Quotation: Jiang X, Zhao Ok, Solar Y, Tune X, Yi C, Xiong T, et al. (2024) The size of zebrafish pectoral fin buds is decided by intercellular Ok+ ranges and consequent Ca2+-mediated signaling through retinoic acid regulation of Rcan2 and Kcnk5b. PLoS Biol 22(3):
e3002565.
https://doi.org/10.1371/journal.pbio.3002565
Educational Editor: Mary C. Mullins, College of Pennsylvania Faculty of Drugs, UNITED STATES
Obtained: July 21, 2023; Accepted: February 27, 2024; Revealed: March 25, 2024
Copyright: © 2024 Jiang et al. That is an open entry article distributed below the phrases of the Creative Commons Attribution License, which allows unrestricted use, distribution, and copy in any medium, offered the unique creator and supply are credited.
Knowledge Availability: All related information are throughout the paper and its Supporting Information information.
Funding: This work was supported by Nationwide Pure Science Basis of China grants (32070820 to CLA; and 32270881 to CLA), the Shanghai Science and Expertise Fee (22ZR1441200 to CLA), and ShanghaiTech College start-up funds. The funding companies didn’t have any position within the examine design, information assortment, information evaluation, resolution to publish or preparation of the manuscript.
Competing pursuits: The authors have declared that no competing pursuits exist.
Abbreviations:
AER,
apical ectodermal ridge; ER,
endoplasmic reticulum; FLIM,
Fluorescence Lifetime Microscopy; hpf,
hours submit fertilization; IRF,
instrument response issue; RA,
retinoic acid; TH,
thyroid hormone; ZPA,
zone of polarizing exercise
Introduction
The scaling of anatomical constructions includes the coordinated management of gene transcription and intracellular communication. An rising variety of findings are linking electrophysiological adjustments to the regulation of developmental phenomena in numerous organic contexts [1,2]; as examples, transmembrane voltage potential influences eye improvement through regulation of Pax6 [3]; Hedgehog signaling is regulated in Drosophila wing discs by cell depolarization [4]; and inactivation of Ok+ inward rectifying channels results in patterning defects within the craniofacial skeleton, vertebrate phalanges, and fly wings through disruption of BMP/Dpp signaling [5,6]. Moreover, gain-of-function mutations in channels that facilitate outward move of intracellular Ok+ are additionally linked to syndromes that generate craniofacial alterations, neurodevelopmental impairments, defects within the improvement of the limbs, and so forth. [7,8]. As discoveries for the involvement of electrophysiological regulation in improvement enhance, a lot stays unknown about how these endogenous electrophysiological adjustments are built-in into the identified molecular mechanisms that regulate improvement.
In zebrafish, mutations that enhance the exercise of Ok+ channels promote allometric development of juvenile and grownup fish fins [9–13], linking intracellular Ok+ within the management of coordinated proportional development of those anatomical constructions. The Ok+ focus is excessive inside cells, so the opening of Ok+-leak channels usually causes an outward move of Ok+ [14]. Within the grownup fin, rising Ok+-leak channel exercise will increase the transcription of a number of morphogens and promotes development [11]. Nevertheless, it stays unknown how Ok+ channel exercise on the cell membrane is managed and relayed throughout the scaling means of fish appendages, and whether or not an analogous electrophysiological management exists within the conserved embryonic vertebrate fin/limb bud developmental program.
Early embryonic limb and pectoral fin buds kind at particular places within the lateral plate mesoderm that expresses an evolutionary conserved profile of morphogens and development components [15]. The event of limb buds and pectoral fin buds provoke when retinoic acid (RA) not directly promotes the distal transcription of Fgf10 [16], which in flip induces the expression of Fgf8 and the formation of the apical ectodermal ridge (AER) within the distal anterior–posterior interface of ectodermal cells of the buds [17,18]. After formation of the AER, the morphogen Sonic hedgehog (Shh) manifests in a gaggle of bud mesenchymal cells referred to as a zone of polarizing exercise (ZPA). The ZPA varieties within the posterior bud mesenchyme and is required for development and patterning by the actions of Shh [19,20]. Management of the AER and ZPA contain communication between these 2 areas, in addition to different regional cell teams through their morphogens/development components (e.g., Bmp4) [21]. Much like early limb bud improvement, activation of those signaling facilities and different morphogens promote pectoral fin bud outgrowth [22]. For the zebrafish pectoral fin bud, development is underway by 28 to30 hours submit fertilization (hpf) [23]. Throughout limb improvement, development persist till the formation of the distal digits [24], however in zebrafish, pectoral fin bud improvement is proscribed to forming the endochondral bone, musculature, and different tissues on the proximal base of the fin [25,26]. Roughly 24 h after initiation of fin bud outgrowth, its improvement has transitioned into the event of a distinct sort of appendage: the finfold, and most of the morphogen/development issue alerts have rearranged to the distal tip of the fin bud by 54 to 56 hpf to generate the finfold after which in the end larval pectoral fins [23,27]. Whereas the fundamental interactions between signaling facilities and signaling molecules throughout early fin/limb bud improvement is known, it’s unknown whether or not adjustments in intracellular Ok+ will coordinately regulate them in an analogous method because it does throughout developmental/regenerative development of fins.
Utilizing a genetic sensor for Ok+ [28] with Fluorescence Lifetime Microscopy (FLIM), we discovered that relative ranges of intracellular Ok+ lower all through the early pectoral fin bud throughout its development. Once we transgenically overexpressed Ok+-leak channels that lower in intracellular Ok+, we elevated bud development and coordinately elevated the expression a number of morphogens (fgf8a, fgf10a, aldh1a2, shha, and bmp4) that management bud improvement. Remedy with RA, a sign that may promote development of vertebrate appendages, decreased intracellular Ok+ ranges within the buds. We discovered that RA induces rcan2 expression and that rcan2 scales fin buds and reduces intracellular Ok+. As well as, we discovered that the Ok+-leak channel kcnk5b is expressed in pectoral fin buds and that it’s required for Rcan2-mediated scaling. We additionally decided that Kcnk5b promotes depolarization and that depolarization is required for enhanced fin bud development. We additional present that Kcnk5b exercise requires IP3R-mediated Ca2+ launch and CaMKK exercise for SHH transcription in vitro and in vivo in addition to for kcnk5b’s enhancement of fin bud development. Thus, we offer a mechanism by which an vital proximal morphogen (RA) can regulate the exercise of a Ok+-leak channel through Rcan2 to advertise Ca2+-mediated scaling of embryonic pectoral fins buds.
Outcomes
Endogenous intracellular Ok+ ranges lower throughout fin bud development
We beforehand confirmed that Kcnk5b promotes allometric development of grownup fins through hierarchical activation of a number of morphogens [11]. The power of a person Ok+ channel to advertise the expression of a number of developmental morphogens within the grownup fin and larva led us to hypothesize that this phenomenon could also be extra broadly practical, and that it regulates the early developmental program conserved in vertebrate fin/limb buds, which is current within the growing pectoral fin buds of zebrafish [22]. Subsequently, we needed to find out whether or not there are endogenous adjustments in intracellular Ok+ within the growing pectoral fin bud.
To measure intracellular Ok+, we used a longtime FRET-based genetic sensor (KIRIN1) that may detect adjustments particularly in intracellular Ok+ ranges [28]. To beat the constraints related to FRET measurements in vivo, we used FLIM, as a result of FLIM doesn’t depend on intensity-based ratios, slightly, FLIM detects adjustments within the intrinsic decay charge of a donor fluorophore (CPF) when it undergoes FRET to the acceptor fluorophore (YFP) throughout the sensor (Fig 1Aa and 1Ab). One can immediately quantify the amassed decay profile of a fluorophore (CFP) after excitation by laser pulses [29] (Fig 1Ba and 1Bb); consequently, we might detect relative variations in intracellular Ok+ attributable to will increase in Ok+-leak channel expression (S1Aa and S1Ab Fig), or by posttranslational regulation of a Ok+-leak channel (S1B Fig) or by completely different Ok+-leak channel mutants which have completely different actions (S1C Fig) [11]. These experiments confirmed that we are able to detect adjustments in relative intracellular Ok+ ranges with the KIRIN1 sensor utilizing FLIM.
Fig 1. Intracellular Ok+ decreases throughout pectoral fin bud development.
(A) FRET-based mechanism of detection of intracellular Ok+ by KIRIN1 sensor and utility of FLIM evaluation. Adjustments in FRET have an effect on the fluorescence lifetime of the donor fluorophore CFP that’s excited by a 2-Photon laser pulse (a). A discount of intracellular Ok+ is detected as increased (longer) lifetime of the CFP, whereas enhance in Ok+ is detected as its decrease (shorter) lifetime (b). (B) We depict these lifetime (nanoseconds, ns) relationships of upper lifetime equating to decrease intracellular Ok+ (a) and the inverse (unique) as 1/lifetime (ns-1) (b) as a way to extra simply relate the development adjustments with what the FLIM-based measurements are indicating for intracellular Ok+. (C) Design of Ok+ sensor (KIRIN1) transgene for transgenic zebrafish line (a). Warmth-shock technique for inducing transgene throughout completely different developmental time factors of the fin bud and subsequently FLIM-FRET (FILM) imaging it (b). (D) FLIM measurements of the CFP in confocal sections of ectoderm (Ecto) and mesenchyme (Mesen) in pectoral fin buds at 48 hpf utilizing a FRET-based sensor for Ok+. Measurements from a transgene containing solely CFP that’s unable to FRET with YFP. (E) Time course of FLIM measurements of intracellular Ok+ within the ectodermal and mesenchymal tissues of the growing pectoral fin buds. (F) Time course of development measurements of the growing fin buds. (G–I) Z-stacks of confocal FLIM planes in fin buds at 32 hpf (G), 48 hpf (H), and 56 hpf (I). Z-stacks cowl distances of 15.2 μm (32 hpf), 18.36 μm (48 hpf), and 10.3 μm (56 hpf). The lifetime worth of every pixel within the ectoderm (white arrowhead) and the mesenchyme (asterisk) is represented by colours in a rainbow scale from 2.8 ns (blue) to three.4 ns (pink). (J) Era of fin buds that mosaically categorical the KIRIN1 sensor and both mCherry or kcnk5b-mCherry (mosaic for two transgenes) and proposed outcomes. (Ka,b) Pictures of cells in fin buds of 56 hpf embryos harboring hsp70:mCherry (pink asterisks) and cells missing the mCherry transgene (white asterisks). Confocal aircraft for mCherry fluorescence (a). FLIM picture of KIRIN1+ cells of the identical confocal aircraft (b). (La,b) Pictures of cells in fin buds of 56 hpf embryos harboring hsp70:kcnk5b-mCherry (pink asterisks) and cells missing the transgene (white asterisks). Confocal aircraft for mCherry fluorescence (a). FLIM picture of KIRIN1+ cells of the identical confocal aircraft (b). (M) FLIM measurements in fin buds of 56 hpf embryos of indicated cell classes; “close to” signifies cells subsequent or distant to mCherry-positive (mCherry+) or kcnk5b-mCherry-positive (kcnk5b+) cells. The graphs are depicted as 1/lifetime (ns-1) to extra simply relate the portrayal of the values to the associated change in intracellular Ok+ (see S1A Fig). Experiments have been repeated 3 or extra time (N ≥ 3). For FLIM, we measured 2 or 3 places in every tissue of 1 fin bud per embryo. Every measured worth is represented as an information level (D, M), aside from E, wherein the info factors are offered as averages and customary deviations of all of the measurements (≥11) at every time level. For fin bud measurement measurements (F), every information level represents 1 fin bud per embryo. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values >0.05 are designated as “not vital” (NS). Numerical information used on this determine are included in S1 Data. FLIM, Fluorescence Lifetime Microscopy; hpf, hours submit fertilization.
