custom-made. It has been improved to work in pairs connected to separate FPs to eradicate background fluorescence and improve sensitivity. (G) dCas9 can be utilized to label DNA sequences in vivo via gRNA focusing on. (H) Zinc fingers might be genetically engineered to focus on DNA sequences in vivo and label them with FPs. Blue arrows signify time in a long time for the reason that sequencing of GFP in 1990, left, to the current, proper. FP, fluorescent protein; POI, protein of curiosity.
These secondary attachment fluorescent methods are typically termed tags [18–21], with one of the crucial fashionable transgenic tag methods being Solar-tag [19,21]. The Solar-tag system makes use of two brief peptides with excessive affinity to 1 one other to recruit FPs to focus on proteins. A fusion protein is constructed consisting of a POI and as much as 24 repeating sequences that permit for complimentary attachment by way of a second FP fusion protein [22,23]. Particularly, a single-chain fragment variable (ScFv) antibody is used which acknowledges GCN4; the GCN4 repeats fused to the POI are used to recruit ScFv-bound FPs to tag the POI for detection (Fig 1B). Solar-tag’s “scaffolding system” produces elevated brightness as a result of a number of fluorescent molecules can concurrently connect to the POI [21]. Llama-tags are one other protein tagging system permitting for sooner visualization in vivo on account of their use of already matured FPs [18]. Llama-tags use optimized nanobody fusion proteins to instantly recruit FPs to POIs (Fig 1C). This mechanism subsequently requires creation of only one transgene for use and permits for transgenic addition of the tags into FP-expressing animals which at the moment are ubiquitous [24,25].
Actual-time transcriptional visualization.
Speedy fluorescence detection by way of pooled mature fluorophores on the prepared has grown in recognition not just for visualizing protein areas, but additionally for detecting gene expression and mRNA dynamics in vivo [19,26,27]. By far the most well-liked methods for real-time transcription detection are the MS2 [28,29] and PP7 [30–33] methods, whereby MS2 coat protein (MCP) and PP7 coat protein (PCP), fused to fluorophores, bind MS2 and PP7 RNA stem loops, respectively. These have been used to generate a broad array of transgenic animals and computational instruments to trace gene expression timing in stay imaging experiments [19,34–39]. Along with these fashionable mRNA visualization choices, a number of different aptamer-coat protein pairs are presently out there for transgenic research together with BglG [40], U1Ap [41], λN22 [42], HTLV-1 Rex [43], TAT/REV [44], and a number of other much less examined variants [40,45,46] (Fig 1D).
The 2-part aptamer-binding eIF4a [47] offers an intriguing twist on the system, offering a dependable, straight-forward methodology to label RNA with out ubiquitous background fluorescence by fusing the two elements to separate fluorophores [48,49] (Fig 1E). Additional, RNA-binding Pumilio [50,51] permits for sequence modification to bind to chosen, engineered eight-nucleotide sequences of RNA and has been applied within the Pumby [52] labeling system to label adjoining mRNA websites to make the most of cut up fluorophores (Fig 1F). The most important limitation of most tag methods being the need for top background fluorescent sign from a pool of overexpressed FPs poised for binding to the tag, these cut up fluorophore approaches current important development by eliminating this extra fluorescent noise. Equally, DNA sequences might be focused transgenically for visualization utilizing dCAS9-FP fusions to focus on sequences in a information RNA directed method [53] (Fig 1G) or by way of conventional engineering of zinc finger-FP fusions [54] (Fig 1H).
2 Transgenic optical sensors
FRET-based sensors.
Past merely labeling mobile elements, a rising array of transgenic optical instruments for measuring subcellular and tissue situations have arisen in current a long time [55,56]. The sphere of transgenic biosensors has, once more, bounded ahead from the earliest strategies, resembling fluorescence restoration after photobleaching (FRAP, Fig 2A), in a mere few a long time [57,58]. These outstanding nano- and micro-scale auto-biological gadgets current a novel alternative within the fields of developmental and illness biology permitting for measurements the place even essentially the most trendy and superior exogenous applied sciences are inadequate [59].
Fig 2. Transgenic optical sensors.
(A) One of many earliest subcellular sensor system, FRAP is used to detect protein diffusion and translation charges in vivo via photobleaching of an FP, adopted by imaging of fluorescent restoration. (B) Fluorophores with overlapping excitation and emission wavelengths can share vitality via FRET when sufficiently shut in proportion to their distance from each other. (C) Separating FRET pairs with molecular springs permits for measurement of extension of the molecular springs, and subsequently, rigidity. (D) Ace Sensors make the most of modified rhodopsin that alters configuration upon formation of membrane potential altering the FRET effectivity of an inserted FP. (E) As a result of flavin-binding FP fluorescence maturation is oxygen unbiased however GFP-based FPs will not be, FluBO reviews O2 ranges by way of O2-dependent FRET effectivity variation. (F) Separating a FRET pair with a caspase recognition sequence permits for detection of caspase exercise by way of SCAT sensors. (G) Steric alteration of the PS3 ε-subunit when certain to ATP permits the ATeam ATP sensor to detect ATP by way of FRET effectivity alteration. (H) Much like FRET, luminescing proteins can generally share vitality. (I) The GAP sensor makes use of shift of GFP excitation wavelength when in proximity to aequorin when certain or unbound to calcium ions. (J) The biochemical domains of some FPs might be remoted, permitting for artificially induced fragility. (Ok) Detection of F-actin polymerization with PriSSM is achieved by wedging a GFP and cpGFP right into a Myosin II motor area such that binding of the motor area to F-actin produces a steric alteration which will increase fluorescence and will increase blue:near-IF excitation ratio. (L) In G-CaMP, a cpGFP is flanked by Calmodulin and a myosin gentle chain fragment which bind each other such that steric situations disrupt fluorescence within the absence of calcium binding. (M) ADP/ATP steric alterations in GlnK fluctuate fused cpGFP optimum excitation wavelength of the Perceval sensor. Inexperienced arrows signify time in a long time for the reason that sequencing of GFP in 1990, left, to the current, proper. FP, fluorescent protein; FRAP, fluorescence restoration after photobleaching; FRET, Förster resonance vitality switch.
Maybe essentially the most versatile of those transgenic devices is proximity detection by way of Förster resonance vitality switch (FRET) [60–62]. On this methodology, 2 proteins are labeled with fluorophores able to FRETing with each other and the space between them might be calculated after they’re in adequate proximity to 1 one other [63] (Fig 2B). Mechanical forces are recognized to be important in a number of developmental and illness pathways [64–71]. A very progressive extension of this methodology is the creation of a number of rigidity sensors [72–77] (Fig 2C). As a result of the space between 2 FRETing fluorophores might be detected optically, fusing suitable fluorophores collectively separated by a molecular spring, typically spectrin [78] or flagelliform [79] repeats, permits for the measurement of forces on the FRET pairs by way of a easy calculation of the measured distance and the recognized spring-like traits of the separator [80–82]. These sensors have been developed in mechanical elements of cells and tissues starting from extracellular matrix and cell–cell connections [83,84], cytoskeletal connections [85,86], and all the way in which into the nucleus and onto the genome itself [75].
An progressive use of FRET in sensors has facilitated the detection of membrane voltage potentials by way of opsin-based Acetabularia opsin (Ace) sensors. Within the Ace sensor methods, an FP is fused to an opsin with which it might probably FRET; when the intracellular aspect of the membrane features a internet optimistic cost, elevated FRET reduces fluorescence of the FP (Fig 2D). Ace sensors now are available in quite a lot of spectral flavors together with pink (VARNAM [87]) and inexperienced (Ace2N-mNeon [88]). One other inventive use of FRET to sense subcellular situations is FluBO [89]. Whereas optical sensing instruments have lengthy been a discipline of inquiry and optimization [90], FluBO cleverly makes use of a flavin-binding FP (FbFP) as donor with a YFP acceptor to detect molecular oxygen (O2) inside cells. YFP is delicate to O2 ranges whereas FbFP will not be, permitting for detection of upper FRET effectivity in O2 wealthy cells (Fig 2E). Caspase exercise can additional be measured utilizing FRET pairs separated by caspase cleavage websites thereby offering a discount in FRET as a readout for caspase exercise [91–97] (see Fig 2F). ATP focus is a crucial subcellular situation which has motivated the creation of a mess of sensors [98]. The transgenic ATeam sensors [99] make the most of modified ATP synthase ε-subunit transversely terminally fused to a donor and acceptor FPs such that binding to ATP [99–101] or MgATP [102] brings the FPs into proximity whereby elevated FRET effectivity might be detected (Fig 2G).
Bioluminescence-based sensors.
A considerably comparable methodology for practical and proximity detection to the FRET-based sensors is the usage of bioluminescent resonant vitality switch (BRET) whereby a bioluminescent molecule is used to shift the emission fluorescence of a fluorophore in adequate proximity by way of molecular transference of the bioluminescent enzyme’s emission to the fluorophore [103,104] (Fig 2H). FRET detection in FRET sensors, whereas exact, has confirmed troublesome to successfully implement in vivo as autofluorescence and comparatively weak signal-to-noise making alternate options like BRET engaging [105,106].
Nonetheless, whereas a panoply of BRET sensors exist [106,107], e.g., Ca2+ sensing with LuCID [108], cAMP sensing with CAMYEL [109,110], cytoskeletal rigidity sensing [111], O2 sensing [112], caspase exercise sensing [113], and POI/POI interplay detection [114], present iterations stay shackled to the requirement of exogenous introduction of their cofactor substrates [115]. As such, these instruments stay distinct from pure transgenic optical instruments. Regardless of this, current advances have been made to attain autoluminescence from bioluminescence methods by introducing genes that in the end catalyze endogenous synthesis of chemical cofactors [116–119]. Conversely, a intelligent employment of Aequorea victorea apo-aequorin permits for Ca2+ sensing with out its bioluminescent cofactor requirement within the GAP (GFP and apo-aequorin) sensors [120]. In these sensors, binding of calcium ions to aequorin shifts the excitation most of GFP permitting for Ca2+ focus quantification by way of ratiometric fluorescence measurement (Fig 2I).
Modified and permutated fluorophore sensors.
A methodological extension of the split-fluorophore mechanism described above for in vivo tagging of RNA with low background is a category of sensors using split-fluorophores [48,49]. In most of those methods, interactions of proteins labeled with complimentary subunits of the cut up fluorophores are detected by fluorescence by way of adequate proximity of the subunits. Whereas this methodology is just like the FRET pair methods above, it has important advantages and downsides compared: split-fluorophores confer a binary interplay measurement and never a distance, however supply greater detection decision with much less background noise and require much less subtle imaging and evaluation strategies than FRET methods. Thus, split-fluorophores have additionally been used to develop binary drive sensors the place the subunits are separated by a versatile linker permitting for subunit separation upon adequate rigidity [76,121].
Fluorophores have additional been modified by rearrangement and insertions of their amino acid sequences to supply varied results, termed circularly permutated fluorophores (cpFPs) [122,123] (Fig 2J). A outstanding software of 1 such cpFP is the pressure sensor PriSSM [124]. Proximity imaging (PRIM) [125]-based pressure sensor module (PriSSM) facilitates detection of F-actin-myosin II pressure by way of ratiometric fluorescence following 490 or 390 nm laser activation. Pressure on PriSSM adjustments the orientations of tandem, contacting GFP and cpGFP FPs making the optimum activation wavelength 390 nm and 490 nm indifferent from F-actin or beneath pressure, respectively, in addition to growing fluorescence beneath pressure (Fig 2K). An extra implementation of a circularly permutated GFP (cpEGFP) was used to assemble a Ca2+ sensor, G-Camp [17,126], which will increase in fluorescence on account of conformational adjustments induced by calcium ion binding (Fig 2L). The lowered effectivity of cpEGFP within the unbound state has the extra, key benefit of accelerating the signal-to-noise ratio of G-CaMP relative to various methods. The Perceval sensor [127–129] permits in vivo measurement of ATP/ADP ratios by way of measurement of fluorescence ratio from a circularly permuted monomeric Venus (cpmVenus) fused to Methanococcus jannaschii GlnK1 when excited by 490 or 405 nm wavelength lasers (Fig 2M). Additional, use of modified FPs to be delicate to different molecules has additionally been achieved as in ClopHensor which makes use of a modified, chloride-ion-sensitive GFP, E2GFP, fused to DsRed-m to detect chloride ranges in cells by ratiometric fluorescence [130,131].
3 Optogenetics: Optical instruments for real-time subcellular manipulations
Activation/Inactivation methods.
Optogenetics is a more moderen method that facilitates handy manipulation of mobile operate [132–136]. With the usage of gentle, proteins might be utilized to have an effect on the operate of the cell. Optogenetics makes use of light-sensitive proteins (both synthetic or naturally occurring) and adjusts their performance via adjusting their secondary, tertiary, or quaternary construction. There’s a myriad of mechanisms that may be affected with optogenetic instruments, however all are initiated by a chromophore or light-absorbing amino acid. Optogenetics is often most well-liked over chemically inducible methods for a mess of causes. Particularly, optogenetic instruments permit for higher specificity via avoidance of secondary chemical results, availability of a number of remoted wavelengths permits for straightforward combinatorial implementations, trendy laser microscopy permits exact 3D localization of impact, and bodily obstacles can typically be overcome via the usage of gentle reasonably than chemical compounds. These instruments can typically be adjusted inside milliseconds permitting for specified mobile manipulation and can be utilized to review a broad vary of cell varieties in stay animals, particularly throughout improvement when tissues depth is minimal and tissue transparency is biggest.
Inactivation of proteins by publicity to particular wavelengths of sunshine is a straight-forward and dependable mechanism to modulate protein operate in vivo. Degrons are naturally occurring peptide markers which facilitate degradation of the protein to which they’re connected [137]. For use as optogenetic instruments, these degradation tags are fused to cryptochromes, enzymes which change conformation in response to gentle to reveal or conceal cryptic domains inside themselves, permitting for concealment and longevity of the goal protein or publicity and degradation of the goal protein in response to gentle.
Whereas optogenetic degrons, as a category, are also known as photo-sensitive degrons, psd is a selected optogenetic degron developed by fusing the photosensitive area of Arabidopsis thaliana phototropin1, gentle oxygen voltage 2 (LOV2), to mouse ornithine decarboxylase carboxy-terminal degron (cODC) [138,139]. When fused to a protein of curiosity (POI), psd publicity to blue gentle induces a conformational change to reveal the cryptic degron, resulting in protease degradation of the assemble which might then be repopulated at nighttime (Fig 3A). One other system developed for triggering protease degradation of fusion proteins by publicity to gentle is the blue gentle inducible degradation (B-LID) methodology [140]. B-LID incorporates a small peptide degron [141] rendered cryptic by fusion to a LOV2 area; on this manner, degradation of fusion proteins is triggered by publicity of the degron upon publicity to blue gentle (see Fig 3A). Photograph-N-degron was lately developed to make the most of light-dependent N-end rule-mediated protein degradation [142]. Mild-induced uncoiling of the Jα helix in LOV2 exposes an N-terminal arginine amino acid and triggers N-end rule degradation [143,144] (see Fig 3A).
Fig 3. Optogenetic instruments for real-time subcellular manipulations.
(A) Degrons are used with LOV2 to yield blue gentle triggering of fusion protein degradation. (B) Photosensitizers are FPs which produce ROS biproducts when excited and can be utilized to distort fused proteins. (C) LANS and LINuS set off reversible nuclear import upon blue gentle publicity by publicity of a cryptic NLS in a LOV2 fusion. (D) LINX and LEXY set off reversible nuclear export upon blue gentle publicity by publicity of a cryptic NES in a LOV2 fusion. (E) iLID/SspB heterodimerize reversibly when uncovered to blue gentle. (F) CRY2/CIB1 reversibly heterodimerize when uncovered to blue gentle. It is very important do not forget that CRY2 additionally oligomerizes beneath blue gentle publicity. (G) Reversible CRY homooligomerization happens when uncovered to blue gentle. (H) COP1/UVR8 reversibly heterodimerizes when uncovered to ultraviolet gentle. At the hours of darkness, UVR8 reversibly kinds homodimers. (I) Optimistic (pMag) and detrimental (nMag) domains inside Magnets instruments dimerize within the presence of blue gentle by way of publicity from cryptic domains inside LOV2 fusions. (J) PixE/PixD heterooligomerizes at nighttime and might be reversibly dissociated into PixD homodimers and PixE monomers by publicity to blue gentle. (Ok) Q-PAS1/BphP1 heterodimers are reversibly shaped when uncovered to far pink gentle, whereas BphP1 homodimers and Q-PAS1 monomers are produced by way of pink gentle or darkness. (L) PhotoCleavable cleaves upon publicity to violet gentle. (M) Opsin membrane transport pumps are used to induce transmembrane pumping of ions when uncovered to applicable wavelengths of sunshine. Orange arrows signify time in a long time for the reason that sequencing of GFP in 1990, left, to the current, proper. FP, fluorescent protein; LANS, light-activated nuclear shuttle; LOV2, gentle oxygen voltage 2; ROS, reactive oxygen species.
Other than degrons, there are different methods for deactivation of proteins. One such methodology is photosensitizers utilized in chromophore-assisted gentle inactivation (CALI) [145,146]. Photosensitizers are chromophores which produce reactive oxygen species (ROS) in response to gentle activation [147]. CALI takes benefit of this manufacturing to inactivate proteins connected to a photosensitizer. The primary genetically encoded photosensitizer was KillerRed, derived from the hydrozoan chromoprotein anm2CP, which produces phototoxic impacts by way of ROS manufacturing in response to inexperienced gentle publicity [148]. KillerRed is an efficient optogenetic software for selectively killing cells and tissues via inexperienced light-mediated ROS manufacturing however can be a great tool in CALI schemes (Fig 3B). As a result of CALI is a extra direct system than degrons, inactivation is considerably extra speedy than degron-based methods however have the disadvantage of manufacturing cytotoxic ROS. Since KillerRed, a number of alternate photosensitizers have been developed to counter KillerRed’s propensity to dimerize and facilitate use of different wavelengths of sunshine [149,150]. LightsOut introduces an AsLOV2 area into the Gal4 transcription issue (TF), broadly used for conditional transgene expression [151,152], between DNA-binding and gene activation domains to suppress Gal4-mediated expression when uncovered to blue gentle [153].
Subcellular translocation methods.
AsLOV2 has additional been applied in light-activated nuclear shuttle (LANS) [154] and light-inducible nuclear localization sign (LINuS) [155] to allow conditional nuclear localization of POIs upon publicity to blue gentle utilizing a cryptic NLS (see Fig 3C). Conversely, the light-inducible nuclear export methods LINX [156,157] and LEXY [158,159] make use of a light-exposable cryptic NES inside an AsLOV2 fusion permitting for blue light-induced nuclear export of POIs (Fig 3D).
The power to reversibly set off Botrytis cinerea BcLOV4 membrane binding by way of blue gentle publicity has enabled its use as a conditional fusion protein localization system whereby blue gentle is used to focus on fusion proteins to the membrane [160–163]. An fascinating, naturally occurring optogenetic software for localization to DNA is Erythrobacter litoralis EL222 which dimerizes and binds to DNA in response to blue gentle [164,165]. This technique is presently utilized in a number of contexts to manage transcription by way of gentle publicity [166–170].
Binding/Polymerization methods.
An enormously helpful optogenetic innovation has been transgenic instruments which facilitate manipulation of binding dynamics of POIs [171]. It’s now doable to make use of these instruments to carry proteins involved with each other, sequester them towards membranes, or launch them by way of publicity to laser gentle (Fig 3E–3K). This mechanism will not be solely of inherent utility in POI practical management but additionally has been proven extensively to be vastly advantageous as a compounding issue along side different optogenetic mechanisms [132].
iLID, an improved and modified variant of LID [141], is a blue gentle inducible dimerizing protein which kinds a dimer with SspB in gentle and dissociates at nighttime [172] (Fig 3E). iLID makes use of an AsLOV2 area fused to an SsrA peptide which is then capable of bind with its binding accomplice, SspB [173], upon publicity by way of reversible, light-induced LOV2 conformational transformation. A deservedly fashionable use of the iLID system is OptoSOS [174–178]. OptoSOS makes use of a membrane-anchored iLID to recruit an SspB-fused SOS to set off a Ras/Erk cascade when uncovered to blue gentle. One other light-triggered dimerization system is cryptochrome 2 (CRY2) and CIB1 [179]. On this system, blue gentle induces heterodimerization of Arabidopsis thaliana CRY2/CIB1 (Fig 3F). Concurrently, nevertheless, this technique additionally causes CRY2 homo-oligomerization creating each a possible experimental problem and a helpful mechanistic enlargement past different hetero/homodimerization methods [180–182] (Fig 3G). Intriguingly, an optogenetic heterodimerization system exists which makes use of UV-B gentle [183]. Within the presence of UV-B, Arabidopsis COP1 binds to UVR8 and can be utilized deliver POIs into proximity to 1 one other in residing cells (Fig 3H).
Equally, heterodimerization might be managed by pink and far-red gentle, as within the phytochrome B (PhyB)/phytochrome interacting issue (PIF) system [184]. Nonetheless, these pink or far-red phytochrome methods require plant chromophore 3-Z phycocyanobilin (PCB) which isn’t naturally synthesized in most animals, making them difficult to make use of in transgenic methods. Nonetheless, PCB might be synthesized in mammalian cells by transgenically introducing 4 genes required to generate it from heme [118,119]. An intriguing use of this technique is SOScat which operates identically to the iLID optoSOS system by way of pink and far-red gentle reasonably than blue, permitting for higher freedom in designing combinatorial transgenic methods [185]. Moreover, Arabidopsis thaliana PhyA/FHY1 behave very equally permitting for the creation of REDMAP which facilitates gene activation beneath pink gentle via heterodimerization and deactivation beneath far-red gentle by way of dissociation [186]. Bacteriophytochrome BphP1 is used as a light-sensitive binding accomplice to Rhodopseudomonas palustris PpsR2 [187]. Extra lately, a truncated variant of PpsR2, Q-PAS1, was engineered to scale back its dimension and mitigate pure oligomerization of PpsR2 [188,189]. On this light-sensitive binding methodology, far-red gentle publicity causes binding of BphP1 to its binding accomplice which is reversed in pink gentle or darkness situations (Fig 3K).
An extra innovation in light-dependent interplay, Magnets, has been developed to induce POIs heterodimerization within the presence of blue gentle [190,191]. Magnets was developed by modifying the Per-Arnt-Sim (PAS) area of the Neurospora crassa LOV protein Vivid which kinds heterodimers when uncovered to blue gentle [192–194]. Modifications have been made to the protein sequence within the binding area to both introduce optimistic (pMag) or detrimental (nMag) amino acids, thereby creating 2 modified Vivid photoreceptors which heterodimerize however don’t homodimerize within the presence of blue gentle (Fig 3I). One other software derived from Vivid is LightOn which makes use of Gal4 fused to Vivid and a p65 transactivation area, GAVP, to induce dimerization and activation of UAS goal gene expression when uncovered to blue gentle [195].
Opposite to the bind-in-the-light/dissociate-in-the-dark scheme of the methods mentioned up to now, there additionally exist methods which make use of the opposite scheme. An instance of that is the Synechocystis PixD/PixE system. At the hours of darkness, PixE and PixD type oligomers containing 5 PixE and 10 PixD proteins which dissociate into PixE monomers and PixD homodimers when uncovered to blue gentle [196] (Fig 3J). Fascinating, bacterial photoactivated adenylyl cyclase (bPAC) [197], which makes use of a BLUF (blue gentle receptor utilizing FAD) gentle sensor area equally to PixD, facilitates light-dependent improve in cAMP ranges. In LOV2 lure and launch of protein (LOVTRAP), a small protein, Zdark (Zdk), was developed to bind to the darkish state of LOV2 [198]. This facilitates the conditional tethering and launch of LOV2 to Zdk beneath darkish or blue gentle publicity, respectively. Considerably orthogonally paralogous to the Zdk/LOV2 binding pair is the Erbin PDZ/LOV2 binding pair, TULIPS [199], whereby a cpePDZ [200] can bind to a brief peptide sequence fused to LOV2 upon blue gentle publicity. A disparate mechanism of light-mediated anchoring management is thru light-controlled protein cleavage. An instance of this technique is PhotoCleavable (PhoCl) [201] (Fig 3L). PhoCl consists of a modified circularly permutated mMaple which spontaneously dissociates upon publicity to violet gentle and disconnection of N- and C- terminal fused POIs.
Opsins.
Maybe the best examples of optogenetic instruments are opsins which have gained immense recognition in neuron research and are lurching in the direction of translational implementation. Opsins are light-sensitive membrane proteins which regulate transmembrane ion switch. This makes them easy, inherent optogenetic instruments whereby membrane potential might be managed by gentle publicity [202] (Fig 3M).
An instance of an opsin optogenetic software is pHoenex, an optical proton pump software [203]. pHoenex makes use of a modified Halorubrum sodomense Archaerhodopsin-3 (Arch3) light-sensitive proton pump fused to an ATPase, vesicular protein Synaptophysin, and optical pH sensor pHluorin [204] to facilitate H+ shuttling to acidify the inside of synaptic vesicles by way of yellow gentle and pH detection by way of blue gentle. Equally, a number of different optogenetic opsin instruments can be found which facilitate light-mediated transmembrane proton pumping [205], sodium ion pumping [206], and chloride ion pumping [207,208]. Moreover, guanylyl and adenylyl cyclase rhodopsins, resembling rhodopsin-guanylyl cyclase (RhGC) [209], rhodopsin cyclic nucleotide phosphodiesterase (Rh-PDE) [210], and guanylyl cyclase rhodopsin (CyclOp) [211], have demonstrated utility in mediating light-dependent cAMP and/or cGMP ranges to exogenously regulate subcellular signaling.