To evaluate the intracellular Ok+ in vivo, we generated a heat-shock-inducible transgenic reporter Tg[hsp70:CFP-KIRIN1-YFP] (hsp70:KIRIN1) line (Fig 1Ca) and used it to detect relative variations in Ok+ ranges through FLIM 6 h after warmth shock (Fig 1Cb). FLIM measurements within the pectoral fin bud confirmed comparatively decrease intracellular Ok+ ranges within the mesenchyme in comparison with ectoderm at 48 hpf (Fig 1D), a time level wherein the expansion of the pectoral fin buds is underway. We subsequently measured a number of time factors throughout pectoral fin bud outgrowth from the identical animals, and we noticed a rise in intracellular Ok+ in each the ectoderm and the mesenchyme between 32 and 34 hpf, however afterwards, intracellular Ok+ ranges decreased to their lowest ranges at round 48 hpf (Fig 1E). Intracellular Ok+ ranges elevated once more between 54 and 56 hpf (Fig 1E). From the plotted time factors, we noticed that though mesenchyme Ok+ ranges remained decrease than the ectoderm, the adjustments in Ok+ in each tissues have been coupled (Fig 1E). These lifetime measurements weren’t influenced by variations in depth ranges (S2D–S2H Fig) or by the strategy for immobilizing the embryos (S2I–S2O Fig). To establish whether or not there’s a relationship between relative adjustments in intracellular Ok+ and development, we measured fin bud sizes between 30 hpf to 56 hpf. From bud space measurements, we noticed incremental development from 30 hpf to 56 hpf (Fig 1F). We additionally noticed 2 pauses in common development between 32 hpf and 34 hpf and 54 hpf and 56 hpf (Fig 1F) that correlated with time factors wherein intracellular Ok+ elevated (Fig 1E). Collectively, these outcomes advised a coordinated regulation of intracellular Ok+ that pertains to fin bud development.
To visualise the relative distribution of intracellular Ok+ throughout pectoral fin bud improvement, we represented the lifetime values alongside a rainbow scale: the decrease the worth (the upper intracellular Ok+ ranges), the extra in the direction of blue; conversely, the upper values (the decrease intracellular Ok+ ranges), the extra in the direction of pink. From the colorized particular person confocal FLIM planes (S3A–S3C Fig) and 3D Z-stacks of FLIM planes by fin buds at 32 hpf (Figs 1Ga–1Ge and S3A), at 48 hpf (Figs 1Ha–1He and S3B), and at 56 hpf (Figs 1Ia–1Ie and S3C), we noticed clear variations in Ok+ ranges between the mesenchyme and ectoderm, however few regional variations inside every tissue. We additionally noticed international decreases in intracellular Ok+ between 32 hpf (Figs 1Ga–1Ge and S3A) and 48 hpf (Figs 1Ha–1He and S3B) when the expansion of the bud is excessive. Conversely, we noticed international will increase in Ok+ between 48 hpf and 56 hpf (Figs 1Ia–1Ie and S3C) when development reduces. From these observations, we suggest that there’s a lower in intracellular Ok+ throughout fin bud development because of will increase in Ok+-leak channel exercise, and these adjustments are linked to the expansion of the bud.
The dearth of regional variations in intracellular Ok+ in both the mesenchyme or ectoderm advised equilibration of Ok+ between the cells of every tissue. To check this speculation, we generated double mosaic embryos by injecting recipient embryos with the heat-shock-inducible hsp70:KIRIN1 and both kcnk5b-mCherry transgene (hsp70:kcnk5b-mCherry) or with mCherry (hsp70:mCherry) transgene as a management (Fig 1J). We recognized mCherry-expressing cells (Fig 1K) or kcnk5b-mCherry cells (Fig 1L) in fin buds after which assessed intracellular Ok+ in mCherry-positive cells (Fig 1Ka, 1La, and 1M and pink asterisks in Fig 1Kb and 1Lb) and the encircling mCherry-negative cells (Fig 1K and 1L, white asterisks). In comparison with cells that harbored the mCherry transgene (Fig 1K, pink asterisks), we noticed that cells expressing kcnk5b-mCherry (Fig 1L, pink asterisks) and cells surrounding the transgene-expressing cells displayed decreases in Ok+ ranges (Figs 1M and S3D). These information indicated that adjustments in intracellular Ok+ are shared such that cells with extra Ok+-leak channel exercise can lower the intracellular Ok+ of neighboring cells.
Overexpression of a Ok+-leak channel in early fin bud coordinately enhances the expression of the vital morphogens to scale proximodistal development
The coordinated lower within the endogenous ranges of Ok+ within the growing pectoral fin bud advised that intracellular Ok+ has a job in bud improvement. Consequently, we wished to know whether or not reducing intracellular Ok+ is adequate to boost the size of the buds. To lower intracellular Ok+, we overexpressed Kcnk5b or Kcnk10 Ok+-leak channel by heat-shock induction of the Tg[hsp70:kcnk5b-GFP] [11] or Tg[hsp70:kcnk10a-GFP] transgenic strains. Six hours after transgene induction at 48 hpf (S4A–S4C Fig), we measured the expansion space of the fin buds and standardized the measurements of every bud to the world of the attention (Fig 2A and 2B) or the otic vesicle (S4D and S4E Fig). From these analyses, we noticed enhanced development of pectoral fin buds attributable to kcnk5b or kcnk10 in comparison with heat-shocked non-transgenic siblings and transgenic Tg[hsp70:GFP] management teams at 32 hpf, 48 hpf, and 56 hpf when standardized to the world of the attention (Fig 2C) or of the world of the otic vesicle (S4F Fig). Collectively, these information indicated that lowering intracellular Ok+ will increase development of the buds.
Fig 2. Ok+-leak channels improve the expansion of pectoral fin buds.
(A, B) Brightfield photos of a post-heat-shocked 48 hpf non-transgenic embryo (A) or post-heat-shocked 48 hpf hsp70:kcnk5b-GFP transgenic embryo (B). The realm of the fin bud (black-dotted line). (C) Measurements of pectoral fin bud areas of heat-shocked teams of non-transgenic (AB), Tg[hsp70:GFP], Tg[hsp70:kcnk5b-GFP], and Tg[hsp70:kcnk10-GFP]. The non-transgenic and Tg[hsp70:kcnk5b-GFP] embryos are siblings. Every measured bud was standardized to the attention space (red-dotted circles in A,B) in the identical embryo. (D–S) Expression of the indicated morphogens. (D, G, J, M) Lateral views of whole-mount in situs of heat-shock management non-transgenic embryos at 48 hpf for fgf8a (D), fgf10a (G), shha (at 52 hpf) (J), aldh1a2 (M) and dorsal view of aldh1a2 (O) or lateral views of heat-shocked Tg[hsp70:kcnk5b-GFP] embryos at 48 hpf for fgf8a (E), fgf10a (H), shha (at 52 hpf) (Ok), aldh1a2 (N), and dorsal view of aldh1a2 (P). White arrows point out proximal expression of aldh1a2. qRT-PCR measurements from fin buds for fgf8a (F), fgf10a (I), shha (L), and aldh1a2 (Q). Lateral views of in situs of heat-shocked non-transgenic fin bud for bmp4 (R) and msx2b (U) and of heat-shocked Tg[hsp70:kcnk5b-GFP] embryos at 48 hpf for bmp4 (S) and msx2b (V). qRT-PCR measurements from fin buds for bmp4 (T), msx2b (W) the in situ experiments have been repeated 3 or extra time (N ≥ 3). Every in situ repeat accommodates 6–12 embryos per replicate. For the fin bud measurement measurements, we measured 1 fin bud and eye per embryo. We measured at the least 4 embryos per experimental repeat. For the qRT-PCRs, we collected 80 fin bud samples per isolation. We assessed gene expression with 3 or extra isolations. Every isolation was assessed in duplicate or triplicate. Every measured worth is represented as an information level. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values ≥0.05 are designated as “not vital” (NS). The size bars are 0.5 mm (A, B), 100 μm (D–P, R–S,U–T). Numerical information used on this determine are included in S2 Data. hpf, hours submit fertilization.
The rise in pectoral fin bud measurement from Ok+-leak channel overexpression indicated that the developmental gene program was affected. We due to this fact examined the expression of chosen morphogens identified to regulate early bud development at 48 hpf. We noticed that in comparison with heat-shocked management siblings (Fig 2D, 2G, 2J, 2M and 2O), transgenic expression of kcnk5b elevated the expression of fgf8a (Fig 2D and 2E), fgf10a (Fig 2G and 2H), shha (Fig 2J and 2K), and aldh1a2 (Fig 2M–2P). We noticed comparable up-regulation of those genes from qRT-PCR analyses of fin buds (Fig 2F, 2I, 2L and 2Q). As a result of earlier findings present that BMP signaling in vertebrate and invertebrate appendages is affected by Ok+ channels [5,6], we additionally assessed the expression of an vital BMP ligand (bmp4) and its down-stream goal msx2b. We noticed that in comparison with in situ controls (Fig 2R and 2U), kcnk5b elevated the expression of each genes within the in buds (Fig 2S and 2V). We additionally noticed comparable up-regulation of those genes by qRT-PCR (Fig 2T and 2W). Thus, the endogenous decreases in intracellular Ok+ throughout fin bud improvement (Fig 1), the coordinated enhance within the expression of a number of vital morphogens by rising the expression of the Ok+-leak channel kcnk5b (Fig 2D–2W), and the improved development the pectoral fin bud by kcnk5b or kcnk10a overexpression (Fig 2C) all implicated intracellular Ok+ as an integral a part of the fin bud development management, particularly, decreased intracellular Ok+ augmented bud proportional development.
Retinoic acid decreases intracellular Ok+ through an Rcan2-mediated mechanism that scales fin buds
The lower in intracellular Ok+ throughout pectoral fin bud development advised that Ok+ ranges may be conscious of developmental alerts that regulate development. One such sign is RA [30–32]. Subsequently, we examined whether or not RA stimulation influenced intracellular Ok+ ranges in growing fin buds at 32 hpf by FLIM measurements of the KIRIN1 transgene 6 h after its heat-shock induction. In comparison with remedy with the solvent DMSO (Fig 3A and 3C), we noticed decreases in Ok+ after 6 h of remedy with 200 nM RA (Fig 3B and 3C), each in ectoderm and mesenchyme cells (Fig 3D). To check whether or not the adjustments in intracellular Ok+ have been particular to RA, we assessed the impact of thyroid hormone (TH), one other nuclear hormone receptor mechanism. Remedy with TH confirmed no vital variations in Ok+ ranges in comparison with DMSO-treated controls (S5C Fig). These information indicated that RA-mediated signaling is adequate to lower intracellular Ok+ in fin bud tissues and that this impact just isn’t induced by all nuclear hormones.
Fig 3. Retinoic acid decreases intracellular Ok+ in pectoral fin buds through transcriptional activation of the calcineurin inhibitor rcan2.