4 Trendy advances in combinatorial, real-time optical strategies
Methods involving easy, direct light-dependent messenger manipulation are fairly fashionable and chic; nevertheless, essentially the most thrilling implementations of transgenic optical instruments are these which contain cooperative implementation of a number of methods concurrently to allow higher management over and comprehension of the molecular organic methods being interrogated. It’s possible that experimental protocols can already be developed to visualise, measure, and manipulate a number of subcellular molecular processes in the identical cell(s) in real- or near-real-time. Certainly, a rising variety of such methods combining a number of transgenic optical instruments and techniques are being developed to mix or enhance upon particular person schemes.
A fancy mix of transgenic optical instruments, the blue light-inducible TEV protease (BLITz) system, combines the CRY2/CIB1 system with the AsLOV2 system to exactly induce transcription upon publicity to blue gentle [212] (Fig 4A). BLITz consists of two elements, a membrane-bound fusion of CIBN (a truncated model of CIB1), the N-terminal portion of the tobacco etch virus protease (TEV), AsLOV2, TEV cleavage web site, and a transcriptional activator, and a soluble portion consisting of CRY2PHR (a truncated model of CRY2) and the C-terminus of TEV (C-TEV). Upon publicity to blue gentle, binding of CRY2PHR to CIBN facilitates interplay of the two elements of TEV whereas concurrently exposing the TEV cleavage web site, permitting cleavage and launch of the transcriptional activator.
Fig 4. Trendy advances in combinatorial, real-time optical strategies.
(A) The BLITZ system makes use of CRY2/CIBN blue light-induced dimerization to attach TEV fragments that are then capable of free a membrane-anchored issue by cleaving a cryptic recognition web site inside a LOV2 area. (B) LANSTRAP and CLASP enhance upon LANS/LiNUS by introducing membrane sequestration of cytoplasmic fusion proteins by way of Zdk/LOV to extra reliably and consistently keep nuclear exclusion within the absence of blue gentle. (C) LINXnano improves upon the LINX/LEXY methods by fusing LINX to a minimal model of SspB, nano, which binds to membrane-tethered iLID within the presence of blue gentle. (D) Amg2 of the BICYCL system reversibly binds BAm Crimson when uncovered to pink gentle and BAm Inexperienced when uncovered to inexperienced gentle, facilitating wavelength-directed transcriptional toggling by recruiting both activators or repressors to genomic regulatory areas. (E) The Blue-OFF system triggers transcriptional silencing and simultaneous degradation of a POI by combining an optogenetic degron fused to the POI with a repressor-bound LOV2 harboring a cryptic DNA-binding protein. (F) iLight makes use of IsPadC red-light-inducible homodimerization and isomeric steric hindrance of fused proteins in darkish or far-red situations to reversibly recruit TFs to gene loci. Crimson arrows signify time in a long time for the reason that sequencing of GFP in 1990, left, to the current, proper. POI, protein of curiosity; TF, transcription issue.
One other software within the trendy genetic engineer’s toolbox is the light-activated reversible inhibition by assembled lure (LARIAT) technique [213]. LARIAT consists of a CIB1-bound CaMKIIα and a CRY2-bound POI. Since CaMKIIα self-oligomerizes, publicity to blue gentle reversibly traps the POI in clusters. The PixE/PixD homo/heterodimer scheme has additionally been used to conditionally induce/dissociate subcellular protein droplets within the PixELLs system [219]. In PixELLs, blue gentle publicity diffuses phase-separated droplets shaped at nighttime via a mix of the liquid–liquid section separation operate of N-terminal intrinsically disordered protein area (IDR) of the human FUS protein (FUSN) [214] and the oligomerization of PixE/PixD beneath darkish situations (see Fig 3J). An analogous mechanism to PixELLs, developed in the identical research, is optoDroplets which makes use of CRY2. In optoDroplets, FUSN is fused to CRY2 to type section separated droplets when blue gentle is utilized to advertise oligomerization of CRY2 [215].
LANSTRAP and CLASP each enhance upon the LANS system for LOV2-mediated nuclear import [216,217] (see Fig 4B). LANSTRAP makes use of a membrane-bound Zdk2 to instantly bind LANS at nighttime, whereas CLASP makes use of a membrane-bound LOV2 to lure a Zdk2:POI:LANS fusion at nighttime, thereby enhancing nuclear exclusion of LANS pre-blue gentle publicity. Alternatively, LINXnano [157] fuses LINX to nano, a truncated model of SspB, which facilitates binding to a membrane-bound iLID in blue gentle (see Fig 4C). This enchancment of the LINX system produces extra full nuclear export by tethering exported LINXnano:POIs throughout the cytoplasm. Much like LANSTRAP and CLASP in operate is iRIS, which sequesters the LOV2:NLS-fused POI within the cytoplasm by way of fusion to Q-PAS1 which facilitates binding to a membrane-anchored BphP1 when uncovered to far-red gentle and nuclear import by NLS publicity when uncovered to blue gentle [218].
Particular Protein Affiliation software giving transcriptional Readout with speedy Kinetics (SPARK) is a software devised to conditionally report cells through which 2 POIs are interacting [219]. One POI is fused to a TEV protease whereas the opposite is fused to a modified LOV2:TEV cleavage web site:TF. Upon publicity to blue gentle, the TEV cleavage web site is uncovered permitting for launch of the TF and activation of reporter gene(s) in cells the place the two POIs are interacting.
Cyanobacteriochrome-based light-inducible dimers (BICYCL) employs a modified light-induced isomerizing GAF (cGMP-specific phosphodies-terases, adenylyl cyclases and FhlA) area derived from Acaryochloris marina AM1_C0023g2, Amg2, to shift binding between 2 binding companions when uncovered to both pink or inexperienced gentle [220]. Binding companions, binder of Amg2-red state (BAmRed) and binder of Amg2-red state (BAmGreen), have been engineered such that Amg2 binds BAmRed when uncovered to pink gentle and to BAmGreen when uncovered to inexperienced gentle, permitting for POI swapping by way of gentle publicity (Fig 4D). The builders of BICYCL confirmed that it may be used to conditionally recruit both a transcriptional repressor or activator to DNA-tethered Amg2 relying on laser coloration selection.
Blue-OFF combines LOV2-mediated transcriptional inhibition with psd-mediated POI degradation to extra utterly eradicate POI presence when uncovered to blue gentle [221] (Fig 4E). LOV2 is used to create a cryptic DNA-binding area and is additional fused to a transcriptional inhibitor (KRAB) to eradicate transcription of the POI which is fused to B-LID for degron-mediated depletion. Analogously, iLight controls gene transcription by way of far pink mediated homodimerization of the photosensory core module of Idiomarina IsPadC (IsPadC-PMC) fused to both LexA408 to dam transcription or Gal4 and VP16 to set off gene expression, which might be reversed by way of publicity to near-infrared gentle [222] (Fig 4F). At the hours of darkness or after publicity to near-infrared gentle, the fused TFs are sterically inhibited by isomeric transformations of IsPadC-PMC.
Dialogue
Whereas there have been many critiques on optical tagging methods, optical biosensors, optogenetics, and biologically expressed imaging instruments extra broadly [223], we’ve targeted this compendium particularly on instruments presently out there to be used in transgenic plant and animal research. In just a few a long time, molecular biology and genetic engineering fields have exploded from advanced transgenics being a hypothetical potentiality to hundreds of transgenic organisms, lots of of fluorescent proteins and tags, scores of subcellular sensors, and dozens of purposes for a number of optogenetic methods.
It is very important rigorously contemplate the restrictions of every class of optical software in addition to for every particular person instrument therein. For instance, labeling with one of many tag methods described right here comes with the drawback of occupying a laser and detector on the microscope merely to label, so it’s generally prudent to make use of a sensor additionally as a label which can include the disadvantages of sluggish fluorescent maturation and low sign. Equally, inside courses of optical instruments, totally different instruments with comparable features have vital benefits and limitations to think about. For instance, degrons require transcription and translation of substitute fusion POIs to recuperate after depletion whereas translocation instruments require mere shuttling throughout the cell to recuperate; nevertheless, degrons extra shortly and utterly deplete POI ranges. Additional, it’s essential to ponder the activation/inactivation occasions of optogenetic instruments which regularly take a pair to tens of minutes to achieve peak activation in addition to their efficiencies of exercise and reversibility which are likely to land everywhere in the map. As well as, when contemplating the usage of sensors or tags together with optogenetic instruments, it’s crucial to weigh the price in photobleaching and phototoxicity towards the reward of maintained optogenetic activation and tailor experiments accordingly. As mentioned within the earlier part, combinatorial purposes of those instruments are being developed which synergistically complement every particular person part’s limitations.
It’s anticipated that we are going to quickly see a surge in improvement of subtle instruments and research monitoring chromatin structure, transcription, translation, protein localization, and mobile responses by way of reporters, tags, and sensors whereas concurrently manipulating transcription or protein ranges or operate in residing transgenic vegetation and animals. One such research has already reached preprint from the lab of Hernan Garcia manipulating transcription components by way of LEXY whereas concurrently monitoring transcription of their targets by way of MS2/MCP [224]. One other ongoing research from the lab of Sanjeevi Sivasankar seeks to mix optogenetics and BioID by fusing CRY2 or CIB1 to 2 halves of a split-TurboID biotinylation enzyme thereby permitting for exact temporal management of the enzyme’s exercise [225].
Moreover, we’ve conceived of some hypothetical developments which we consider might be helpful if developed. First, exact spatiotemporal management of epigenetic states has the potential to be a useful software for interrogating improvement and illness and might feasibly be achieved by AsLOV2-toggling of histone modifiers and genomic focusing on by way of gRNAs and dCAS9 (see Fig 5A). Second, combining tags with different methods is bound to quickly acquire recognition; utilizing SUN-tag to sequester transcription components conditioned upon pre-cellularization situations within the early Drosophila embryo, e.g., could possibly be helpful for particular enquiries into early improvement (see Fig 5B). Third, the BLITZ system can feasibly be modified to manage overexpressed transgenic transcription components which might in any other case accumulate in methods like LINXnano and confound expectations of spatiotemporal precision (see Fig 5C).
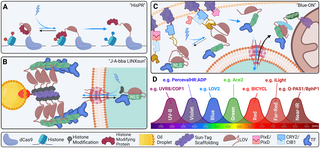
Fig 5. Future instructions of transgenic optical instruments.
(A) Histone modification by CRISPR (“HisPR”): LOV2 can hypothetically be used to conditionally expose a cryptic histone modifying enzyme focused to genomic loci by dCAS9. (B) Jabba-trapped LINX by way of SunTag (“J-A-bba LINXsun”): Jabba-trap was developed to lure fusion proteins on lipid droplets dispersed all through the pre-cellularization Drosophila embryo [226]. Along with LINX and SunTag, it could be doable to exactly set off nuclear export of TFs by way of blue gentle till gastrulation begins. (C) “Blue-ON”: Blue gentle releases a membrane-caged caspase fragment by way of PixE/PixD motion. Caspase fragments assemble in comparable style to BLITZ beneath blue gentle publicity by way of iLID/SspB motion to launch an NLS-fused nuclear issue by way of caspase recognition sequence publicity by LOV2. LANS:TF fusion is uptaken by the nucleus whereas uncovered to blue gentle as in LANS. At the hours of darkness, the LANS assemble is exported from the nucleus by way of NES, sequesters at mitochondrial-bound Zdk, and is degraded by way of fused degron sequence. Whereas fairly advanced, this technique would produce close to full silencing of overexpression fashions at nighttime; additional, as soon as a transgenic is produced in a single species, solely the TF wants get replaced to supply instruments for some other goal. (D) At the moment, it could be doable to make use of as much as 7 transgenic optical instruments concurrently. LANS, light-activated nuclear shuttle; TF, transcription issue.
Unhazardous blue and pink gentle are actually most helpful for in vivo research in transgenic animals, advanced methods encorporating sorts of optogenetic instruments would require a number of lasers for use in live performance. It’s thrilling that a number of methods have already been developed which implement bohemian wavelengths for activation, together with far-red, near-infrared, violet, and UV-B (see Fig 5D). With applicable experimental design and controls, it’s hypothetically doable to concurrently use a handful of various lasers to have an effect on totally different optical instruments and obtain more and more advanced targets.
We’ll undoubtedly see additional exponential progress of the sector of transgenic optical software improvement and software within the coming years. Neurological and metabolic issues stand to learn vastly from future advances which is able to undoubtedly embrace translational purposes with medical potential. Encouragingly, advances in ex utero [227–229] and in utero [230,231] experimental strategies and applied sciences promise to deliver optogenetics to mammalian embryonic improvement, an vital step towards this aim. Additional, the genetically encoded gadgets described herein supply our greatest likelihood at supplementing our inadequate understanding of primary cell, developmental, and illness mechanisms which have hitherto remained inaccessible. This overview is supposed to be a quick almanac of the instruments out there to those that will develop and use these future fashions and gadgets.