(A, B) FLIM photos of growing fin buds from embryos handled for six h with DSMO-treated (A) or with 200 nM RA-treated (B) 32 hpf embryos. (C) FLIM measurements from cells in growing buds at 32 hpf of the indicated remedy teams. (D) FLIM measurements of cells within the ectoderm (Ecto) or the mesenchyme (Mesen) of buds at 32 hpf handled both with DMSO or RA. (E) In situ photos of dorsal view (a) and lateral view (b) for kcnk5b expression in 48 hpf embryos. Pectoral fin buds (pfb), ectoderm (e), and mesenchyme (m). (F) Consultant lateral view photos of pectoral fin buds of in situ experiments for kcnk5b expression at 32 hpf (a), 56 hpf (b), and 72 hpf (c). (G) Consultant lateral views of pectoral fin buds after in situ experiments for rcan2 expression at 32 hpf (a), 34 hpf (b), 48 hpf (c), and 56 hpf (d) embryos. (H) qRT-PCR of rcan2 from remoted pectoral fin buds of 48–52 hpf embryos handled with DMSO or 200 nM RA for six h. (I) In situ for rcan2 expression in fin buds of 48 hpf embryos after 6 h DMSO remedy. Ectoderm is indicated by (e), and mesenchyme is indicated by (m). (J) In situ photos for rcan2 expression in fin buds of 48 hpf embryo after 6 h 200 nM RA remedy. Ectoderm (e), mesenchyme (m). (Ok, L) Pectoral fin buds of heat-shocked non-transgenic embryo (Ok) or heat-shocked transgenic hsp70:rcan2-mCherry sibling (L). (M) Measured fin buds have been standardized to the world of an eye fixed in the identical embryo. (N) FLIM measurements of intracellular Ok+ in pectoral fin bud cells from heat-shocked non-transgenic and mCherry controls and transgenic rcan2-mCherry-expressing siblings. (O–R) Consultant fin buds of embryos expressing empty sgRNA vector “EV” (O), rcan2 sgRNA “KO” (P), mCherry mRNA with rcan2 sgRNA “KO+mCherry” (Q), mutated rescue rcan2 mRNA with rcan2 sgRNA “KO+rcan2” (R). (S) Ratios of fin bud areas to eye areas of embryos of the indicated experimental teams. (T) FLIM measurements of intracellular Ok+ within the pectoral fin bud cells in management and CRISPR-Cas9 knockout of rcan2 in embryos. Experiments have been repeated 3 or extra time (N ≥ 3). For FLIM, we measured 2 or 3 places in every tissue of 1 fin bud per embryo. Every measured worth is represented as an information level (C, D, N, T). For the fin bud measurement measurements, we measured 1 fin bud per embryo (M) and a pair of fin buds per embryo (S). For the qRT-PCR experiments, we collected 80 fin bud samples per isolation. Three or extra isolations have been measured. Every isolation was measured in duplicate or triplicate. Every isolation is represented as an information level. Every in situ repeat contained 6–12 embryos per replicate. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values >0.05 are designated as “not vital” (NS). Scale bars symbolize 50 μm (E–G, I–L, O–R). Numerical information used on this determine are included in S3 Data. FLIM, Fluorescence Lifetime Microscopy; hpf, hours submit fertilization; RA, retinoic acid.
RA regulates gene transcription through particular intracellular receptors [33]. Consequently, the relative discount in Ok+ by RA means that RA regulates the transcription of a number of Ok+ channels. As a result of Kcnk5b is concerned in grownup fin scaling [9], we carried out in situ hybridization experiments to find out whether or not kcnk5b is current in rising fin buds of 48 hpf embryos. We noticed expression primarily in distal pharyngeal pouches (Fig 3Ea), and the mesenchyme of the rising fin buds (Fig 3Ea and 3Eb). We subsequently assessed kcnk5b expression throughout fin bud development at 34 hpf, at 56 hpf, and at 72hpf, a time level wherein bud development has already ceased. We noticed kcnk5b expression primarily within the mesenchyme at 32 hpf (Fig 3Fa) and at 56 hpf (Fig 3Fb), however by 72 hpf, the channel expression was tough to detect within the bud (Fig 3Fc). As a result of remedy with RA decreases intracellular Ok+ ranges, we reckoned that RA could also be enhancing kcnk5b transcription. Subsequently, we examined whether or not RA-mediated lower in intracellular Ok+ was associated to an up-regulation of kcnk5b expression. We didn’t observe a rise in RA-treated embryos by qRT-PCR (S5A Fig) regardless of a big enhance in a identified RA-activated gene (S5B Fig) [34].
We beforehand confirmed that inhibition of calcineurin will increase the exercise of Kcnk5b to lower intracellular Ok+ and improve the size of the grownup fin [11]. Since Kcnk5b is current within the embryonic pectoral fin bud (Fig 3E and 3F), we hypothesized that RA could lower intracellular Ok+ by posttranslational regulation of Kcnk5b by the inhibition of calcineurin. RCAN proteins are well-documented in vivo inhibitors of calcineurin [35–37]. From in situ hybridization experiments, we detected expression at 32 hpf (Fig 3Ga), at 34 hpf (Fig 3Gb), and 48 hpf (Fig 3Gc). Nevertheless, rcan2 expression lowered by 56 hpf (Fig 3Gd). As a result of RA remedy decreased intracellular Ok+ and Rcan2 inhibits calcineurin, which subsequently might suppress calcineurin-mediated inhibition of Kcnk5b, we examined whether or not rcan2 expression is altered by RA. We noticed that in comparison with DMSO controls, RA remedy elevated rcan2 expression by qRT-PCR (Fig 3H), and from in situ hybridization experiments, we noticed staining in DMSO-treated buds (Fig 3I) that turned extra intense after RA remedy (Figs 3J and S5D). We additionally noticed comparable phenomena within the grownup zebrafish fins: RA remedy decreased intracellular Ok+ ranges (S5G Fig) and elevated rcan2 transcription (S5H Fig). Moreover, we detected rcan2 transcripts and protein within the blastemas and distal regenerating interray tissue of regenerating fins (S5I–S5K Fig), the place development happens. We didn’t observe protein expression in fin rays instantly after amputation when regenerative development has not but commenced (S5J and S5K Fig).
The power of RA to up-regulate rcan2, which inhibits a phosphatase that suppresses the Ok+-leak channel Kcnk5b (current and scales fin buds and grownup fins [9,11]) led us to hypothesize that Rcan2 could improve proportional development of fin buds. To check whether or not rising Rcan2 enhances pectoral fin bud development, we generated a fish line harboring the hsp70:rcan2-mCherry transgene to overexpress rcan2 by heat-shock induction (S5L–S5O Fig). In comparison with heat-shocked non-transgenic management siblings (Fig 3K and 3M) and overexpressed mCherry (Fig 3M), induction of rcan2 enhanced the expansion space of the fin buds (Fig 3L and 3M). We additionally noticed enhanced development by transgenic overexpression of rcan2 throughout grownup fin regeneration (S5P–S5V Fig). To find out whether or not rcan2-enhanced development correlated with lowered intracellular Ok+, we generated double-transgenic fish that harbored hsp70:rcan2-mCherry and hsp70:KIRIN1 transgenes. After heat-shock induction of the transgenes, we noticed that rcan2 expression decreased intracellular Ok+ ranges in comparison with controls (Fig 3N). Collectively, these information confirmed that induction of rcan2 expression was adequate to boost development within the fin bud and grownup fin in addition to cut back their intracellular Ok+ ranges in vivo.
We then assessed whether or not focusing on rcan2 by CRISPR-Cas9 (S5W–S5Z Fig) affected fin bud measurement and intracellular Ok+. In comparison with management embryos (Fig 3O and 3S), we noticed that rcan2-targeted embryos displayed lowered fin bud development (Fig 3P, 3Q and 3S), which may very well be rescued by overexpression of rcan2 mRNA (Fig 3R and 3S) that harbored mutations in its wobble bases to impair interactions between the overexpressed mRNA and the genome-targeting sgRNA (S5Za and S5Zb Fig). We additionally assessed caudal fins of CRISPR-Cas9-targeted juvenile fish, which additionally displayed shorter fins in comparison with wild sort (S5A and S5B Fig). Once we assessed the impact of focusing on rcan2 on intracellular Ok+ ranges by focusing on rcan2 in Tg[hsp70:KIRIN1] embryos, we noticed that focusing on rcan2 elevated intracellular Ok+ (Fig 3T) and rescue with rcan2 mRNA returned intracellular Ok+ again to regulate ranges (Fig 3T). Collectively, these outcomes from embryonic fin buds and post-metamorphosis caudal fins indicated the involvement of rcan2 in Ok+-channel-mediated scaling of embryonic and grownup appendages.
Rcan2-mediated scaling requires Kcnk5b exercise through its Serine 345
Calcineurin limits fin proportional development by inhibiting Kcnk5b by serine 345 [11]. Since Rcan2 inhibits calcineurin [35–37], and kcnk5b is expressed within the growing pectoral fin buds (Fig 3E and 3F), we first assessed the significance of kcnk5b in fin bud development by CRISPR-targeting kcnk5b (S6A–S6G Fig). In comparison with controls (Fig 4A and 4E), fin buds of kcnk5b-targeted embryos have been contracted by 48 hpf (Fig 4B and 4E). This phenotype was rescued by overexpression of kcnk5b mRNA (Fig 4D and 4E) that was mutated to impair interplay with the focusing on sgRNA (S6H–S6J Fig). Once we overexpressed rcan2 in kcnk5b-targeted embryos, we noticed that whereas rcan2 overexpression in non-targeted embryos enhanced fin bud development (Fig 4F, 4G and 4I), rcan2 required kcnk5b to boost development (Fig 4H and 4I). We beforehand confirmed that calcineurin inhibits Kcnk5b through serine 345 within the cytoplasmic tail of the channel [11], so we examined whether or not the kcnk5bS345A (the mutant with the bottom Ok+-leak exercise) blunts rcan2 enhancement of fin bud development in vivo. In comparison with management embryos overexpressing rcan2 with the wild-type channel (kcnk5bS345) (Fig 4J and 4L), the calcineurin-dephosphorylated mimic kcnk5bS345A channel lowered rcan2-mediated fin bud development (Fig 4K and 4L). Collectively, these information indicated that rcan2-enhanced development requires Kcnk5b and that the Kcnk5b channel that mimics calcineurin-mediated inhibition of the channel prevents the improved development attributable to overexpression of the endogenous calcineurin inhibitor rcan2.
Fig 4. Rcan2-mediated lower in intracellular Ok+ includes Kcnk5b.
(A–D) Consultant 48 hpf embryos and enlarged panels of fin buds of management Cas9 and empty sgRNA vector (EV) embryo (A), CRISPR-targeted kcnk5b embryo (B), CRISPR-targeted kcnk5b embryo overexpressing GFP (C), CRISPR-targeted kcnk5b embryo overexpressing kcnk5b*-GFP that harbors altered wobble bases to impair interplay with focusing on sgRNA (D). (E) Fin bud-to-eye space ratios of the indicated genotypes. (F–I) Consultant 48 hpf embryos and enlarged panels of fin buds of management Cas9 and empty sgRNA vector (EV) non-Tg embryo (F), rcan2-mCherry-expressing embryo and enlarged panel of the fin bud (G) CRISPR-targeted kcnk5b embryo overexpressing rcan2-mCherry (H). (I) Fin bud-to-eye space ratios of the indicated genotypes. (J, Ok) Consultant 48 hpf embryos and enlarged panel of the fin buds expressing wild-type kcnk5bS345-GFP and rcan2-mCherry (J), or expressing calcineurin-dephosphorylated mimic kcnk5bS345A-GFP and rcan2-mCherry (Ok). (L) Fin bud-to-eye ratios present that rcan2-mediated enlargement of fin buds is impaired by kcnk5bS345A mutant. Experiments have been repeated 3 or extra time (N ≥ 3). For the fin bud measurement measurements, we measured 1 fin bud (E, I) or 2 fin buds (L) per embryo at 48 hpf and never at a later time factors to keep away from incorporating measurements of the finfold development that begin round 56 hpf. Every measured worth is represented as an information level. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values >0.05 are designated as “not vital” (NS). Scale bars equal 100 μm (A–D, F–H, J, Ok). Numerical information used on this determine are included in S4 Data. hpf, hours submit fertilization.
Kcnk5b-enhanced development includes cell depolarization
Kcnk5b is a two-pore Ok+ leak channel whose exercise alters {the electrical} membrane potential on the plasma membrane of cells. The lower in intracellular Ok+ throughout fin bud development (Fig 1) and the significance of Kcnk5b for development (Fig 4) counsel there are vital Ok+-associated adjustments in membrane potential throughout fin bud improvement. To find out whether or not membrane potential alters throughout fin bud improvement, we used DiSBAC2(3), a dye that will increase its fluorescence because it enters cells when channels open to depolarize the cells [38]. We assessed DiSBAC2(3) fluorescence utilizing time-correlated single photon counting photodetectors (identical used for FLIM) that rely the emitted photons per pixel in a confocal aircraft. From measurements at completely different places in confocal planes of a number of fin buds, we noticed the least quantity of depolarization at 32 hpf within the ectoderm (Fig 5A) and the mesenchyme (Fig 5B). Afterward, depolarization elevated by 42 hpf with common highest ranges at 48 hpf and 56 hpf (Fig 5A and 5B). Whereas the averages elevated, these averages represented a broad distribution of depolarization ranges significantly at 48 hpf and 56 hpf. To visualise the distribution of relative variations in membrane potential on the chosen time factors, we depicted the photon-count info of every pixel in mid-level confocal planes of the fin buds utilizing a rainbow coloration scale wherein pink represented the best ranges of DiSBAC2(3) fluorescence (depolarization) and inexperienced to blue depicted decrease ranges. We noticed that 32 hpf constantly confirmed the bottom ranges of depolarization (Fig 5C’ and 5C”). DiSBAC2(3) fluorescence incrementally elevated all through the fin buds at 42 hpf (Fig 5D’), 48 hpf (Fig 5E’), and 56 hpf (Fig 5F’). As indicated by the variance in our measurements, we additionally noticed fin buds at these time factors that displayed decrease ranges of fluorescence or variegated patterns excessive and low fluorescence (Fig 5D”, 5E” and 5F”). We posit that the noticed variation in relative membrane potential values after 32 hpf could stem from oscillations across the common membrane potential, though we can’t rule out variations in dye penetration. In any case, the mixed measurement information present a collective enhance in depolarization.
Fig 5. Kcnk5b enhanced depolarization is required for enhanced fin bud development.
(A, B) DiSBAC2(3) fluorescence measurements of the ectoderm (A) and mesenchyme cells (B) of growing pectoral fin buds at 32, 42, 48, and 56 hpf. (C–F) Confocal photos of growing fin buds displaying the intensities fluorescence as photon counts per pixel at 32 hpf (C), 42 hpf (D), 48 hpf (E), and 56 hpf (F). Colours symbolize the vary of counted photons per pixel. Blue representing the bottom stage of counted photons, and pink representing their highest counts (as much as 750 photons or extra). Pictures representing excessive photon rely (C’, D’, E’, F’) and low photon rely (C”, D”, E”, F”). The entire publicity vary was set at 1,500 counts (1.5). (G) Distribution of counted photons from DiSBAC2(3) within the confocal aircraft of the consultant fin bud at 32 hpf. (H) Distribution of counted photons from DiSBAC2(3) within the confocal aircraft of the consultant fin bud at 32 hpf. (I) Evaluation of DiSBAC2(3) fluorescence depth as counted photons for fin buds expressing mCherry or kcnk5b-mCherry. (J) Evaluation of DiSBAC2(3) fluorescence depth of mCherry-expressing or kcnk5b-mCherry fin buds at 32 hpf after treating for 4 h as indicated: ethanol (Et), DMSO, 10 μm vinpocetine (vin), 40 μm dibucaine (dib). (Ok) Consultant picture of pectoral fin bud of non-transgenic 48 hpf AB fish after warmth shock at 32 hpf and begin of remedy at 36 hpf with the drug solvents Ethanol (Et) and DMSO. (L) Consultant picture of pectoral fin buds of 48 hpf AB fish warmth shocked at 32 hpf and begin of remedy with 10 μm Vinpocetine (Vin) and 40 μm Dibucane (Dib) at 36 hpf. (M) Consultant picture of pectoral fin buds of 48 hpf transgenic Tg[hsp70:kcnk5b-GFP] after warmth shock at 32 hpf and begin of remedy with EtOH and DMSO at 36 hpf. (N) Consultant picture of pectoral fin buds of 48 hpf transgenic Tg[hsp70:kcnk5b-GFP] after warmth shock at 32 hpf and begin of remedy with 10 μm Vin and 40 μm Dib at 36 hpf. (O) Evaluation of pectoral fin bud development at 48 hpf expressing both AB and kcnk5b-GFP within the indicated remedy teams. Experiments have been repeated 3 or extra time (N ≥ 3). Every repeat contained 6 or extra embryos; one fin bud was measured per embryo. For the DiSBAC2(3) fluorescence measurements, 6 impartial factors have been measured from completely different 4 places in every fin bud, distal, anterior, posterior, and proximal, after which averaged to symbolize an information level (A, B, I, J). For the fin bud measurement measurements, we measured 1 fin bud and eye per embryo. We measured at the least 15 embryos per repeat, and every measurement is 1 information level (O). P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values >0.05 are designated as “not vital” (NS). Scale bars equal 100 μm. Numerical information used on this determine are included in S5 Data. hpf, hours submit fertilization.
To evaluate the impact of Kcnk5b-mediated lower in intercellular Ok+ on the membrane potential, we overexpressed kcnk5b-mCherry or mCherry at 32 hpf, the time level with the bottom depolarization ranges (Fig 5A–5C), after which measured membrane potential. We noticed that in comparison with mCherry-expressing fin buds (Fig 5G and 5I), overexpression of kcnk5b-mCherry considerably elevated depolarization (Fig 5H and 5I). These outcomes indicated that Kcnk5b exercise promotes depolarization.
The noticed enhance in depolarization attributable to kcnk5b, a leak channel that ought to promote hyperpolarization, advised that different channels whose exercise causes depolarization, corresponding to Na+ channels, have been concerned. We due to this fact examined whether or not Kcnk5b-mediated depolarization required Na+-channel exercise. We expressed both management mCherry or kcnk5b-mCherry after which handled these fish with the Na+-channel inhibitors Vinpocetine (a broad voltage-gated sodium channel inhibitor, together with the TTX-insensitive channels) and Dibucane (broad sodium channel inhibition) (S7E–S7J Fig). We noticed that inhibition of Na+ channels decreased depolarization within the fin bud in addition to prevented Kcnk5b-induced enhance in depolarization (Fig 5J). To check whether or not Na+-channel-mediated depolarization is required for kcnk5b-enchanced development, we overexpressed kcnk5b-GFP and assess the impact of impairing depolarization utilizing the Na+-channel inhibitors. In comparison with the improved fin bud sizes of control-treated kcnk5b-GFP-transgenic fish (Fig 5M and 5O), remedy of kcnk5b-GFP-expressing fish with the Na+-channel inhibitors impaired Kcnk5b-induced fin bud development (Fig 5N and 5O). Collectively, these information indicated that Kcnk5b exercise promotes depolarization through Na+ channels and that Kcnk5b-induce depolarization is required for Kcnk5b-enhanced development.
IP3R-mediated Ca2+ launch is required for Kcnk5b-induced shha expression and fin bud scaling
Our observations that Ok+-leak channels enhance the expression of vital morphogens advised that the channels are doing so by a number of signaling mechanisms. Because the intracellular accumulation of second messengers is concerned in lots of signaling mechanisms, we assessed whether or not Kcnk5b exercise alters the degrees of explicit second messengers. We used HEK293 cells for an preliminary evaluation, as a result of we beforehand noticed that Kcnk5b induces SHH in these cells [11], a morphogen vital for the event of early fin/limb buds [39]. To find out whether or not cAMP or cGMP ranges change in response to Kcnk5b, we used FLIM with a longtime FRET-based sensor for every [40,41]. In comparison with controls teams of unstimulated cells or cells stimulated with forskolin that produces cAMP (S8A Fig) or cells stimulated with SNAP to provide cGMP (S8B Fig), Kcnk5b didn’t considerably alter the intracellular ranges of cAMP or cGMP (S8A and S8B Fig). We then assessed intracellular Ca2+ utilizing the GCaMP6s sensor [42], and we noticed that in comparison with cells transfected with management plasmid (Fig 6A and 6D), transfection with Kcnk5b led to vital will increase in Ca2+ (Fig 6B and 6D). An inactive mutant model of the Kcnk5b channel, Kcnk5bMut (S8C Fig), additionally didn’t enhance GCaMP6s exercise (Fig 6C and 6D). To additional assess the connection between Kcnk5b and intracellular Ca2+, we transfected cells with GCaMP6s and both with Kcnk5b-mCherry or with mCherry. We subsequently cultured the cells for twenty-four h, FACS sorted each transfection teams for mCherry fluorescence, after which plated every group of sorted cells at 100% confluency (Fig 6Ea and 6Eb). mCherry-transfected cells that didn’t constantly show GCaMP6s fluorescence (Fig 6Ea, 6Ec, 6Ee, 6Eg arrow versus asterisk and 6F), whereas Kcnk5b-mCherry+ cells at all times confirmed excessive GCaMP6s exercise (Fig 6Eb, 6Ed, 6Ef, 6Eh asterisks and 6F). We noticed this phenomenon 100% of the time in all confocal photos (Fig 6G). Collectively, these outcomes point out that Kcnk5b exercise promotes an increase in intracellular Ca2+.
Fig 6. Kcnk5b exercise induces IP3R-mediated Ca2+ launch from the ER.
(A) GCaMP6s fluorescence in HEK293 cells. (B) GCaMP6s fluorescence in HEK293 cells expressing Kcnk5-mCherry. (C) GCaMP6s fluorescence in HEK293 cells expressing Kcnk5BSM-mCherry. (D) Fluorescence measurements of indicated teams. (E) Consultant confocal photos of mCherry-transfected, GCaMP6s-transfected cells exhibiting brightfield with merged fluorescence from mCherry and GCaMP6s (a) or mCherry (c) or GCaMP6s (e) or merged mCherry-GCaMP6s (g), and Kcnk5b-mCherry-transfected, GCaMP6s-transfected cells: brightfield with merged fluorescence from mCherry and GCaMP6s (b) or Kcnk5b-mCherry (d) or GCaMP6s (f) or merged mCherry-GCaMP6s (h). (F) Measurements of GCaMP6s fluorescence depth from the indicated experimental transfection teams. (G) Percents of mCherry-GCaMP6s-double optimistic over the full variety of mCherry-positive cells in every group relate the frequency of GCaMP6s-positive cells within the mCherry or Kcnk5b-mCherry teams. (H) GCaMP6s fluorescence within the pectoral fin buds of transgenic fish harboring each Tg[Cca.actb:GCaMP6s] and Tg[hsp70:kcnk5b-mCherry]. (I) GCaMP6s fluorescence in embryos harboring the secure transgenic fish line Tg[Cca.actb:GCaMP6s] that mosaically categorical patches of mCherry or kcnk5b-mCherry (designated mCherry+) for the ectoderm (a) and mesenchyme (b). (J) Diagram of IP3R inhibition by 2-APB. (Ok) qRT-PCR of SHH expression in HEK293 cells transfected both with GFP or Kcnk5b-GFP after 20-h remedy with 2-APB on the indicated concentrations. (L) Evaluation of pectoral fin bud measurement at 48 hpf after 4 h of remedy with 13 μm 2-APB. (M) Expression of shha in pectoral fin buds of heat-shocked non-transgenic sibling embryos after 4-h remedy with DMSO (a) or 13 μm 2-APB (b), of heat-shocked transgenic Tg[hsp70:kcnk5b-mCherry] siblings after 4-h remedy with DMSO (c) or 13 μm 2-APB (d). (N) Pixel space of in situ staining of shha within the indicated remedy teams. (O) qRT-PCR of shha expression in remoted fin buds of the indicated teams. Experiments have been repeated 3 or extra time (N ≥ 3). For cell tradition experiments, every repeat contained duplicate or triplicate wells, and 10 or extra cells have been measured per nicely. Every information level represents 1 cell (D, F, G). For fin bud fluorescence measurements, we measured 2 or 3 places in every tissue of 1 fin bud per embryo (H, I). For fin bud space measurements, we measured the world of 1 fin bud and eye per embryo. We measured at the least 4 embryos per repeat (L). For the qRT-PCR experiments, we collected 80 fin bud samples per isolation. Three or extra isolations have been measured. Every isolation was measured in duplicate or triplicate. Every measured worth is represented as an information level. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values >0.05 are designated as “not vital” (NS). Scale bars equal 100 μm (A–C), 10 μm (E), 0.5 μm (M). Numerical information used on this determine are included in S6 Data. ER, endoplasmic reticulum; hpf, hours submit fertilization.