Acknowledgments
We want to categorical our gratitude to Dr. Eric Wieschaus of Princeton College and Clarissa Pasciliao of the College of Toronto for his or her invaluable recommendation, in addition to all of the members of the Koromila lab for his or her constructive suggestions through the preparation of this manuscript. All figures have been created in BioRender and Adobe Illustrator.
References
- 1.
Bouwman BAM, Crosetto N, Bienko M. The period of 3D and spatial genomics. Tendencies Genet. 2022 Oct 1;38(10):1062–75. pmid:35680466 - 2.
Soroczynski J, Risca VI. Technological advances in probing 4D genome group. Curr Opin Cell Biol. 2023 Oct 1;84:102211. pmid:37556867 - 3.
Van Gijtenbeek LA, Kok J. Illuminating Messengers: An Replace and Outlook on RNA Visualization in Micro organism. Entrance Microbiol. 2017;8:1161. pmid:28690601 - 4.
Asp M, Bergenstråhle J, Lundeberg J. Spatially Resolved Transcriptomes—Subsequent Technology Instruments for Tissue Exploration. Bioessays. 2020;42(10):1–16. pmid:32363691 - 5.
Ding J, Sharon N, Bar-Joseph Z. Temporal modelling utilizing single-cell transcriptomics [Internet]. Vol. 23, Nature Evaluations Genetics. Nature Publishing Group; 2022 [cited 2023 Sep 7]. p. 355–68. Accessible from: https://www.nature.com/articles/s41576-021-00444-7. pmid:35102309 - 6.
Tian L, Chen F, Macosko EZ. The increasing vistas of spatial transcriptomics [Internet]. Vol. 41, Nature Biotechnology. Nature Publishing Group; 2023 [cited 2023 Sep 7]. p. 773–82. Accessible from: https://www.nature.com/articles/s41587-022-01448-2. pmid:36192637 - 7.
Cui M, Cheng C, Zhang L. Excessive-throughput proteomics: a methodological mini-review [Internet]. Vol. 102, Laboratory Investigation. Nature Publishing Group; 2022 [cited 2023 Sep 7]. p. 1170–81. Accessible from: https://www.nature.com/articles/s41374-022-00830-7. pmid:35922478 - 8.
Bennett HM, Stephenson W, Rose CM, Darmanis S. Single-cell proteomics enabled by next-generation sequencing or mass spectrometry [Internet]. Vol. 20, Nature Strategies. Nature Publishing Group; 2023 [cited 2023 Sep 7]. p. 363–74. Accessible frem: https://www.nature.com/articles/s41592-023-01791-5. - 9.
Vandereyken Ok, Sifrim A, Thienpont B, Voet T. Strategies and purposes for single-cell and spatial multi-omics [Internet]. Vol. 24, Nature Evaluations Genetics. Nature Publishing Group; 2023 [cited 2023 Sep 7]. p. 494–515. Accessible from: https://www.nature.com/articles/s41576-023-00580-2. pmid:36864178 - 10.
Brodbelt JS. Deciphering combinatorial post-translational modifications by top-down mass spectrometry [Internet]. Vol. 70, Present Opinion in Chemical Biology. Annual Evaluations; 2022 [cited 2023 Sep 7]. p. 157–79. Accessible from: https://www.annualreviews.org/doi/abs/10.1146/annurev-biophys-092721-085421. pmid:35779351 - 11.
Karch KR, Snyder DT, Harvey SR, Wysocki VH. Native Mass Spectrometry: Current Progress and Remaining Challenges [Internet]. Vol. 51, Annual Assessment of Biophysics Annual Evaluations; 2022 [cited 2023 Sep 7]. p. 157–79. Accessible from: https://www.annualreviews.org/doi/abs/10.1146/annurev-biophys-092721-085421. pmid:34982572 - 12.
Kang MG, Rhee HW. Molecular Spatiomics by Proximity Labeling. Acc Chem Res [Internet]. 2022 [cited 2023 Sep 7];2022:37. Accessible from: https://pubs.acs.org/doi/full/10.1021/acs.accounts.2c00061. pmid:35512328 - 13.
Suzuki Y, Kadomatsu Ok, Sakamoto Ok. In the direction of the in vivo identification of protein-protein interactions. J Biochem [Internet]. 2023 Might 29 [cited 2023 Sep 7];173(6):413–5. pmid:36821413 - 14.
Shkel O, Kharkivska Y, Kim YK, Lee JS. Proximity Labeling Methods: A Multi-Omics Toolbox. Chem Asian J. 2022 Jan 17;17(2). pmid:34850572 - 15.
Prasher DC, Eckenrode VK, Ward WW, Prendergast FG, Cormier MJ. Main construction of the Aequorea victoria green-fluorescent protein. Gene. 1992 Feb 15;111(2):229–33. pmid:1347277 - 16.
Riani YD, Matsuda T, Takemoto Ok, Nagai T. Inexperienced monomeric photosensitizing fluorescent protein for photo-inducible protein inactivation and cell ablation. BMC Biol [Internet]. 2018 Apr 30 [cited 2023 Aug 16];16(1):1–12. Accessible from: https://link.springer.com/articles/10.1186/s12915-018-0514-7. - 17.
Nakai J, Ohkura M, Imoto Ok. A excessive signal-to-noise Ca2+ probe composed of a single inexperienced fluorescent protein. Nat Biotechnol [Internet]. 2001;19:137–141. Accessible from: http://biotech.nature.com. pmid:11175727 - 18.
Bothma JP, Norstad MR, Alamos S, Garcia HG. LlamaTags: A Versatile Instrument to Picture Transcription Issue Dynamics in Dwell Embryos. Cell [Internet]. 2018 [cited 2023 Jul 18];173(7):1810–1822.e16. pmid:29754814 - 19.
Toran P, Smolina I, Driscoll H, Ding F, Solar Y, Cantor CR, et al. Labeling native bacterial RNA in stay cells. Cell Res. 2014;24(7):894–897. pmid:24732010 - 20.
Marques SM, Slanska M, Chmelova Ok, Chaloupkova R, Marek M, Clark S, et al. Mechanism-Based mostly Technique for Optimizing HaloTag Protein Labeling. JACS Au [Internet]. 2022 [cited 2023 Jul 19];2(6):1324–37. pmid:35783171 - 21.
Tanenbaum ME, Gilbert LA, Qi LS, Weissman JS, Vale RD. A protein tagging system for sign amplification in gene expression and fluorescence imaging. Cell [Internet]. 2014 Oct 10 [cited 2023 Aug 10];159(3):635. pmid:25307933 - 22.
Harmansa S, Affolter M. Protein binders and their purposes in developmental biology. Improvement (Cambridge) [Internet]. 2018 Jan 15 [cited 2023 Sep 6];145(2). pmid:29374062 - 23.
Takemoto Ok, Matsuda T, Sakai N, Fu D, Noda M, Uchiyama S, et al. SuperNova, a monomeric photosensitizing fluorescent protein for chromophore-assisted gentle inactivation. Sci Rep [Internet]. 2013 Sep 17 [cited 2023 Sep 6];3(1):1–7. Accessible from: https://www.nature.com/articles/srep02629. pmid:24043132 - 24.
De Meyer T, Muyldermans S, Depicker A. Nanobody-based merchandise as analysis and diagnostic instruments. Tendencies Biotechnol [Internet]. 2014 Might 1 [cited 2023 Oct 2];32(5):263–70. Accessible from: http://www.cell.com/article/S0167779914000419/fulltext. pmid:24698358 - 25.
Xu J, Kim AR, Cheloha RW, Fischer FA, Li JSS, Feng Y, et al. Protein visualization and manipulation in Drosophila via the usage of epitope tags acknowledged by nanobodies. Elife. 2022 Jan 1;11. pmid:35076390 - 26.
Yiu HW, Demidov VV, Toran P, Cantor CR, Broude NE. RNA detection in stay bacterial cells utilizing fluorescent protein complementation triggered by interplay of two RNA aptamers with two RNA-binding peptides. Prescription drugs. 2011;4(3):494–508. - 27.
van Gijtenbeek LA, Kok J. Illuminating messengers: An replace and outlook on RNA visualization in micro organism. Entrance Microbiol [Internet]. 2017 [cited 2023 Jul 10];8(JUN):1–19. Accessible from: www.frontiersin.org. pmid:28690601 - 28.
Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH, Lengthy RM. Localization of ASH1 mRNA particles in residing yeast. Mol Cell. 1998;2(4):437–445. pmid:9809065 - 29.
Tutucci E, Vera M, Biswas J, Garcia J, Parker R, Singer RH. An improved ms2 system for correct reporting of the mrnA life cycle. 2018;15(1). - 30.
Lim F, Peabody DS. RNA recognition web site of PP7 coat protein. Nucleic Acids Res [Internet]. 2002 Oct 10 [cited 2023 Sep 6];30(19):4138. pmid:12364592 - 31.
Heinrich S, Sidler CL, Azzalin CM, Weis Ok. Stem-loop RNA labeling can have an effect on nuclear and cytoplasmic mRNA processing. RNA [Internet]. 2017 Feb 1 [cited 2023 Sep 6];23(2):134–41. pmid:28096443 - 32.
Fukaya T, Lim B, Levine M. Speedy Charges of Pol II Elongation within the Drosophila Embryo. Curr Biol. 2017 Might 8;27(9):1387–91. pmid:28457866 - 33.
Levo M, Raimundo J, Bing XY, Sisco Z, Batut PJ, Ryabichko S, et al. Transcriptional coupling of distant regulatory genes in residing embryos. Nature [Internet]. 2022 Might 4 [cited 2023 Aug 14];605(7911):754–60. Accessible from: https://www.nature.com/articles/s41586-022-04680-7. pmid:35508662 - 34.
Viushkov VS, Lomov NA, Rubtsov MA, Vassetzky YS. Visualizing the Genome: Experimental Approaches for Dwell-Cell Chromatin Imaging [Internet]. Vol. 11, Cells. Multidisciplinary Digital Publishing Institute; 2022 [cited 2023 Aug 14]. p. 4086. Accessible from: https://www.mdpi.com/2073-4409/11/24/4086/htm. pmid:36552850 - 35.
Vinter DJ, Hoppe C, Ashe HL. Dwell and glued imaging of translation websites at single mRNA decision within the Drosophila embryo. STAR Protoc [Internet]. 2021 [cited 2023 Jul 18];2(3):100812. pmid:34585149 - 36.
Koromila T, Stathopoulos A. Distinct Roles of Broadly Expressed Repressors Assist Dynamic Enhancer Motion and Change in Time. Cell Rep [Internet]. 2019;28(4):855–863.e5. pmid:31340149 - 37.
Koromila T, Gao F, Iwasaki Y, He P, Pachter L, Peter Gergen J, et al. Odd-paired is a pioneer-like issue that coordinates with zelda to manage gene expression in embryos. Elife. 2020 Jul 1;9:1–71. pmid:32701060 - 38.
Ohishi H, Shimada S, Uchino S, Li J, Sato Y, Shintani M, et al. STREAMING-tag system reveals spatiotemporal relationships between transcriptional regulatory components and transcriptional exercise. Nat Commun [Internet]. 2022 Dec 20 [cited 2023 Oct 10];13(1):1–19. Accessible from: https://www.nature.com/articles/s41467-022-35286-2. - 39.
Birnie A, Plat A, Korkmaz C, Bothma JP. Exactly timed regulation of enhancer exercise defines the binary expression sample of Fushi tarazu within the Drosophila embryo. Curr Biol. 2023 Jul 24;33(14):2839–2850.e7. pmid:37116484 - 40.
Chen J, Nikolaitchik O, Singh J, Wright A, Bencsics CE, Coffin JM, et al. Excessive effectivity of HIV-1 genomic RNA packaging and heterozygote formation revealed by single virion evaluation. Proc Natl Acad Sci U S A. 2009 Aug 11;106(32):13535–40. pmid:19628694 - 41.
Takizawa PA, Vale RD. The myosin motor, Myo4p, binds Ash1 mRNA by way of the adapter protein, She3p. Proc Natl Acad Sci U S A [Internet]. 2000 Might 9 [cited 2023 Sep 6];97(10):5273–8. pmid:10792032 - 42.
Schönberger J, Hammes UZ, Dresselhaus T. In vivo visualization of RNA in vegetation cells utilizing the λN22 system and a GATEWAY-compatible vector sequence for candidate RNAs. Plant J [Internet]. 2012 Jul 1 [cited 2023 Sep 6];71(1):173–81. pmid:22268772 - 43.
Ye J, Silverman L, Lairmore MD, Inexperienced PL. HTLV-1 Rex is required for viral unfold and persistence in vivo however is dispensable for mobile immortalization in vitro. Blood [Internet]. 2003 Dec 12 [cited 2023 Sep 6];102(12):3963. pmid:12907436 - 44.