To find out whether or not Kcnk5b has the identical impact on intercellular Ca2+ ranges in vivo, we generated double-transgenic fish that harbored Tg[hsp70:kcnk5b-mCherry] and a CGaMP6s Ca2+ reporter below the management of the β-actin promoter Tg[Cca.actb:GCaMP6s] [43]. We assessed intracellular Ca2+ within the fin buds of heat-shocked transgenic kcnk5b-mCherry embryos at 48 hpf and their non-transgenic siblings as controls. We noticed that Ca2+ ranges have been increased in kcnk5b-mCherry in comparison with non-transgenic embryos (Fig 6H). We noticed comparable outcomes after we assessed mosaic embryos harboring the secure transgenic fish line Tg[Cca.actb:GCaMP6s] that mosaically categorical patches of mCherry or kcnk5b-mCherry (Fig 6Ia and 6Ib). Collectively, these information indicated that Kcnk5b usually promotes the rise in intracellular Ca2+ ranges in vivo, and so they advised Ca2+ mediates the growth-inducing impact of Kcnk5b. Will increase in Ca2+ attributable to the lower in intracellular Ok+ might happen from extracellular sources through Ca2+ channels within the plasma membrane and/or from intracellular sources such because the endoplasmic reticulum (ER) [44]. To find out the significance of those 2 sources, we examined whether or not inhibiting launch of Ca2+ from every supply impaired Kcnk5b-induced transcription of SHH in HEK293 cells. After inhibiting T- and L-type Ca2+ channels on the plasma membrane (S8D Fig), we didn’t detect a big impact on Kcnk5b-induced SHH transcription (S8E and S8F Fig). Nevertheless, after inhibiting IP3R-mediated launch of Ca2+ from the ER (Fig 6J), we noticed decreased Kcnk5b-induced SHH transcription dose-dependently (Fig 6K).
To find out whether or not IP3R-mediated Ca2+ launch was required for the Kcnk5b-enhanced development of pectoral fin buds, we inhibited IP3R exercise for 4 h and assessed bud development in heat-shocked non-transgenic and transgenic Tg[hsp70:kcnk5b-GFP] siblings. We noticed that the IP3R inhibitor blocked the expansion phenotype attributable to expression of kcnk5b at 48 hpf (Fig 6L). This lower in development was related to the impairment of Kcnk5b-induced enhance of shha expression domains (Fig 6M and 6N) and qRT-PCR (Fig 6O).
The significance of Ca2+ launch for shha expression and fin bud development indicated that a number of Ca2+-dependent kinases have been concerned. Subsequently, we examined which of the Ca2+-activated CaM kinases have been required for Kcnk5b-enhanced expression of SHH in HEK293 cells. Whereas we noticed no vital results by inhibiting CaMKII and CaMKIV (S9A Fig), we did observe that inhibition of CaMKK impaired Kcnk5b-enhanced SHH expression (Fig 7A). We then examined whether or not inhibiting CaMKK within the fin buds has comparable results on shha expression in vivo. We noticed that inhibiting CaMKK decreased the expression of shha by in situ (Fig 7B and 7C) and by qRT-PCR (Fig 7D). Inhibiting CaMKK additionally impaired Kcnk5b-enhanced development (Fig 7E). Moreover, overexpressing the human CaMKK2 (Fig 7F) or the zebrafish camkk1b (Fig 7G) was adequate to extend SHH expression in HEK cells. Collectively, these outcomes indicated that Ca2+ is required for Kcnk5b-induced transcription of shha and that CaMKK is a vital a part of Ca2+ regulation of shha expression and pectoral fin bud development.
Fig 7. Kcnk5b requires CaMKK for development and SHH/shha expression.
(A) SHH expression in HEK293 cells transfected with GFP or Kcnk5b-GFP after 20-h remedy on the indicated concentrations of the CaMKK inhibitor STO-609 on the indicated concentrations. (B) Expression of shha in pectoral fin buds from heat-shocked non-transgenic siblings after 6-h remedy with DMSO (a) or 24 μm STO-609 (b) and from transgenic Tg[hsp70:kcnk5b-mCherry] siblings after remedy with DMSO (c) or STO-609 (d). (C) The ratios of every in situ staining space of shha to the world of its corresponding fin bud for the indicated teams. (D) qRT-PCR of shha expression in remoted fin buds within the indicated management and experimental teams. (E) Graphed evaluation of fin-bud-area-to-eye-area measurement ratios. (F, G) Distinction in SHH expression in HEK293 cells transfected with mCherry or human CaMKK2-mCherry (F) or camkk1b-mCherry (G). Experiments have been repeated 3 or extra time (N ≥ 3). For cell qRT-PCR experiments, every RNA isolation per nicely was measured in duplicate or triplicate. Every measured worth is represented as an information level (A, F, G). For fin bud fluorescence measurements, we measured 2 or 3 places in every tissue of 1 fin bud per embryo. For fin bud space measurements, we measured the world of 1 fin bud and eye per embryo. We measured at the least 4 embryos per repeat (E). For fin bud qRT-PCR experiments, we collected 80 fin bud samples per isolation. Three or extra isolations have been measured. Every isolation was measured in duplicate or triplicate samples (D). Every measured worth is represented as an information level. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values >0.05 are designated as “not vital” (NS). Scale bars equal 0.5 μm (B). Numerical information used on this determine are included in S7 Data.
Dialogue
Ok+-channel exercise as a mechanism for regulating development of vertebrate fin/limb buds
The fish mutants longfin, one other lengthy fin, kcn13j, and rapunzel exhibit that rising the expression or the posttranslational exercise of a Ok+ channel (Kcnh2a, Kcn13j, Kcnk5b, or Kcc4a) induces allometric development of grownup fin [9,10,12,13]. The observations that completely different Ok+ channels improve development assist the conclusion that adjustments in intracellular Ok+ is what alters scaling. Whereas Kcnk5b and different Ok+-leak channels cut back intracellular Ok+ ranges [45], Kcn13j Ok+-rectifying channels facilitate Ok+ entry into the cells [46]. A proof for why channels that launch Ok+ and a channel that restores intracellular Ok+ each promote development is present proof that implies that Ok+ leak channels act immediately in tissues of the fin, whereas the Ok+-rectifying channel acts within the dermomyotome of somites to have an effect on the later development of grownup fins [13]. The worth of our in vivo FLIM measurements of intracellular Ok+ is that they supply direct proof of a lower in relative intracellular Ok+ ranges in pectoral fin bud tissues throughout outgrowth (Fig 1D–1I), which helps the conclusion that development of the pectoral fin buds includes an general discount of intracellular Ok+ in bud tissues. This conclusion can be supported by a number of observations: (1) the improved proportions from transgenic overexpression of various Ok+-leak channels (Fig 2); (2) the expression of Kcnk5b throughout the fin bud outgrowth and its down regulation as fin bud improvement ceases (Fig 3E and 3F); (3) the smaller fin bud measurement in CRISPR-targeted kcnk5b embryos (Fig 4A–4E); (4) the lower in intracellular Ok+ by RA remedy (Fig 3A–3D); (5) the improved bud measurement (Fig 3K–3M); (6) the decreased Ok+ (Fig 3N) from rcan2 overexpression (which boosts development); (7) rcan2-induced enhanced development requires kcnk5b (Fig 4); and (8) CRISPR focusing on of rcan2 each decreased bud measurement and elevated Ok+ (Fig 3S and 3T). We additionally noticed rcan2 expression and its comparable results on scaling and regulating intracellular Ok+ in grownup caudal fins.
The endogenous expression profiles of kcnk5b (Fig 3E and 3F) and rcan2 (Fig 3G) present that each are current throughout the development of the fin bud. Rcan2 can be current within the grownup fin blastema the place it could possibly regulate Kcnk5b and improve development (S5I–S5K Fig). Whereas the amassed information point out that the expression of rcan2 coincides with kcnk5b in fin bud and fin development, between 32 hpf and 34 hpf, we noticed a rise in intracellular Ok+ (Fig 1E) regardless of the presence of rcan2 and kcnk5b. We posit that there’s one other mechanism regulating Kcnk5b or one other Ok+ channel or Ok+ pumps that facilitates the rise in Ok+. We suggest that the rise in Ok+ between 32 hpf and 34 hpf (Fig 1E) explains the pause in bud development throughout this era (Fig 1F). We suspect the kcnk5b expression at 56 hpf (Fig 3Fb) implies that this channel is concerned in different phenomena corresponding to early tissue differentiation, whereas the decreased rcan2 at 56 hpf (Fig 3Gd) signifies that the RA-Rcan2 mechanism now not promotes Kcnk5b exercise and, consequently, intracellular Ok+ ranges enhance.
Zebrafish fin bud improvement happens inside a 24-h interval, by which period it has transitioned into finfold development, which initiates out of the distal-most area of the bud to kind the pectoral fins of the larval fish. As a result of this transition between the conserved vertebrate fin/limb bud developmental stage, we primarily centered our characterizations to 48 hpf as a way to outline the contribution of intracellular Ok+ regulation within the scaling of the conserved embryonic bud construction. Thus, the adjustments in fin bud measurement attributable to knockout or overexpression of kcnk5b or rcan2 usually are not giant, however they’re biologically vital. We attribute the variance that we noticed on the measured time factors to pure variations in development throughout the slender window of pectoral fin bud improvement.
There are a number of signaling facilities/zones throughout the growing fin buds [22], but we noticed few distinctions within the distribution of relative intracellular Ok+ ranges within the early buds (Figs 1G–1I and S3). We suggest this is because of hole junctions. Hole junctions are vital for limb bud improvement and fin development [47–49], and the shared distribution of comparatively comparable Ok+ ranges through hole junctions can clarify the coordinated regulation of every morphogen all through the bud. We noticed that the first distinction in relative ranges of intracellular Ok+ exists between the mesenchyme and the ectoderm (Fig 1D and 1E). These variations could also be because of an ECM barrier between the tissues, fewer gap-junctional connections and/or variations in molecular alerts that affect Ok+ channel expression/exercise. As a result of cells use intracellular Ok+ to regulate resting membrane potential [45,50], any variations in Ok+ ranges between these tissues counsel that they’ve distinct electrophysiological properties.
Regarding the in vivo electrophysiology, since Ok+-leak channels usually end result within the outward move of Ok+ ions, this could hyperpolarize the membrane potential because the cytoplasmic aspect turns into extra negatively charged from the outflow of positive-charged ions. Nevertheless, we noticed that depolarization will increase throughout outgrowth (Fig 5A–5F) whereas intracellular Ok+ decreases (Fig 1E). Moreover, after we overexpress kcnk5b within the fin buds, we elevated depolarization (Fig 5G–5I). We suggest that the elevated Ok+-channel exercise first hyperpolarizes the membrane potential, and this enhance in membrane potential instantly prompts Na+ channels to depolarize the cells. This speculation suits with our findings that we impair the rise in depolarization (Fig 5J) in addition to Kcnk5b-enhanced development with Na+ channel inhibitors (Fig 5K–5O). We posit that the rise in intracellular Ok+ by 56 hpf immediately facilitates the noticed depolarization from the buildup of Ok+ on the membrane.
Integration of Kcnk5b right into a mechanism of fin bud improvement and development
The coordinated adjustments in intracellular Ok+ (Fig 1) argue that there are widespread mechanisms regulating Ok+ dynamics. We recognized retinoic acid-regulated signaling as one such mechanism (Fig 3), since RA was adequate to lower intracellular Ok+ within the ectoderm and mesenchyme (Fig 3A–3D). We beforehand confirmed that calcineurin inhibition induces allometric development of grownup fins [51] by rising Kcnk5b exercise [11]. Our findings that kcnk5b and the endogenous calcineurin inhibitor regulator of calcineurin 2 (rcan2) are current in pectoral fin buds (Fig 3E–3G) and that each are vital in figuring out the dimensions of the fin buds (Figs 3 and 4) together with our discovering that RA will increase rcan2 expression (Fig 3H–3J) counsel a mechanism that includes calcineurin-mediated antagonism of Kcnk5b exercise to scale the buds (Figs 3 and 4). Our outcomes assist this mannequin: (1) overexpression of the calcineurin inhibitor rcan2 decreased intracellular Ok+ and will increase development of fin buds (Fig 3M and 3N) and of grownup fins (S5AA and S5BB); (2) overexpression of kcnk5b (which decreases intracellular Ok+) enhanced the developmental transcription and development of the fin buds (Fig 2B–2W) and of grownup fins [11]; and (3) a discount in Kcnk5b exercise by knockout or by level mutation impairs rcan2-induced bud development (Fig 4F–4L).