Das AT, Harwig A, Berkhout B. The HIV-1 Tat Protein Has a Versatile Function in Activating Viral Transcription. J Virol [Internet]. 2011 Sep 15 [cited 2023 Sep 6];85(18):9506. pmid:21752913 - 45.
Rentmeister A, Mannack LVJC, Eising S. Present strategies for visualizing RNA in cells. F1000Res [Internet]. 2016 [cited 2023 Sep 6];5. - 46.
Ozawa T, Natori Y, Sato M, Umezawa Y. Imaging dynamics of endogenous mitochondrial RNA in single residing cells. Nat Strategies. 2007;4(5):413–419. pmid:17401370 - 47.
Suzuki C, Garces RG, Edmonds KA, Hiller S, Hyberts SG, Wagner AM. PDCD4 inhibits translation initiation by binding to eIF4A utilizing each its MA3 domains. Proc Natl Acad Sci U S A [Internet]. 2008 Mar 4 [cited 2023 Sep 6];105(9):3274–9. pmid:18296639 - 48.
Feng S, Sekine S, Pessino V, Li H, Leonetti MD, Huang B. Improved cut up fluorescent proteins for endogenous protein labeling. Nat Commun [Internet]. 2017 Aug 29 [cited 2023 Sep 7];8(1):1–11. Accessible from: https://www.nature.com/articles/s41467-017-00494-8. - 49.
Furman JL, Badran AH, Shen S, Stains CI, Hannallah J, Segal DJ, et al. Systematic analysis of split-fluorescent proteins for the direct detection of native and methylated DNA. Bioorg Med Chem Lett. 2009 Jul 15;19(14):3748–51. pmid:19457665 - 50.
Wang X, McLachlan J, Zamore PD, Corridor TMT. Modular recognition of RNA by a human Pumilio-homology area. Cell [Internet]. 2002 Aug 23 [cited 2023 Aug 14];110(4):501–12. Accessible from: http://www.cell.com/article/S0092867402008735/fulltext. pmid:12202039 - 51.
Tilsner J, Linnik O, Christensen NM, Bell Ok, Roberts IM, Lacomme C, et al. Dwell-cell imaging of viral RNA genomes utilizing a Pumilio-based reporter. Plant J [Internet]. 2009 Feb 1 [cited 2023 Aug 14];57(4):758–70. pmid:18980643 - 52.
Adamala KP, Martin-Alarcon DA, Boydena ES. Programmable RNA-binding protein composed of repeats of a single modular unit. Proc Natl Acad Sci U S A [Internet]. 2016 Might 10 [cited 2023 Sep 7];113(19):E2579–88. pmid:27118836 - 53.
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, et al. Dynamic imaging of genomic loci in residing human cells by an optimized CRISPR/Cas system. Cell [Internet]. 2013 Dec 19 [cited 2023 Oct 10];155(7):1479–91. Accessible from: http://www.cell.com/article/S0092867413015316/fulltext. pmid:24360272 - 54.
Lindhout BI, Fransz P, Tessadori F, Meckel T, Hooykaas PJJ, van der Zaal BJ. Dwell cell imaging of repetitive DNA sequences by way of GFP-tagged polydactyl zinc finger proteins. Nucleic Acids Res [Internet]. 2007 Aug 15 [cited 2023 Oct 10];35(16):e107–e107. pmid:17704126 - 55.
Borisov SM, Wolfbeis OS. Optical biosensors. Chem Rev. 2008 Feb;108(2):423–61. pmid:18229952 - 56.
Wang H, Jing M, Li Y. Lighting up the mind: genetically encoded fluorescent sensors for imaging neurotransmitters and neuromodulators. Curr Opin Neurobiol. 2018 Jun 1;50:171–8. pmid:29627516 - 57.
Cole NB, Smith CL, Sciaky N, Terasaki M, Edidin M, Lippincott-Schwartz J. Diffusional Mobility of Golgi Proteins in Membranes of Dwelling Cells. Science (1979). 1996;273(5276):797–801. pmid:8670420 - 58.
Zhang Y, Avalos JL. Conventional and novel instruments to probe the mitochondrial metabolism in well being and illness. Wiley Interdiscip Rev Syst Biol Med. 2017;9(2):1373. pmid:28067471 - 59.
Wang M, Da Y, Tian Y. Fluorescent proteins and genetically encoded biosensors. Chem Soc Rev. 2023 Feb 20;52(4):1189–214. pmid:36722390 - 60.
Hochreiter B, Garcia AP, Schmid JA. Fluorescent proteins as genetically encoded FRET biosensors in life sciences. Sensors (Switzerland). 2015;15(10):26281–26314. pmid:26501285 - 61.
Wu L, Huang C, Emery BP, Sedgwick AC, Bull SD, He XP, et al. Förster resonance vitality switch (FRET)-based small-molecule sensors and imaging brokers. Chem Soc Rev. 2020;49:5110–5139. - 62.
Llères D, James J, Swift S, Norman DG, Lamond AI. Quantitative evaluation of chromatin compaction in residing cells utilizing FLIM-FRET. J Cell Biol. 2009;187(4):481–496. pmid:19948497 - 63.
Day RN. Visualization of Pit-1 Transcription Issue Interactions within the Dwelling Cell Nucleus by Fluorescence Resonance Power Switch Microscopy. Mol Endocrinol. 1998 Sep 1;12(9):1410–9. pmid:9731708 - 64.
Dupont S, Wickström SA. Mechanical regulation of chromatin and transcription. Vol. 23, Nature Evaluations Genetics. Nature Publishing Group; 2022. p. 624–43. pmid:35606569 - 65.
Fenelon KD, Hopyan S. Structural elements of nuclear integrity with gene regulatory potential. Curr Opin Cell Biol. 2017;48:63–71. pmid:28641117 - 66.
Wang C, Yang J. Mechanical forces: The lacking hyperlink between idiopathic pulmonary fibrosis and lung most cancers. Eur J Cell Biol. 2022 Jun 1;101(3):151234. pmid:35569385 - 67.
Vignes H, Vagena-Pantoula C, Prakash M, Fukui H, Norden C, Mochizuki N, et al. Extracellular mechanical forces drive endocardial cell quantity lower throughout zebrafish cardiac valve morphogenesis. Dev Cell. 2022;57(5):598–609.e5. pmid:35245444 - 68.
Zuela-Sopilniak N, Lammerding J. Can’t deal with the stress? Mechanobiology and illness. Tendencies Mol Med. 2022 Sep 1;28(9):710–25. pmid:35717527 - 69.
Sanfeliu-Cerdán N, Lin LC, Dunn AR, Goodman MB, Krieg M. Visualizing Neurons Below Stress In Vivo with Optogenetic Molecular Pressure Sensors. Strategies Mol Biol. 2023:239–266. pmid:36587102 - 70.
Maurer M, Lammerding J. The Driving Pressure: Nuclear Mechanotransduction in Mobile Operate, Destiny, and Illness. Annu Rev Biomed Eng. 2019;21:443–468. pmid:30916994 - 71.
Alisafaei F, Jokhun DS, Shivashankar GV, Shenoy VB. Regulation of nuclear structure, mechanics, and nucleocytoplasmic shuttling of epigenetic components by cell geometric constraints. Proc Natl Acad Sci U S A. 2019;116(27):13200–13209. pmid:31209017 - 72.
Yang C, Zhang X, Guo Y, Meng F, Sachs F, Guo J. Mechanical dynamics in stay cells and fluorescence-based drive/rigidity sensors. Biochim Biophys Acta Mol Cell Res. 2015;1853(8):1889–1904. pmid:25958335 - 73.
Grashoff C, Hoffman BD, Brenner MD, Zhou R, Parsons M, Yang MT, et al. Measuring mechanical rigidity throughout vinculin reveals regulation of focal adhesion dynamics. Nature. 2010 Jul 8;466(7303):263–6. pmid:20613844 - 74.
Carley E, Stewart RK, Zieman A, Jalilian I, King DE, Zubek A, et al. The linc advanced transmits integrin-dependent rigidity to the nuclear lamina and represses epidermal differentiation. Elife. 2021 Mar 1;10. - 75.
Fenelon KD, Thomas E, Samani M, Zhu M, Tao H, Solar Y, et al. Transgenic drive sensors and software program to measure drive transmission throughout the mammalian nuclear envelope in vivo. Biol Open. 2022;11(11). pmid:36350289 - 76.
Guo J, Sachs F, Meng F. Fluorescence-based drive/rigidity sensors: A novel software to visualise mechanical forces in structural proteins in stay cells. Antioxid Redox Sign. 2014;20(6):986–999. pmid:24205787 - 77.
Jin X, Rosenbohm J, Minnick G, Esfahani AM, Safa BT, Yang R. Cell characterization by nanonewton drive sensing. In: Robotics for Cell Manipulation and Characterization. Tutorial Press; 2023. p. 245–70. - 78.
Das R, Lin LC, Català-Castro F, Malaiwong N, Sanfeliu-Cerdán N, Porta-De-la-Riva M, et al. An uneven mechanical code ciphers curvature-dependent proprioceptor exercise. Sci Adv. 2021 Sep 1;7(38):4617–34. pmid:34533987 - 79.
Brenner MD, Zhou R, Conway DE, Lanzano L, Gratton E, Schwartz MA, et al. Spider Silk Peptide Is a Compact, Linear Nanospring Perfect for Intracellular Stress Sensing. Nano Lett. 2016 Mar 9;16(3):2096–102. pmid:26824190 - 80.
Value AL, Ringer P, Chrostek-Grashoff A, Grashoff C. Tips on how to Measure Molecular Forces in Cells: A Information to Evaluating Genetically-Encoded FRET-Based mostly Stress Sensors. Cell Mol Bioeng. 2015;8(1):96–105. pmid:25798203 - 81.
Gadella TWJ. Fret and Flim Methods. van der Vliet PC, Pillai S, editors. Elsevier; 2009. p. 1–534. - 82.
Ishikawa-Ankerhold HC, Ankerhold R, Drummen GPC. Superior fluorescence microscopy techniques-FRAP, FLIP, FLAP, FRET and FLIM. Molecules. 2012;17(4):4047–4132. pmid:22469598 - 83.
Borghi N, Sorokina M, Shcherbakova OG, Weis WI, Pruitt BL, Nelson WJ, et al. E-cadherin is beneath constitutive actomyosin-generated rigidity that’s elevated at cell-cell contacts upon externally utilized stretch. Proc Natl Acad Sci U S A. 2012;109(31):12568–12573. pmid:22802638 - 84.
Haas AJ, Zihni C, Ruppel A, Hartmann C, Ebnet Ok, Tada M, et al. Interaction between Extracellular Matrix Stiffness and JAM-A Regulates Mechanical Load on ZO-1 and Tight Junction Meeting. Cell Rep. 2020 Jul 21;32(3):107924. pmid:32697990 - 85.
Tao H, Zhu M, Lau Ok, Whitley OKW, Samani M, Xiao X, et al. Oscillatory cortical forces promote three dimensional cell intercalations that form the murine mandibular arch. Nat Commun. 2019 Apr 12;10(1):1–18. - 86.
Arsenovic PT, Ramachandran I, Bathula Ok, Zhu R, Narang JD, Noll NA, et al. Nesprin-2G, a Element of the Nuclear LINC Complicated, Is Topic to Myosin-Dependent Stress. Biophys J [Internet]. 2016;110(1):34–43. pmid:26745407 - 87.
Kannan M, Vasan G, Huang C, Haziza S, Li JZ, Inan H, et al. Quick, in vivo voltage imaging utilizing a pink fluorescent indicator. Nat Strategies. 2018 Nov 12;15(12):1108–16. pmid:30420685 - 88.
Gong Y, Huang C, Li JZ, Grewe BF, Zhang Y, Eismann S, et al. Excessive-speed recording of neural spikes in awake mice and flies with a fluorescent voltage sensor. Science (1979) [Internet]. 2015 Dec 11 [cited 2023 Aug 17];350(6266):1361–6. Accessible from: www.sciencemag.org/content/350/6266/1357/suppl/DC1. pmid:26586188 - 89.
Potzkei J, Kunze M, Drepper T, Gensch T, Jaeger KE, Büchs J. Actual-time dedication of intracellular oxygen in micro organism utilizing a genetically encoded FRET-based biosensor. BMC Biol. 2012 Mar 22;10(1). pmid:22439625 - 90.
Papkovsky DB, Dmitriev RI. Organic detection by optical oxygen sensing. Chem Soc Rev. 2013;42:8700. pmid:23775387 - 91.
Tyas L, Brophy VA, Pope A, Rivett AJ, Tavaré JM. Speedy caspase-3 activation throughout apoptosis revealed utilizing fluorescence-resonance vitality switch. EMBO Rep [Internet]. 2000 Sep 1 [cited 2023 Sep 11];1(3):266–70. pmid:11256610 - 92.
Suzuki M, Shindo Y, Yamanaka R, Oka Ok. Dwell imaging of apoptotic signaling circulation utilizing tunable combinatorial FRET-based bioprobes for cell inhabitants evaluation of caspase cascades. Sci Rep [Internet]. 2022 Dec 7 [cited 2023 Sep 11];12(1):1–12. Accessible from: https://www.nature.com/articles/s41598-022-25286-z. - 93.
Wu X, Simone J, Hewgill D, Siegel R, Lipsky PE, He L. Measurement of two caspase actions concurrently in residing cells by a novel twin FRET fluorescent indicator probe. Cytometry A. 2006 Jun;69(6):477–86. pmid:16683263 - 94.
Takemoto Ok, Nagai T, Miyawaki A, Miura M. Spatio-temporal activation of caspase revealed by indicator that’s insensitive to environmental results. J Cell Biol [Internet]. 2003 Jan 20 [cited 2023 Sep 11];160(2):235–43. pmid:12527749 - 95.
Bozza WP, Di X, Takeda Ok, R Rosado LA, Pariser S, Zhang B. The Use of a Stably Expressed FRET Biosensor for Figuring out the Efficiency of Most cancers Medication. PLoS ONE [Internet]. 2014 [cited 2023 Sep 11];9(9):e107010. Accessible from: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0107010. pmid:25188024 - 96.
Onuki R, Nagasaki A, Kawasaki H, Baba T, Uyeda TQP, Taira Ok. Affirmation by FRET in particular person residing cells of the absence of serious amyloid β-mediated caspase 8 activation. Proc Natl Acad Sci U S A [Internet]. 2002 Nov 12 [cited 2023 Sep 13];99(23):14716–21. Accessible from: https://www.pnas.org/doi/abs/10.1073/pnas.232177599. - 97.
Kominami Ok, Nagai T, Sawasaki T, Tsujimura Y, Yashima Ok, Sunaga Y, et al. In Vivo Imaging of Hierarchical Spatiotemporal Activation of Caspase-8 throughout Apoptosis. PLoS ONE [Internet]. 2012 Nov 21 [cited 2023 Sep 13];7(11):e50218. Accessible from: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0050218. pmid:23185580 - 98.
Ng S, Lim HS, Ma Q, Gao Z. Optical aptasensors for adenosine triphosphate. Theranostics. 2016;6(10):1683–1702. pmid:27446501 - 99.
Imamura H, Huynh Nhat KP, Togawa H, Saito Ok, Iino R, Kato-Yamada Y, et al. Visualization of ATP ranges inside single residing cells with fluorescence resonance vitality transfer-based genetically encoded indicators. Proc Natl Acad Sci U S A. 2009 Sep 15;106(37):15651–6. pmid:19720993 - 100.
Trevisiol A, Saab AS, Winkler U, Marx G, Imamura H, Möbius W, et al. Monitoring ATP dynamics in electrically lively white matter tracts. Elife. 2017 Apr 17;6. pmid:28414271 - 101.
Kishikawa JI, Fujikawa M, Imamura H, Yasuda Ok, Noji H, Ishii N, et al. MRT Letter: Expression of ATP Sensor Protein in Caenorhabditis elegans. Microsc Res Tech. 2012. pmid:22038755 - 102.
De Col V, Fuchs P, Nietzel T, Elsässer M, Voon CP, Candeo A, et al. ATP sensing in residing plant cells reveals tissue gradients and stress dynamics of vitality physiology. Elife. 2017 Jul 18;6. pmid:28716182 - 103.
Prinz A, Diskar M, Herberg FW. Software of bioluminescence resonance vitality switch (BRET) for biomolecular interplay research. Vol. 7, ChemBioChem. John Wiley & Sons, Ltd; 2006. p. 1007–12. pmid:16755626 - 104.
Pfleger KDG, Eidne KA. Illuminating insights into protein-protein interactions utilizing bioluminescence resonance vitality switch (BRET). Nat Strategies. 2006;3:165–174. pmid:16489332 - 105.
Eder D, Basler Ok, Aegerter CM. Difficult FRET-based E-Cadherin drive measurements in Drosophila. Sci Rep. 2017;7(1). pmid:29057959 - 106.
Boute N, Jockers R, Issad T. The usage of resonance vitality switch in high-throughput screening: BRET versus FRET. Tendencies Pharmacol Sci. 2002 Aug 1;23(8):351–4. pmid:12377570 - 107.
Wu Y, Jiang T. Developments in FRET- and BRET-Based mostly Biosensors [Internet]. Vol. 13, Micromachines. Multidisciplinary Digital Publishing Institute; 2022 [cited 2023 Aug 17]. p. 1789. Accessible from: https://www.mdpi.com/2072-666X/13/10/1789/htm. pmid:36296141 - 108.
Erdenee E, Ting AY. A Twin-Goal Actual-Time Indicator and Transcriptional Integrator for Calcium Detection in Dwelling Cells. ACS Synth Biol [Internet]. 2022 Mar 18 [cited 2023 Aug 22];11(3):1086–95. pmid:35254056 - 109.
Valkovic AL, Leckey MB, Whitehead AR, Hossain MA, Inoue A, Kocan M, et al. Actual-time examination of cAMP exercise at relaxin household peptide receptors utilizing a BRET-based biosensor. Pharmacol Res Perspect. 2018 Oct 1;6(5). pmid:30263124 - 110.
Jiang LI, Collins J, Davis R, Lin KM, DeCamp D, Roach T, et al. Use of a cAMP BRET Sensor to Characterize a Novel Regulation of cAMP by the Sphingosine 1-Phosphate/G13 Pathway. J Biol Chem. 2007 Apr 6;282(14):10576–84. pmid:17283075 - 111.
Ramirez MP, Anderson MJM, Kelly MD, Sundby LJ, Hagerty AR, Wenthe SJ, et al. Dystrophin missense mutations alter focal adhesion rigidity and mechanotransduction. Proc Natl Acad Sci U S A [Internet]. 2022 Jun 21 [cited 2023 Jul 24];119(25):e2205536119. Accessible from: https://www.pnas.org/doi/abs/10.1073/pnas.2205536119. pmid:35700360 - 112.
Oshino R, Oshino N, Tamura M, Kobilinsky L, Probability B. A delicate bacterial luminescence probe for O2 in biochemical methods. Biochim Biophys Acta. 1972 Jun 26;273(1):5–17. - 113.
Den Hamer A, Dierickx P, Arts R, De Vries JSPM, Brunsveld L, Merkx M. Brilliant Bioluminescent BRET Sensor Proteins for Measuring Intracellular Caspase Exercise. ACS Sens [Internet]. 2017 Jun 23 [cited 2023 Aug 13];2(6):729–34. Accessible from: https://pubs.acs.org/sharingguidelines. pmid:28670623 - 114.
Xu Y, Piston DW, Johnson CH. A bioluminescence resonance vitality switch (BRET) system: Software to interacting circadian clock proteins. Proc Natl Acad Sci U S A [Internet]. 1999 Jan 5 [cited 2023 Aug 13];96(1):151–6. pmid:9874787 - 115.
Wu Y, Jiang T. Developments in FRET- and BRET-Based mostly Biosensors. Vol. 13, Micromachines. Multidisciplinary Digital Publishing Institute; 2022. p. 1789. pmid:36296141 - 116.
Khakhar A, Starker CG, Chamness JC, Lee N, Stokke S, Wang C, et al. Constructing customizable auto-luminescent luciferase-based reporters in vegetation. Elife. 2020 Mar 1;9. - 117.
Mitiouchkina T, Mishin AS, Somermeyer LG, Markina NM, Chepurnyh TV, Guglya EB, et al. Crops with genetically encoded autoluminescence. Nat Biotechnol. 2020;38(8):944–946. pmid:32341562 - 118.
Müller Ok, Engesser R, Timmer J, Zurbriggen MD, Nagy F, Weber W. Synthesis of phycocyanobilin in mammalian cells. Chem Commun. 2013 Sep 10;49(79):8970–2. pmid:23963496 - 119.
Zhou Y, Kong D, Wang X, Yu G, Wu X, Guan N, et al. A small and extremely delicate pink/far-red optogenetic change for purposes in mammals. Nat Biotechnol. 2022 Oct 4;40(2):262–72. pmid:34608325 - 120.
Rodriguez-Garcia A, Rojo-Ruiz J, Navas-Navarro P, Aulestia FJ, Gallego-Sandin S, Garcia-Sancho J, et al. GAP, an aequorin-based fluorescent indicator for imaging Ca2+ in organelles. Proc Natl Acad Sci U S A. 2014 Feb 18;111(7):2584–9. pmid:24501126 - 121.
Yang C, Zhang X, Guo Y, Meng F, Sachs F, Guo J. Mechanical dynamics in stay cells and fluorescence-based drive/rigidity sensors. Biochim Biophys Acta Mol Cell Res [Internet]. 2015 [cited 2023 Jul 24];1853(8):1889–904. pmid:25958335 - 122.
Baird GS, Zacharias DA, Tsien RY. Round permutation and receptor insertion inside inexperienced fluorescent proteins. Proc Natl Acad Sci U S A. 1999 Sep 28;96(20):11241–6. pmid:10500161 - 123.
Topell S, Hennecke J, Glockshuber R. Circularly permuted variants of the inexperienced fluorescent protein. FEBS Lett. 1999 Aug 27;457(2):283–9. pmid:10471794 - 124.
Iwai S, Uyeda TQP. Visualizing myosin-actin interplay with a genetically-encoded fluorescent pressure sensor. Proc Natl Acad Sci U S A. 2008 Nov 4;105(44):16882–7. pmid:18971336 - 125.
De Angelis DA, Miesenböck G, Zemelman BV, Rothman JE. PRIM: Proximity imaging of inexperienced fluorescent protein-tagged polypeptides. Proc Natl Acad Sci U S A. 1998 Oct 13;95(21):12312–6. pmid:9770483 - 126.
Tian L, Hires SA, Mao T, Huber D, Chiappe ME, Chalasani SH, et al. Imaging neural exercise in worms, flies and mice with improved GCaMP calcium indicators. Nat Strategies. 2009;6(12):875–881. pmid:19898485 - 127.
Berg J, Hung YP, Yellen G. A genetically encoded fluorescent reporter of ATP/ADP ratio. Nat Strategies. 2009;6(2):161. pmid:19122669 - 128.
Li J, Yu Q, Ahooghalandari P, Gribble FM, Reimann F, Tengholm A, et al. Submembrane ATP and Ca2+ kinetics in α-cells: Surprising signaling for glucagon secretion. FASEB J. 2015 Aug 1;29(8):3379–88. - 129.
Tantama M, Martínez-François JR, Mongeon R, Yellen G. Imaging vitality standing in stay cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio. Nat Commun [Internet]. 2013 [cited 2023 Jul 29];4. Accessible from: www.nature.com/naturecommunications. pmid:24096541 - 130.
Diuba AV, Samigullin DV, Kaszas A, Zonfrillo F, Malkov A, Petukhova E, et al. CLARITY evaluation of the Cl/pH sensor expression within the mind of transgenic mice. Neuroscience. 2020 Jul 15;439:181–94. pmid:31302264 - 131.
Batti L, Mukhtarov M, Audero E, Ivanov A, Paolicelli O, Zurborg S, et al. Transgenic mouse traces for non-invasive ratiometric monitoring of intracellular chloride. Entrance Mol Neurosci. 2013 Might 21;6(MAY):44911. pmid:23734096 - 132.
Fischer AAM, Kramer MM, Radziwill G, Weber W. Shedding gentle on present tendencies in molecular optogenetics. Curr Opin Chem Biol. 2022 Oct 1;70. pmid:35988347 - 133.
Kichuk TC, Carrasco-López C, Avalos JL, José Avalos CL. Lights up on organelles: Optogenetic instruments to manage subcellular construction and group. WIREs Mech Dis. 2020. pmid:32715616 - 134.
Chia N, Lee SY, Tong Y. Optogenetic instruments for microbial artificial biology. Biotechnol Adv. 2022 Oct 1;59. pmid:35398205 - 135.
Labella ML, Sigrist S, Jørgensen EM. Placing genetics into optogenetics: Knocking out proteins with gentle. In: Optogenetics. 2013:79–90. - 136.
Ledri M, Andersson M, Wickham J, Kokaia M. Optogenetics for controlling seizure circuits for translational approaches. Neurobiol Dis. 2023 Aug 1;184:106234. pmid:37479090 - 137.
Usherenko S, Stibbe H, Muscò M, Essen LO, Kostina EA, Taxis C. Photograph-sensitive degron variants for tuning protein stability by gentle. BMC Syst Biol [Internet]. 2014 Nov 18 [cited 2023 Jul 20];8(1):128. pmid:25403319 - 138.
Usherenko S, Stibbe H, Muscò M, Essen LO, Kostina EA, Taxis C. Photograph-sensitive degron variants for tuning protein stability by gentle. BMC Syst Biol. 2014 Nov 18;8(1):128. pmid:25403319 - 139.