The expression patterns of rcan2 and kcnk5b have been primarily within the mesenchyme of the buds (Fig 3E–3G). We interpreted these findings to imply that intracellular Ok+ is increased within the ectoderm due to very low rcan2 and kcnk5b ranges or due to the absence of those 2 genes and the presence of one other RA-regulated Ok+ channel within the ectoderm. Variations within the expression of Ok+ channels between the ectoderm and the mesenchyme can clarify their variations in Ok+ ranges (Fig 1D and 1E). RA does regulate the expression different morphogens and development components in fin/limb buds, so the linked adjustments in intracellular Ok+ between the ectoderm and mesenchyme will be coupled by different RA-mediated actions that have an effect on the exercise or expression of different Ok+-channels within the ectoderm.
The coordinated regulation of intracellular Ok+ had some specificity, since RA did regulate intracellular Ok+ whereas a distinct nuclear hormone (thyroid hormone) didn’t have any impact (S5C Fig). We interpret these outcomes to imply that TH3 signaling doesn’t immediately regulate the Kcnk5b-mediated development mechanism though thyroid hormones in different organic contexts can induce Rcan2 and/or promote development [52,53]. We suggest that any thyroid-mediated development happens through one other molecular mechanism and never this electrophysiological one. Alternatively, there are endogenous components current that restrict the results of this hormone, since it could possibly promote metamorphosis [54], which wants happen later because the fish ends its larval stage [55]. One other chance is that in early embryonic fin bud and homeostatic development of the grownup fin, the binding websites that thyroid nuclear hormone receptors require are both not accessible or are absent from the regulatory areas of the zebrafish rcan2. Contemplating the significance of thyroid hormone-mediated expression of rcan2 within the differentiation of osteoblasts [56], and contemplating that development of the fin bud and regenerating fin wants cell proliferation earlier than tissue differentiation, our findings could spotlight one among these potentialities.
Ok+-leak channels scale utilizing Ca2+
An vital query is how do Ok+-leak channels scale fin buds. A part of the reply includes IP3R-induced Ca2+ launch from the ER. Nevertheless, our earlier findings confirmed that inhibition of the Ca2+-dependent phosphatase calcineurin will increase the exercise of Kcnk5b [11]. Whereas this discovering seems incongruent with our present discovering that IP3R-mediated enhance in intracellular Ca2+ is required for development, the noticed adjustments in SHH expression from our IP3R inhibition experiments (Fig 6K) supply a proof: we noticed that milder inhibition of IP3R (Fig 6K, 30 μm 2-APB) enhances Kcnk5b-induced expression of SHH, which we posit as lowering the pool of Ca2+ wanted for calcineurin’s inhibition of Kcnk5b, whereas higher IP3R inhibition decreases SHH (Fig 6K, ≥105 μm 2-APB, and 6M–6O) by impairing different Ca2+-dependent enzymes wanted for SHH expression, corresponding to CaMKK (Fig 7).
Our findings are according to different proof that time to the significance of Ca2+ within the development of appendages. The L-type Ca2+ channel Cav1.2 may cause syndactyly, wherein the bones of the digits improperly fuse, when mutations trigger this channel to remain open longer and enhance Ca2+ within the sarcoplasmic reticulum (muscle ER) [57]. Conversely, knock-out of Cav1.2 within the limb mesenchyme results in shorter limbs because of impaired skeletal improvement [58]. In Drosophila wing discs, disruption of proteins that preserve ER Ca2+ shops—such because the Serca2 Ca2+ pump, the Orai Ca2+ channel within the plasma membrane, the Best2 Cl– channel within the ER membrane, or Stim, a scaffold protein that colocalizes Orai and Best2—results in mispatterned, stunted grownup wings [59].
The ER has the most important intracellular Ca2+ retailer, and the discharge of Ca2+ from the ER into the cytoplasm and energetic pumping of Ca2+ again into the ER happens at regulated frequencies to generate oscillating cytoplasmic waves [60]. Ca2+ oscillations coordinate mesenchyme cell motion within the growing buds of feathers [61], and an analogous phenomenon could regulate the expansion of the fin buds [12,62]. Our commentary that Ok+ is shared between cells (Fig 1K–1M) prompts questions on which ions are concerned within the coordinated management of genes and allometric development (Fig 2). We suggest that the sustained transcription that’s wanted for extended allometric development includes a sustained stimulus. Primarily based on our observations, the lower in intracellular Ok+ stays comparatively fixed because the fin bud grows (Fig 1E and 1F). Decreases in Ok+ might enhance intracellular Ca2+ by rising the amplitude or the length of Ca2+ launch from the ER. We posit that it enhances length, since CaMKK exercise is required, and this enzyme wants sustained durations of Ca2+ that obtain its two-step activation course of: Ca2+ have to be current lengthy sufficient to work together with calmodulin after which permit the Ca2+-calmodulin complicated to activate CaMKK. We conclude that Ok+ is an overarching long-term sign that adjusts IP3R-mediated Ca2+ launch to control development.
Intracellular Ok+ in coordinated regulation of morphogens throughout improvement
An vital query is why use Ok+ channels to scale constructions. One chance is that particular Ok+ channels have interactions with particular growth-regulating receptors. The Thromboxane receptor interacts with the Ok+ miniK channel to control the channel’s exercise [63]. Trimeric GPCRs and different membrane-associated signaling molecules work together with channels to affect channel operate [64–66]. Nevertheless, we at present imagine that particular channel–receptor interactions don’t clarify our observations, as a result of the same allometric development phenotypes will be induced by completely different Ok+ channels (Figs 2C and S4D–S4F) [9–12] that possible don’t work together with the identical growth-promoting receptors. A second chance is that altering intracellular Ok+ ranges alters the electrophysiology of cells to advertise development. Adjustments in intracellular Ok+ are identified to change the electrophysiological properties of cells [45], and such adjustments might alter the actions of pro-growth transmembrane receptors or membrane-associated signaling cascade parts with out direct interactions. A associated chance is that intracellular Ok+ is distributed all through the cytoplasm, so adjustments in intracellular Ok+ might affect components past the plasma membrane, such because the IP3R. Adjustments in Ok+ might alter actions of different transduction cascade parts by influencing ionic interplay with charged amino acids in proteins or between them. Such a Ok+-mediated regulatory mechanism wouldn’t essentially be an on-off change, however may very well be an amplifier that augments the actions of signaling parts that improve the expression of current developmental alerts (Fig 2D–2W). The ionic “amplifier” mechanism suits with the commentary that CRISPR-targeting of kcnk5b didn’t stop development of the fin buds; as a substitute, it simply lowered their proportions (Fig 4A–4E).
Ok+ channels within the broader context of improvement and illness
In a broader context of improvement, there are a number of findings that hyperlink completely different Ok+ outward-flow channels to human syndromes that harbor limb defects. The voltage-gated KCNH1 (Kv10.1), the two-pore channel KCNK4 (TRAAK/TREK) and the small-conductance Ca2+-activated KCNN3 (SK3/KCa2.3) are all Ok+ channels whose enhance in actions can result in hypoplasia/aplasia of the distal phalanges, in addition to result in alterations in cranial-facial options and neuropathies [8,67–70]. Conversely, mutations that impair KCNK9 exercise produce bilateral hand contractures and talipes equinovarus ft [71,72]. Comparability between these findings and our findings exhibits the range within the physiological actions of Ok+ channels that lower intracellular Ok+. Whereas the completely different phenotypes can manifest from development defects, we suspect that the variations are because of tissue-specific actions, since even the identical channel in numerous cells can facilitate the transcription of various genes [11].
Along with channels concerned in outward Ok+ move, inwardly rectifying Ok+ channels are vital for physiology. Whereas many discoveries hyperlink their significance to behavioral phenomena [73], mutations in KCNJ6 (GIRK2) or KCNJ13 (Kir7.1) end in extreme cranial-facial malformations together with mental disabilities [74,75]. Nevertheless, it’s unclear how most of the detrimental phenotypes are because of KCNKJ6’s dysfunctional Ok+ move, because the solely characterised mutation that causes the lack of Ok+ selectivity, additionally beneficial properties Ca2+ permeability [76]. With reference to development, elevated expression of Kcnj2 is linked to hypoplasia of distal digit constructions [77], and elevated exercise of one other inward rectifying channel Kcnj13 has additionally been linked to enhanced development of the grownup fins, though it seems to provoke this defect through its embryonic actions [13].
Ok+ channels are additionally linked to tumor formation and most cancers. A number of completely different cancers harbor up-regulated expression and/or exercise of Ok+-leak channels. Expression and exercise of KCNK5 is up-regulated in some breast most cancers cell strains. Signaling from Estrogen Receptor-α (ERα) has been discovered to advertise breast most cancers [78]. ERα signaling can up-regulate the expression of KCNK5, and blocking KCNK5 exercise impairs the cell proliferation attributable to activated ERα [79]. KCNK9 expression can be elevated in quite a lot of breast most cancers tumors, and experimentally overexpressing this channel promotes tumor formation in vivo [80]. A hyperlink between KCNN4 up-regulation and cell proliferation has been proven in easy muscle cells of the vasculature [81]. A number of most cancers cell strains harbor elevated expression of KCNH1, and rising this channel’s expression in cells can remodel them into cancer-like cells [82]. KCNH5 is extremely up-regulated in medulloblastomas, and focused down-regulation of this channel lowered blastoma development in vivo [83].
Tumor tissues and cell strains also can have lowered expression of the potassium channel regulator KCNRG [84], which reduces Ok+ currents throughout the plasma membrane [85,86] and reduces cell proliferation [84]. It’s hypothesized that KCNRG reduces Ok+ channel expression, thereby limiting the discharge of intracellular Ok+ by within the plasma membrane [86]. Alongside these strains, decreased expression of Kv1.3, a rectifying Ok+ channel concerned in restoring intracellular Ok+ ranges, was noticed to be down-regulated in breast adenocarcinoma cell line MCF-7 [87].
Our information are according to different discovering that present that Ok+ channels that cut back intracellular Ok+ to advertise development. Nevertheless, ours and others’ findings additionally point out that Ok+ channel exercise regulates greater than cell proliferation, since Ok+ channel exercise regulates the transcription of a number of morphogens in numerous areas of the growing buds and grownup fins [11,88]. If Kcnk channels acted solely as oncogenes within the fin buds and grownup fins, then they’d possible produce tumors slightly than foster coordinated allometric development of the whole anatomical constructions [9,11] or reversed polarity of regenerative outgrowth in grownup fins [88]. Moreover, the coordinated patterning alterations in craniofacial and limb constructions by faulty Ok+ channels additionally counsel that these channels regulate greater than cell proliferation.
The findings that particular person dysfunctional Ok+ channels produce compound defects counsel that Ok+ channels have numerous actions through completely different molecular mechanisms. Nevertheless, within the majority of circumstances, it stays unclear how these channels are concerned. In the end, to get a greater understanding for a way particular Ok+ channels regulate the formation and development of particular tissues and what regulates their channel contributions, future tissue-specific focusing on of particular Ok+ channels is required together with in vivo assessments of their electrophysiological actions.
In abstract, we suggest that RA regulates intracellular Ok+ through an Rcan2-mediated enhance in Kcnk5b exercise to advertise fin bud development. The ensuing lower in intracellular Ok+ ranges causes IP3R-mediated Ca2+ launch that enhances shha transcription and different morphogens both immediately or by Shha manufacturing (Fig 8). Thus, our observations combine Ok+-channel-regulated scaling into identified molecular controls of fin/limb bud improvement. Whereas our proposed mechanism describes how one Ok+-leak channel is linked into developmental alerts, particulars of the mechanisms nonetheless must be outlined. Given the range of ion channels and the significance of Ok+ in regulating the electrophysiological properties of cells, our findings could have broader implications in organ scaling and ailments which might be attributable to the lack of proportional development. Continued work will decide the extent and variety in how the electrochemical properties of cells interface with the molecular controls that govern organ improvement and proportional development.