Renicke C, Schuster D, Usherenko S, Essen LO, Taxis C. A LOV2 domain-based optogenetic software to manage protein degradation and mobile operate. Chem Biol. 2013;20(4):619–626. pmid:23601651 - 140.
Bonger KM, Rakhit R, Payumo AY, Chen JK, Wandless TJ. Normal methodology for regulating protein stability with gentle. ACS Chem Biol. 2014 Jan 17;9(1):111–5. pmid:24180414 - 141.
Bonger KM, Chen LC, Liu CW, Wandless TJ. Small-molecule displacement of a cryptic degron causes conditional protein degradation. Nat Chem Biol. 2011;7(8):531–537. pmid:21725303 - 142.
Stevens LM, Kim G, Koromila T, Steele JW, McGehee J, Stathopoulos A, et al. Mild-dependent N-end rule-mediated disruption of protein operate in Saccharomyces cerevisiae and Drosophila melanogaster. PLoS Genet. 2021 Might 17;17(5):e1009544. pmid:33999957 - 143.
Varshavsky A. N-degron and C-degron pathways of protein degradation. Proc Natl Acad Sci U S A. 2019 Jan 8;116(2):358–66. pmid:30622213 - 144.
Heo AJ, Bin KS, Kwon YT, Ji CH. The N-degron pathway: From primary science to therapeutic purposes. Vol. 1866, Biochimica et Biophysica Acta—Gene Regulatory Mechanisms. Elsevier; 2023. p. 194934. pmid:36990317 - 145.
Ryu KA, Kaszuba CM, Bissonnette NB, Oslund RC, Fadeyi OO. Interrogating organic methods utilizing visible-light-powered catalysis. Vol. 5, Nature Evaluations Chemistry. Nature Publishing Group; 2021. p. 322–37. pmid:37117838 - 146.
Takemoto Ok, Sekiya T. Optical manipulation of molecular operate by chromophore-assisted gentle inactivation. Proc Jpn Acad Ser B. 2021 Apr 9;97(4):197–209. pmid:33840676 - 147.
Liao JC, Roider J, Jay DG. Chromophore-assisted laser inactivation of proteins is mediated by the photogeneration of free radicals. Proc Natl Acad Sci U S A. 1994 Mar 29;91(7):2659–63. pmid:8146171 - 148.
Bulina ME, Chudakov DM, Britanova OV, Yanushevich YG, Staroverov DB, Chepurnykh TV, et al. A genetically encoded photosensitizer. Nat Biotechnol. 2006 Dec 20;24(1):95–9. pmid:16369538 - 149.
Riani YD, Matsuda T, Takemoto Ok, Nagai T. Inexperienced monomeric photosensitizing fluorescent protein for photo-inducible protein inactivation and cell ablation. BMC Biol. 2018 Apr 30;16(1):1–12. - 150.
Takemoto Ok, Matsuda T, Sakai N, Fu D, Noda M, Uchiyama S, et al. SuperNova, a monomeric photosensitizing fluorescent protein for chromophore-assisted gentle inactivation. Sci Rep. 2013 Sep 17;3(1):1–7. pmid:24043132 - 151.
Duffy JB. GAL4 system in Drosophila: A fly geneticist’s Swiss military knife. Genesis. 2002;34(1–2):1–15. pmid:12324939 - 152.
Sadowski I, Ma J, Triezenberg S, Ptashne M. GAL4-VP16 is an unusually potent transcriptional activator. Nature. 1988;335(6190):563–564. pmid:3047590 - 153.
Zhu L, McNamara HM, Toettcher JE. Mild-switchable transcription components obtained by direct screening in mammalian cells. Nat Commun. 2023 Jun 2;14(1):1–16. - 154.
Yumerefendi H, Dickinson DJ, Wang H, Zimmerman SP, Bear JE, Goldstein B, et al. Management of protein exercise and cell destiny specification by way of light-mediated nuclear translocation. PLoS ONE. 2015;10(6):1–19. pmid:26083500 - 155.
Niopek D, Benzinger D, Roensch J, Draebing T, Wehler P, Eils R, et al. Engineering light-inducible nuclear localization indicators for exact spatiotemporal management of protein dynamics in residing cells. 2014 Jul 14;5(1). - 156.
Lerner AM, Yumerefendi H, Goudy OJ, Strahl BD, Kuhlman B. Engineering Improved Photoswitches for the Management of Nucleocytoplasmic Distribution. ACS Synth Biol [Internet]. 2018 Nov 15 [cited 2023 Sep 7];7(12):2898–907. pmid:30441907 - 157.
Yumerefendi H, Lerner AM, Parker Zimmerman S, Hahn Ok, Bear JE, Strahl BD, et al. Mild-induced nuclear export reveals speedy dynamics of epigenetic modifications. Nat Chem Biol [Internet]. 2016 [cited 2023 Aug 14];12. Accessible from: www.nature.com/naturechemicalbiology. pmid:27089030 - 158.
Kögler AC, Kherdjemil Y, Bender Ok, Rabinowitz A, Marco-Ferreres R, Furlong EEM. Extraordinarily speedy and reversible optogenetic perturbation of nuclear proteins in residing embryos. Dev Cell [Internet]. 2021;56(16):2348–2363.e8. Accessible from: https://www.sciencedirect.com/science/article/pii/S1534580721005943. pmid:34363757 - 159.
Niopek D, Wehler P, Roensch J, Eils R, Di Ventura B. Optogenetic management of nuclear protein export. Nat Commun. 2016;7:1–9. pmid:26853913 - 160.
Qiao J, Peng H, Dong B. Improvement and Software of an Optogenetic Manipulation System to Suppress Actomyosin Exercise in Ciona Dermis. Int J Mol Sci. 2023;24(6). pmid:36982781 - 161.
Berlew EE, Kuznetsov IA, Yamada Ok, Bugaj LJ, Boerckel JD, Chow BY. Single-Element Optogenetic Instruments for Inducible RhoA GTPase Signaling. Adv Biol. 2021 Sep 1;5(9):2100810. pmid:34288599 - 162.
Hannanta-Anan P, Glantz ST, Chow BY. Optically inducible membrane recruitment and signaling methods. Vol. 57, Present Opinion in Structural Biology. 2019. p. 84–92. pmid:30884362 - 163.
Berlew EE, Kuznetsov IA, Yamada Ok, Bugaj LJ, Chow BY. Optogenetic Rac1 engineered from membrane lipid-binding RGS-LOV for inducible lamellipodia formation. Photochem Photobiol Sci. 2020 Mar 1;19(3):353–61. pmid:32048687 - 164.
Nash AI, McNulty R, Shillito ME, Swartz TE, Bogomolni RA, Luecke H, et al. Structural foundation of photosensitivity in a bacterial light-oxygen-voltage/ helix-turn-helix (LOV-HTH) DNA-binding protein. Proc Natl Acad Sci U S A. 2011 Jun 7;108(23):9449–54. pmid:21606338 - 165.
Rivera-Cancel G, Motta-Mena LB, Gardner KH. Identification of pure and synthetic DNA substrates for light-activated LOV-HTH transcription issue EL222. Biochemistry. 2012 Dec 18;51(50):10024–34. pmid:23205774 - 166.
Gligorovski V, Sadeghi A, Rahi SJ. Multidimensional characterization of inducible promoters and a extremely light-sensitive LOV-transcription issue. Nat Commun. 2023 Jun 27;14(1):1–18. - 167.
Wang Z, Yan Y, Zhang H. Design and Characterization of an Optogenetic System in Pichia pastoris. ACS Synth Biol. 2022 Jan 21;11(1):297–307. pmid:34994189 - 168.
Zhao EM, Lalwani MA, Lovelett RJ, Garciá-Echauri SA, Hoffman SM, Gonzalez CL, et al. Design and Characterization of Speedy Optogenetic Circuits for Dynamic Management in Yeast Metabolic Engineering. ACS Synth Biol. 2020 Dec 18;9(12):3254–66. pmid:33232598 - 169.
Zhao EM, Lalwani MA, Chen JM, Orillac P, Toettcher JE, Avalos JL. Optogenetic Amplification Circuits for Mild-Induced Metabolic Management. ACS Synth Biol. 2021 Might 21;10(5):1143–54. pmid:33835777 - 170.
Zhao EM, Zhang Y, Mehl J, Park H, Lalwani MA, Toettcher JE, et al. Optogenetic regulation of engineered mobile metabolism for microbial chemical manufacturing. Nature. 2018 Mar 21;555(7698):683–7. pmid:29562237 - 171.
Pathak GP, Strickland D, Vrana JD, Tucker CL. Benchmarking of optical dimerizer methods. ACS Synth Biol. 2014 Nov 21;3(11):832–8. pmid:25350266 - 172.
Guntas G, Hallett RA, Zimmerman SP, Williams T, Yumerefendi H, Bear JE, et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and exercise of signaling proteins. Proc Natl Acad Sci U S A. 2015 Jan 6;112(1):112–7. pmid:25535392 - 173.
Lungu OI, Hallett RA, Choi EJ, Aiken MJ, Hahn KM, Kuhlman B. Designing Photoswitchable Peptides Utilizing the AsLOV2 Area. Chem Biol. 2012 Apr 20;19(4):507–17. pmid:22520757 - 174.
Keenan SE, Blythe SA, Marmion RA, Djabrayan NJV, Wieschaus EF, Shvartsman SY. Speedy Dynamics of Sign-Dependent Transcriptional Repression by Capicua. Dev Cell. 2020 Mar 23;52(6):794–801.e4. pmid:32142631 - 175.
Johnson HE, Goyal Y, Pannucci NL, Schüpbach T, Shvartsman SY, Toettcher JE. The Spatiotemporal Limits of Developmental Erk Signaling. Dev Cell. 2017 Jan 23;40(2):185–92. pmid:28118601 - 176.
Goglia AG, Wilson MZ, Jena SG, Silbert J, Basta LP, Devenport D, et al. A Dwell-Cell Display for Altered Erk Dynamics Reveals Ideas of Proliferative Management. Cell Syst. 2020 Mar 25;10(3):240–253.e6. pmid:32191874 - 177.
Farahani PE, Lemke SB, Dine E, Uribe G, Toettcher JE, Nelson CM. Substratum stiffness regulates Erk signaling dynamics via receptor-level management. Cell Rep. 2021;37(13). pmid:34965432 - 178.
Johnson HE. Software of Optogenetics to Probe the Signaling Dynamics of Cell Destiny Resolution-Making. In: Computational Modeling of Signaling Networks. Springer; 2023. p. 315–26. - 179.
Kennedy MJ, Hughes RM, Peteya LA, Schwartz JW, Ehlers MD, Tucker CL. Speedy blue-light-mediated induction of protein interactions in residing cells. Nat Strategies. 2010;7(12):973–975. pmid:21037589 - 180.
Bugaj LJ, Choksi AT, Mesuda CK, Kane RS, Schaffer D V. Optogenetic protein clustering and signaling activation in mammalian cells. Nat Strategies. 2013 Feb 3;10(3):249–52. pmid:23377377 - 181.
Singh AP, Wu P, Ryabichko S, Raimundo J, Swan M, Wieschaus E, et al. Optogenetic management of the Bicoid morphogen reveals quick and sluggish modes of hole gene regulation. Cell Rep. 2022 Mar 22;38(12):110543. pmid:35320726 - 182.
McDaniel SL, Gibson TJ, Schulz KN, Fernandez Garcia M, Nevil M, Jain SU, et al. Continued Exercise of the Pioneer Issue Zelda Is Required to Drive Zygotic Genome Activation. Mol Cell. 2019 Apr 4;74(1):185–195.e4. pmid:30797686 - 183.
Crefcoeur RP, Yin R, Ulm R, Halazonetis TD. Ultraviolet-B-mediated induction of protein-protein interactions in mammalian cells. Nat Commun. 2013 Apr 30;4(1):1–7. pmid:23653191 - 184.
Yang X, Jost APT, Weiner OD, Tang C. A lightweight-inducible organelle-targeting system for dynamically activating and inactivating signaling in budding yeast. Mol Biol Cell. 2013 Aug 1;24(15):2419–30. pmid:23761071 - 185.
Toettcher JE, Weiner OD, Lim WA. Utilizing Optogenetics to Interrogate the Dynamic Management of Sign Transmission by the Ras/Erk Module. Cell. 2013 Dec 5;155(6):1422–34. pmid:24315106 - 186.
Lan TH, He L, Huang Y, Zhou Y. Optogenetics for transcriptional programming and genetic engineering. Tendencies Genet [Internet]. 2022 Dec 1 [cited 2023 Sep 6];38(12):1253–70. Accessible from: http://www.cell.com/article/S0168952522001408/fulltext. pmid:35738948 - 187.
Braatsch S, Johnson JA, Noll Ok, Beatty JT. The O2-responsive repressor PpsR2 however not PpsR1 transduces a light-weight sign sensed by the BphP1 phytochrome in Rhodopseudomonas palustris CGA009. FEMS Microbiol Lett. 2007 Jul 1;272(1):60–4. pmid:17456182 - 188.