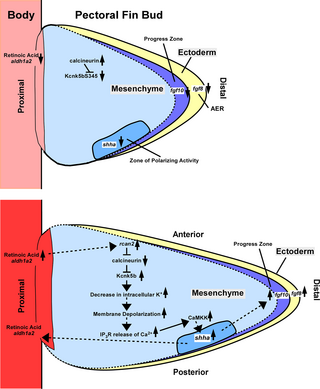
Fig 8. Mannequin.
RA signaling induces the transcription of rcan2, an endogenous inhibitor of calcineurin, to alleviate calcineurin inhibition of Kcnk5b channel exercise. Kcnk5b decreases intracellular Ok+ ranges within the mesenchyme that enhance depolarization to advertise IP3R-mediated Ca2+ launch from the ER. The rise in intracellular Ca2+ prompts CaMKK, and each are required for the elevated transcription of shha and enhanced development of the pectoral fin buds by Kcnk5b. ER, endoplasmic reticulum; RA, retinoic acid.
Supplies and strategies
Cloning
Constructs have been designed both by customary restriction enzyme or by homologous recombination strategies. KIRIN1 was synthesized (Genewiz) and cloned into MCS area of pcDNA6-myc-6xHIS-tag plasmid (Invitrogen, V22120) or pBluescript (VWR, 95040–830) harboring the hsp70 zebrafish promoter by 2 miniTol2 websites (transgenic vector). We cloned kcnk5b-GFP, GFP, kcnk10a-GFP, kcnk5b-mCherry, CaMKK2-mCherry, camkk1b-mCherry, and mCherry, rcan2-mCherry into pcDNA-myc-6xHIS-tag or pBluescript II vector (Invitrogen AM1344) for expression in cells or fish. We cloned kcnk5b-GFP, rcan2-mCherry and mCherry into pXT7 vector (Addgene, #32995) for mRNA injection.
Zebrafish husbandry
AB pressure fish have been raised in 10 L tanks with continually flowing water, 26°C customary light-dark cycle (Model and colleagues) [89] HaiSheng aquarium system. Fish embryos and larva have been raised in 1× E3 medium (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4, 10%–5% Methylene Blue) till 10 to 12 dpf, then transferred to aquarium water tanks to develop. Transgenic strains established by screening for GFP, CFP, and mCherry expression after warmth shock. Experiments used female and male fish equally. Normally, embryos have been immobilized by including Tricane (MESAB) to E3 at a ultimate focus of 4 μm for two min (or 200 nM blebbistatin for 30 min the place indicated) after which embedded in 1% low-melting agarose for 10 to 30-min imaging periods. The place indicated, embryos have been immobilized with Blebbistatin in 200 nM ultimate focus for 30 min as a substitute of Tricane.
Ethics assertion
Fish experiments have been compliant to the final animal welfare pointers and protocols (#20200903003) permitted by legally licensed animal welfare committee, ShanghaiTech Animal Welfare Committee.
Era of transgenic strains
Zebrafish embryos have been collected at one-cell stage for plasmid injection. Transgenic strains harboring the hsp70:KIRIN1, hsp70:GFP hsp70:kcnk5b-GFP, hsp70:kcnk5b-mCherry, hsp70:mCherry, hsp70:kcnk10a-mCherry, or hsp70:rcan2-mCherry transgene plasmids have been created by injecting 300 μg/μl of every assemble along with mRNA of Tol2 transposase [90]. Constructive embryos have been screened after 37°C warmth shock for 1 h. Constructive embryos have been introduced as much as grownup fish and screened by crossing with wild-type fish. Then, the optimistic F1 and successive generations have been screened by heat-shocking and figuring out fluorescence expression. Grownup fish have been 6 months to a yr previous, until indicated in another way within the textual content. Embryonic phases that have been used are indicated within the textual content.
Development measurements
Embryos have been first staged by the hour till 30 hpf and subsequently by the revealed head to physique angles because the embryos continued develop [30 hpf (angle 85°), 32 hpf (90°), 34 hpf (97.5°), 36 hpf (105°), 38 hpf (110°), 40 hpf (115°), 44 hpf (125°), 46 hpf (130°), until 48 hpf (135°) [91]. Embryos have been subsequently measured each 2 h. At every designated stage, the embryos have been imaged utilizing a Zeiss Stereoscope. The fin bud areas, eye areas, and otic vesicle areas have been measured utilizing Zen 3.4 (Blue version) software program by outlining every anatomical construction with this system’s contour instrument and measuring the encompassed space.
Warmth-shock induction of transgenes
Dad or mum fish of heat-shock-driven transgenic strains have been both outcrossed (rcan2, kcnk5b) as hemizygosity to same-strain wild-type fish or in-crossed (KIRIN1, GCaMP6s) for homozygosity. Progeny have been collected in 1xE3 and raised at 28°C. All embryos have been display for his or her respective transgenes by a single warmth shock at 37°C for 1 h at 24 hpf and chosen for fluorescence. We decided that transgene expression was highest at 6 h submit warmth shock, so we designed our warmth shock regimens accordingly. For FLIM imaging, transgenic strains Tg[hsp70:KIRIN1] embryos have been warmth shocked for 1 h at 37°C imaged 4 to six h later for every time level of the time course experiments. Tg[hsp70:kcnk5b-GFP] embryos and non-transgenic siblings have been warmth shocked twice: as soon as at 36 hpf and once more at 42 hpf for 1 h at 37°C to maximise goal gene expression after which collected for the in situ and qRT-PCR experiments at 48 hpf. For pectoral fin bud development experiments, Tg[hsp70:kcnk5b-GFP] and Tg[hsp70:rcan2-mCherry] embryos and non-transgenic siblings have been warmth shocked at 36 hpf and once more at 42 hpf after which imaged at 48 hpf to maximise their expression over a good portion of the pectoral fin bud development interval. For depolarization inhibitor therapies and subsequent DiSBaC2(3) experiments, Tg[hsp70:kcnk5b-GFP] embryos and non-transgenic siblings have been warmth shocked at 26 hpf and imaged at 32 hpf. For depolarization inhibitor therapies and subsequent measurements of fin bud measurement, Tg[hsp70:kcnk5b-GFP] embryos and non-transgenic siblings have been warmth shocked at 32 hpf as soon as to keep away from the center issues induced by Na+ channel inhibition. For grownup Tg[hsp70:rcan2-mCherry] and non-transgenic fish throughout caudal fin regeneration, the entire fish was incubated in 38.5°C aquarium water for 12 min as soon as a day throughout every particular experiment.
In situ hybridization
mRNA probes have been made out of RT-PCR merchandise remoted from 2 dpf zebrafish embryos or caudal fins from 6-month-old adults. The primer sequences for producing the revealed probes [92] are listed in Table 1. mRNA probes have been made out of RT-PCR merchandise from fish larva (5 dpf) or regenerating (3 dpa) grownup caudal fins. In vitro transcription reagents have been from Promega. Embryos and tail fins have been incubated in 4% PFA in 1xPBS at 4°C in a single day with light rocking. Samples have been dehydrated by incubation for 15 min in methanol at room temperature after which incubated in 100% methanol for ≥2 h at −20°C. Samples have been then rehydrated utilizing the reversed dehydration collection of methanol/1xPBS options (75%, 50%, 25%). Samples have been then incubated greater than 4× in 1xPBS to take away all methanol, and subsequently incubated in 10 μm Proteinase Ok for 10 min at RT. Samples have been then incubated 20 min in 4% PFA/1xPBS to inactivate the Proteinase Ok. Samples have been incubated in 1xPBS 6× 10 min to take away the PFA, then incubated in pre-hybridization buffer for 3 h at 65°C. Samples have been subsequently incubated within the hybridization resolution containing 200 ng/ml of every mRNA probe ≥14 h at 65°C. Samples then have been washed with successive wash steps to take away unbound probe and put together for antibody incubation: as soon as 2xSSC/75% deionized formalin at 65°C, as soon as 2xSSC/50% deionized formalin at 65°C, as soon as 2xSSC/25% deionized formalin at 65°C. 2xSSC at 65°C, twice 0.2xSSC at 65°C, 6 instances 1xPBST (1xPBS with Tween-20), as soon as in blocking resolution [2% bovine albumin (Sigma-Aldrich, A3294-100G), 2% Sheep Serum (Meilunbio, M134510)] at RT for 4 h. Samples have been incubated with Anti-digoxigenin-AP Fab Fragment (Sigma-Aldrich, 11093274910, RRID: AB_514497) in blocking resolution ≥14 h at 4°C. Samples have been then washed 6 instances with 1xPBST, subsequently incubated in [0.1 M Tris-HCl (pH 9.5), 0.1 M NaCl, 0.05 M MgCl2] 3 instances for 30 min, after which in Nitro Blue Tetrazolium (Sigma-Aldrich, N6639-1G) and 5-bromo-4-chloro-3-indolyl phosphate (Sigma-Aldrich, 136149) in [0.1 M Tris-HCl (pH 9.5), 0.1 M NaCl, 0.05 M MgCl2] at RT ≥8 h. Samples photos below VS120-S6-W (OLYMPUS). For sign areas, photos of the shha in situs have been transferred to Fiji ImageJ (RRID SCR_003070). The stained areas have been traced utilizing “free ROI” after which quantitated utilizing the “measure” operate below the “evaluation” menu to calculate the variety of pixels contained within the stained space. For sign depth evaluation, the sign area and adjoining unstained areas have been measured by “free ROI” choice and the imply depth pixel values have been decided from the “measure” operate below the evaluation menu by subtracting the imply worth of the unstained area from the imply worth of the sign area for every fin bud.
qRT-PCR
For transgenic expression in fish embryos, the embryos have been heat-shocked twice at 6-h intervals at 32 hpf and 42 hpf to maximise power transgene results on developmental gene expression. Fin buds have been remoted through the use of tip of a micropipette to softly take away the yolk of the 48 hpf embryos. The heads have been eliminated utilizing the small lancet iris knife (Zhongyuan Healthcare Tools, Cat # hyl10.31372714). The principle physique of the embryo was eliminated through the use of tweezers to repair the physique (neural tube going through up) and chopping the fin buds and residual yolk membrane from the physique on each aspect with a small lancet iris knife. The fin buds and residual yolk membrane have been collected within the TRIZOL (CWBio, 03917). Over 40 embryos (80 fin buds) have been collected for every pattern group. For grownup caudal fin tissues, we warmth shocked grownup fish as soon as for 12 min at 38.5°C 6 h earlier than the tissue isolation. After anesthetizing grownup fish with 0.4% Tricane, we surgically remoted the distal caudal fin tissues from the fin tip together with the distalmost bone bifurcation. For qRT-PCR experiments from HEK293 cells, we remoted RNA from 90% confluent 6 cm nicely per replicate. The mRNA from fish or cultured HEK293 cells was remoted utilizing Trizol (CWBio, 03917). Then, 1 μg mRNA was used for the reverse transcription to cDNA utilizing 4× gDNA wiper Combine, 5× HiScript III qRT SuperMix (Vazyme, L/N 7E350C9). qRT-PCR was carried out utilizing 2× ChamQ Common SYBR qPCR Grasp Combine (Vazyme, L/N TE342F9) with QuantStudio3 machine (Thermo Fisher). The cycle process was at 50.0°C for two min, 95°C for 10 min within the stage 1; 95°C for 15 s, 55.0°C for 20 s for 40 routine within the stage 2; 95°C for 15 s, 60°C for 1 min, 95°C for 15 s within the Soften Curve. Samples have been standardized utilizing the detected ef1a expression for fish mRNA isolation and GAPDH for human cell line mRNA isolations. Please see the “Table 1” on this part for the primer sequences of every gene.