Redchuk TA, Kaberniuk AA, Verkhusha VV. Close to-infrared light-controlled methods for gene transcription regulation, protein focusing on and spectral multiplexing. Nat Protoc. 2018 Apr 26;13(5):1121–36. pmid:29700485 - 189.
Redchuk TA, Omelina ES, Chernov KG, Verkhusha V V. Close to-infrared optogenetic pair for protein regulation and spectral multiplexing. Nat Chem Biol. 2017 Mar 27;13(6):633–9. pmid:28346403 - 190.
Benedetti L, Marvin JS, Falahati H, Guillén-Samander A, Looger LL, De Camilli P. Optimized vivid-derived magnets photodimerizers for subcellular optogenetics in mammalian cells. Elife. 2020;9:1–49. pmid:33174843 - 191.
Kawano F, Suzuki H, Furuya A, Sato M. Engineered pairs of distinct photoswitches for optogenetic management of mobile proteins. Nat Commun. 2015 Feb 24;6(1):1–8. pmid:25708714 - 192.
Zoltowski BD, Crane BR. Mild activation of the LOV protein vivid generates a quickly exchanging dimer. Biochemistry. 2008 Jul 8;47(27):7012–9. pmid:18553928 - 193.
Zoltowski BD, Schwerdtfeger C, Widom J, Loros JJ, Bilwes AM, Dunlap JC, et al. Conformational switching within the fungal gentle sensor vivid. Science (1979). 2007 Might 18;316(5827):1054–7. pmid:17510367 - 194.
Vaidya AT, Chen CH, Dunlap JC, Loros JJ, Crane BR. Construction of a light-activated LOV protein dimer that regulates transcription. Sci Sign. 2011 Aug 2;4(184). pmid:21868352 - 195.
Wang X, Chen X, Yang Y. Spatiotemporal management of gene expression by a light-switchable transgene system. Nat Strategies. 2012;9(3):266–269. pmid:22327833 - 196.
Yuan H, Bauer CE. PixE promotes darkish oligomerization of the BLUF photoreceptor PixD. Proc Natl Acad Sci U S A. 2008 Aug 19;105(33):11715–9. pmid:18695243 - 197.
Stierl M, Stumpf P, Udwari D, Gueta R, Hagedorn R, Losi A, et al. Mild modulation of mobile cAMP by a small bacterial photoactivated adenylyl cyclase, bPAC, of the soil bacterium Beggiatoa. J Biol Chem. 2011;286(2):1181–1188. pmid:21030594 - 198.
Wang H, Vilela M, Winkler A, Tarnawski M, Schlichting I, Yumerefendi H, et al. LOVTRAP: An optogenetic system for photoinduced protein dissociation. Nat Strategies. 2016 Jul 18;13(9):755–8. pmid:27427858 - 199.
Strickland D, Lin Y, Wagner E, Hope CM, Zayner J, Antoniou C, et al. TULIPs: tunable, light-controlled interacting protein tags for cell biology. Nat Strategies [Internet]. 2012 Apr 4;9(4):379–84. Accessible from: https://www.nature.com/articles/nmeth.1904. pmid:22388287 - 200.
Huang J, Koide A, Makabe Ok, Koide S. Design of protein operate leaps by directed area interface evolution. Proc Natl Acad Sci U S A [Internet]. 2008 Might 6 [cited 2023 Sep 13];105(18):6578–83. Accessible from: www.pnas.orgcgidoi10.1073pnas.0801097105. pmid:18445649 - 201.
Zhang W, Lohman AW, Zhuravlova Y, Lu X, Wiens MD, Hoi H, et al. Optogenetic management with a photocleavable protein, Phocl. Nat Strategies. 2017 Mar 13;14(4):391–4. pmid:28288123 - 202.
Kato HE, Kamiya M, Sugo S, Ito J, Taniguchi R, Orito A, et al. Atomistic design of microbial opsin-based blue-shifted optogenetics instruments. Nat Commun [Internet]. 2015 Might 15 [cited 2023 Sep 7];6(1):1–10. Accessible from: https://www.nature.com/articles/ncomms8177. pmid:25975962 - 203.
Rost BR, Schneider F, Grauel MK, Wozny C, Bentz C, Blessing A, et al. Optogenetic acidification of synaptic vesicles and lysosomes. Nat Neurosci. 2015 Dec 9;18(12):1845–52. pmid:26551543 - 204.
Miesenböck G, De Angelis DA, Rothman JE. Visualizing secretion and synaptic transmission with pH-sensitive inexperienced fluorescent proteins. Nature. 1998 Dec 30;394(6689):192–5. pmid:9671304 - 205.
Shevchenko V, Mager T, Kovalev Ok, Polovinkin V, Alekseev A, Juettner J, et al. Inward H+ pump xenorhodopsin: Mechanism and various optogenetic strategy. Sci Adv [Internet]. 2017 [cited 2023 Sep 11];3(9). pmid:28948217 - 206.
Inoue Ok, Ono H, Abe-Yoshizumi R, Yoshizawa S, Ito H, Kogure Ok, et al. A lightweight-driven sodium ion pump in marine micro organism. Nat Commun [Internet]. 2013 Apr 9 [cited 2023 Sep 11];4(1):1–10. Accessible from: https://www.nature.com/articles/ncomms2689. pmid:23575682 - 207.
Feroz H, Ferlez B, Lefoulon C, Ren T, Baker CS, Gajewski JP, et al. Mild-Pushed Chloride Transport Kinetics of Halorhodopsin. Biophys J [Internet]. 2018 Jul 7 [cited 2023 Sep 11];115(2):353. pmid:30021110 - 208.
Govorunova EG, Sineshchekov OA, Li H, Wang Y, Brown LS, Spudich JL. RubyACRs, nonalgal anion channelrhodopsins with extremely red-shifted absorption. Proc Natl Acad Sci U S A [Internet]. 2020 Sep 15 [cited 2023 Sep 13];117(37):22833–40. pmid:32873643 - 209.
Scheib U, Stehfest Ok, Gee CE, Körschen HG, Fudim R, Oertner TG, et al. The rhodopsin-guanylyl cyclase of the aquatic fungus Blastocladiella emersonii permits quick optical management of cGMP signaling. Sci Sign [Internet]. 2015 [cited 2024 Feb 5];8(389). Accessible from: www.SCIENCESIGNALING.org. pmid:26268609 - 210.
Yoshida Ok, Tsunoda SP, Brown LS, Kandori H. A singular choanoflagellate enzyme rhodopsin displays lightdependent cyclic nucleotide phosphodiesterase exercise. J Biol Chem [Internet]. 2017 Might 5 [cited 2024 Feb 5];292(18):7531–41. Accessible from: http://www.jbc.org/article/S0021925820428637/fulltext. pmid:28302718 - 211.
Gao S, Nagpal J, Schneider MW, Kozjak-Pavlovic V, Nagel G, Gottschalk A. Optogenetic manipulation of cGMP in cells and animals by the tightly light-regulated guanylyl-cyclase opsin CyclOp. Nat Commun [Internet]. 2015 [cited 2024 Feb 5];6. Accessible from: www.nature.com/naturecommunications. pmid:26345128 - 212.
Lee D, Creed M, Jung Ok, Stefanelli T, Wendler DJ, Oh WC, et al. Temporally exact labeling and management of neuromodulatory circuits within the mammalian mind. Nat Strategies. 2017 Apr 3;14(5):495–503. pmid:28369042 - 213.
Lee S, Park H, Kyung T, Kim NY, Kim S, Kim J, et al. Reversible protein inactivation by optogenetic trapping in cells. Nat Strategies. 2014;11(6):633–636. pmid:24793453 - 214.
Patel A, Lee HO, Jawerth L, Maharana S, Jahnel M, Hein MY, et al. A Liquid-to-Strong Part Transition of the ALS Protein FUS Accelerated by Illness Mutation. Cell. 2015 Aug 27;162(5):1066–77. pmid:26317470 - 215.
Dine E, Gil AA, Uribe G, Brangwynne CP, Toettcher JE. Protein Part Separation Gives Lengthy-Time period Reminiscence of Transient Spatial Stimuli. Cell Syst. 2018 Jun 27;6(6):655–663.e5. pmid:29859829 - 216.
Yumerefendi H, Wang H, Ickinson DD, Erner AL, Malkus P, Goldstein B, et al. Mild-Dependent Cytoplasmic Recruitment Enhances the Dynamic Vary of a Nuclear Import Photoswitch. Chembiochem. 2018. pmid:29446199 - 217.
Chen SY, Osimiri LC, Chevalier M, Bugaj LJ, Nguyen TH, Greenstein RA, et al. Optogenetic Management Reveals Differential Promoter Interpretation of Transcription Issue Nuclear Translocation Dynamics. Cell Syst. 2020;11(4):336–353.e24. pmid:32898473 - 218.
Redchuk TA, Omelina ES, Chernov KG, Verkhusha VV. Close to-infrared optogenetic pair for protein regulation and spectral multiplexing. Nat Chem Biol [Internet]. 2017 Mar 27 [cited 2023 Aug 26];13(6):633–9. Accessible from: https://www.nature.com/articles/nchembio.2343 pmid:28346403 - 219.
Kim MW, Wang W, Sanchez MI, Coukos R, von Zastrow M, Ting AY. Time-gated detection of protein-protein interactions with transcriptional readout. Elife. 2017;30:6. pmid:29189201 - 220.
Jang J, Tang Ok, Youn J, McDonald S, Beyer HM, Zurbriggen MD, et al. Engineering of bidirectional, cyanobacteriochrome-based light-inducible dimers (BICYCL)s. Nat Strategies [Internet]. 2023 Feb 23 [cited 2023 Aug 22];20(3):432–41. Accessible from: https://www.nature.com/articles/s41592-023-01764-8. pmid:36823330 - 221.
Baaske J, Gonschorek P, Engesser R, Dominguez-Monedero A, Raute Ok, Fischbach P, et al. Twin-controlled optogenetic system for the speedy down-regulation of protein ranges in mammalian cells. Sci Rep. 2018 Oct 9;8(1):1–10. - 222.
Kaberniuk AA, Baloban M, Monakhov MV, Shcherbakova DM, Verkhusha VV. Single-component near-infrared optogenetic methods for gene transcription regulation. Nat Commun [Internet]. 2021 Jun 23 [cited 2023 Aug 16];12(1):1–12. Accessible from: https://www.nature.com/articles/s41467-021-24212-7. - 223.
Sittewelle M, Ferrandiz N, Fesenko M, Royle SJ. Genetically encoded imaging instruments for investigating cell dynamics at a look. J Cell Sci. 2023;136(7):1–7. pmid:37039102 - 224.
Zhao J, Lammers NC, Alamos S, Kim YJ, Martini G, Garcia HG. Optogenetic dissection of transcriptional repression in a multicellular organism. bioRxiv [Internet]. 2023 Apr 28 [cited 2023 Sep 7];2022.11.20.517211. Accessible from: https://www.biorxiv.org/content/10.1101/2022.11.20.517211v2. - 225.
Shafraz O, Marie C, Davis O, Sivasankar S. Mild Activated BioID (LAB): an optically activated proximity labeling system to review protein-protein interactions. bioRxiv [Internet]. 2023 Might 6 [cited 2023 Sep 7];2022.10.22.513249. - 226.
Vendor CA, Cho CY, O’Farrell PH. Speedy embryonic cell cycles defer the institution of heterochromatin by eggless/SetDB1 in Drosophila. Genes Dev [Internet]. 2019 Apr 1 [cited 2023 Aug 22];33(7–8):403–17. Accessible from: http://genesdev.cshlp.org/content/33/7-8/403.full. pmid:30808658 - 227.
Oldak B, Aguilera-Castrejon A, Hanna JH. Current insights into mammalian pure and artificial ex utero embryogenesis. Curr Opin Genet Dev. 2022;77:101988. pmid:36179582 - 228.
Neurovirulence of the Australian outbreak Japanese Encephalitis virus genotype 4 is decrease. [cited 2024 Feb 5]. - 229.
Aguilera-Castrejon A, Oldak B, Shani T, Ghanem N, Itzkovich C, Slomovich S, et al. Ex utero mouse embryogenesis from pre-gastrulation to late organogenesis. Nature [Internet]. 2021 Mar 17 [cited 2024 Feb 5];593(7857):119–24. Accessible from: https://www.nature.com/articles/s41586-021-03416-3. - 230.
Zhang L, Getz SA, Bordey A. Twin in Utero Electroporation in Mice to Manipulate Two Particular Neuronal Populations within the Growing Cortex. Entrance Bioeng Biotechnol. 2022 Jan 12;9:814638. pmid:35096799 - 231.
Huang Q, Cohen MA, Alsina FC, Devlin G, Garrett A, McKey J, et al. Intravital imaging of mouse embryos. Science (1979) [Internet]. 2020 Apr 10 [cited 2024 Feb 5];368(6487):181–6. pmid:32273467