Protein extraction and western blot
Zebrafish have been anesthetized with 0.4% Tricaine and fin ideas (fin tissues from the distalmost edge of every fin to the two distalmost bone segments of the bone rays) have been surgically remoted and homogenized in RIPA buffer with protease inhibitors (Thermo/Pierce), then incubated at 4°C for 1 h below fixed shaking. Lysates have been centrifuge at 13,000r for 15 min at 4°C. Protein focus was decided utilizing the BCA technique package (MDbio). Proteins have been denatured (incubated at 95°C for 10 min in loading buffer after which transferred to ice instantly previous to loading (50 μg/lane) in a ten% SDS-PAGE Acrylamide-Bis (29:1) gel). After electrophoresis, gel contents have been transferred onto 0.2 μm PVDF membrane. After switch, the membrane incubated in 5% skim milk/0.5% TBST for 1.5 h at room temperature after which washed 3× 5 min room temperature. Blots have been then incubated with Rcan2 major antibody (1:1,000) in a single day at 4°C. Blots have been then wash in 1xTBST 3× 5 min after which incubated in secondary antibody (1:1,000) in blocking buffer for two h at room temperature. Blots have been then washed 3× 5 min. Protein bands have been detected utilizing ECL chemiluminescence detection reagent (Perkin Elmer) and luminescence was detected utilizing an Amersham Imager 600 chemiluminescence imager (Common Electrical).
Cell tradition
HEK293 cell cultures (from ATCC.org) have been incubated at 37°C, 5% CO2, 95% humidity in incubators (Thermo Fisher, FORMA STERI-CYCLE i160) in DMEM medium (Gibco,1199506) with 10% FBS (Gibco,1009914) and 1% penicillin streptomycin (Gibco, 15140122). The id of the cell strains was not authenticated, and mycoplasma was not detected. Cells have been cut up to 50% density and transfected with Lipofectamin (Invitrogen, 11668–019) 12 h later. Expression for the transfected constructs was evaluated by expression of fluorescent marker. All cultures examined unfavourable for mycoplasma, which we examined by gathering 1 ml of DMEM cultured over 24 h with HEK293 cells, and centrifuge at 12,000 rpm for 1 min. Then, do the PCR as following primers: MGSO-TGCACCATCTGTCACTCTGTTAACCTC, GPO-3-GGGAGCAAACAGGATTAGATACCCT.
FRET-FLIM detection and evaluation
HEK293 cells have been transfected with 1 μg of pcDNA-kcnk5b-GFP; pcDNA6-mT2-CUTie-YFP [40] or pcDNA-CFP-cGi500-YFP [41]. The transgenic fish [hsp70:KIRIN1] have been warmth shock at 37°C for 1 h for the sensor to induce peak transgene expression 6 h later. Embryos have been incubated in 250 μm PTU (1-phenyl 2-thiourea, Sigma) to stop pigmentation beginning at 24 hpf. Ten minutes earlier than imaging, embryos have been anesthetized in 4 μm MESAB (or 200 nM blebbistatin for 30 min, the place indicated) after which embedded in 1% agarose earlier than imaging. For different imaging experiments, the time factors have been indicated within the textual content or determine legends. For FLIM measurements of the grownup caudal fins, grownup fish (roughly 6 months previous) have been anesthetized with 1% MESAB after which positioned on moist plastic Petri dish and imaged for five min after which positioned again into aquarium water to regain consciousness. Measurements have been from 3 completely different places within the mesenchyme of caudal fins: 1 at every of the two lobe ideas and 1 within the distal midline between the two lobes. Every measurement is 1 information level. For the embryonic developmental time course of fluorescence lifetime imaging (FLIM-FRET) measurements, every embryo was imaged at 32 hpf, 35 hpf, 38 hpf, 42 hpf, 48 hpf, and 56 hpf. Fluorescence lifetime imaging measurements have been made by photon counting the fluorescence emission of both CFP (KIRIN1, cAMP, cGMP) utilizing a 2-photon-confocal Hyperscope (Scientifica, United Kingdom) and PMT-hybrid 40 MOD 5 photon detectors (Picoquant, Germany). We decided publicity instances for every aircraft based mostly on the variety of photons detected particular person pixels. In every fin bud, we usually measured 3 areas throughout the mesenchyme and ectoderm by deciding on areas of curiosity at distal, proximal, and central places throughout the buds. Every area contained a number of pixels of that ranged from 1 to three cells in measurement. For every measurement, we used thresholds of a number of hundred photons per pixel to beat background photon emissions and forestall any affect within the FLIM measurements. The lifetime curves have been analyzed from graphically chosen areas or cells within the confocal aircraft. The lifetime curves have been decided (curve becoming) from the decay of the counted photon emissions after single laser pulses (2-photon Chameleon Extremely II Ti Sapphire laser) utilizing the algorithm
for base computations of multi-exponential reconvolution with corrections for instrument response time (instrument response issue, IRF) and removing of background alerts <100/pixel in this system SymPhoTime 64, model 2.4 (Picoquant, Germany, RRID: SCR_016263). The fitted curves represented 2 part (fluorophores FRET at <10 nm and fluorophores no FRET >10 nm) equations with χ2 values of 1 ± 0.19 for accuracy. To visualise variations in spatial distribution of every pixel lifetime worth, particular person lifetime values have been distributed alongside a rainbow scale wherein boundary lifetime values have been assigned: blue lowest boundary and pink highest boundary. FRET effectivity (on which our FLIM measurements are based mostly) utilizing our donor-only (CFP) and sensor-based donor + acceptor (CFP-YFP) Ok+-detection information in Fig 1A. The ensuing common effectivity was 37.9%. We entered this calculated effectivity in a standardized equation [RDA = R0 x 6√(1-Efficiency)2/Efficiency)] to calculate the efficient distance between the two fluorophores for FRET from our measurements, (4.59 Å) after which to match this to the reference distance of maximal FRET of CFP and YFP (4.7 Å) [93]. RDA is calculated distance between donor and acceptor fluorophores. R0 is the gap required for 50% effectivity. The closeness our calculated and reference distances indicated significant effectivity of the Ok+-sensor in our in vivo measurements.
KIRIN1 sensor
As beforehand revealed, KIRIN1 sensor was generated by fusing CFP and YFP to the N- and C-termini of the Ok+-binding protein (Kbp) from E. coli [28]. The Kbp protein usually maintains an elongated conformation that retains the donor and acceptor fluorophores aside, which prevents FRET between the fluorophores. When Ok+ binds to the bacterial OsmY (BON) area within the N-terminus and the Lysin motif (LysM) within the C terminus, it brings the two fluorophores collectively to permit FRET [28]. This ends in a discount within the lifetime of the donor (CFP) fluorophore, which will be detected by FLIM.
DiSBAC2(3) staining and measurements
DiSBAC2(3) was diluted to a ten μm focus in 1× E3 medium that contained 250 μm focus of PTU (1-phenyl 2-thiourea, Sigma, P7629). Embryos have been put into PTU at 24 hpf to stop the pigmentation of the embryos. Three hours earlier than imaging, embryos have been incubated in diluted DiSBAC2(3) at RT below light rocking. They have been immobilized by including blebbistatin to a ultimate focus of 200 nM for 30 min earlier than imaging, embedded in 1% low-melting agarose gel, after which imaged below a 2-photon-confocal Hyperscope (Scientifica, UK) at 900 nm to maximise the detection of the fluorescence from the DiSBAC2(3) dye and reduce fluorescence of mCherry below 2-photon stimulation (www.fpbase.org). Photon counts have been detected utilizing PMT-hybrid 40 MOD 5 photon detectors (Picoquant, Germany) that detect photons (present a numerical rely) for every pixel. We used blebbistatin, as a result of it inhibits muscle motion by inhibiting myosin exercise and doesn’t goal any ion channel. We imaged every embryo for 3 min and measured 6 pixels in particular areas in every fin bud (distal mesenchyme, anterior mesenchyme, proximal mesenchyme, and posterior mesenchyme). The 6 pixels from every area have been averaged and every common represents an information level within the graph; 32 hpf have been chosen for imaging the impact of kcnk5b overexpression, due to the low ranges of depolarization at this stage (Fig 5A–5C).
CRISPR knockouts for rcan2 and kcnk5b
The ChopChop on-line instrument (http://chopchop.cbu.uib.no/) was used to design sgRNAs to restrict predicted off-target sgRNA chopping. The fourth exon of rcan2 was focused by the sgRNA sequence is 5′-AACCGACTGGAGGTGAGCCA-3′ ordered from IDT with Alt-R gRNA modification. The primary exon in kcnk5b was focused by the sgRNA sequence 5′-TGCGATATTCCAAATCCTCG-3′ and 5′-GGTCTTGGGCGAAATTATTG-3′ ordered by IDT with Alt-R gRNA modification, and 1 μl sgRNAs (400 ng/μl) and Cas9 (500 ng/μl) protein (Genscript, Cat. #Z03469) have been co-injected into single-cell zebrafish embryos. After imaging for experiments, every embryo was numbered and raised individually to five dpf. Genomic DNA was remoted from every 5 dpf larva individually. After genomic DNA extraction by alkaline lysis buffer, the genomic area surrounding the goal websites was PCR amplified utilizing primers ahead: 5′-CAACACACTCTCTGGCTTTCAG-3′ and reverse: 5′-TGAACTGCATAGTTGAGATGGG-3′ for rcan2 or ahead: 5′-ATCACCAGAAACTTGGGAGTGT-3′ and reverse: 5′-GCTTTTGCGTGAGAACTACCAT-3′ for kcnk5b and set for sequencing (Genewiz.com.cn).
In vivo mRNA overexpression experiments
Mutated rcan2-P2A-mcherry, kcnk5b-GFP and kcnk5bS345A-GFP have been cloned into pXT7 vector (from Lin Haifan and Bao Baolong lab) and both linearized with Xba1 (Anza Thermo Scientific #ER0682) or amplified by PCR to make templates. For rcan2, the unique goal sequence 5′-AACCGACTGGAGGTGAGCCA-3′ was mutated to 5′-ATCCAACCGGTGGGCTTCCG-3′. For kcnk5b, the unique target-2 sequence 5′-TGCGATATTCCAAATCCTCGAGG-3′ was mutated to 5′-TGCAATCTTTCAGATACTAGAAG-3′, and the unique target-14 sequence 5′-GGTCTTGGGCGAAATTATTGAGG-3′ was mutated to 5′-GGTATTAGGAGAGATCATCGAAG-3′ to impair interplay between the sgRNA focusing on sequences and the rescue mRNAs with out altering coded amino acids. The linear template was transcribed to mRNA utilizing T7 MESSAGE MACHINE (Invitrogen, #AM1344). Concentrations injected into embryos: 1 μl Cas9 protein, 1 μl sgRNA, 1 μl rcan2-P2A-mCherry mRNA/kcnk5b-P2A-GFP (700 ng/1 μl), mCherry mRNA (375 ng/1 μl), 1 μl empty sgRNA (T7-sgRNA scaffold, 2,000 ng/μl) or kcnk5bS345-GFP/kcnk5bS345A-GFP (200 ng/μl).
Mosaic analyses in fin buds
Double mosaic embryos have been created by injecting 120 μg/μl of hsp70:kcnk5b-mCherry and 400 ng/μl into Tg[hsp70:KIRIN1] individually into 1–4 cell AB embryos. Embryos solely expressing kcnk5b-mCherry mosaically have been injected with hsp70:kcnk5b-mCherry into the transgenic Tg[Cca.actb:GCaMP6s] line. Injected embryos and larva have been raised in 1xE3 medium (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4, 10%–5% Methylene Blue). The mosaic larvae have been chosen by screening for mCherry and CFP expression for kcnk5 and KIRIN1 transgenes or mCherry and GFP expression for kcnk5b and GCaMP6s transgenes within the fin buds 4 to six h after heat-shock induction.
Small molecule therapies
All inhibitors (aside from Vinpocetine) have been dissolved in DMSO: Retinoic acid (Abcone, R26255) 10 mM inventory focus, 2-APB (IP3R inhibitor, ENZO, ALX-400-045-M100) 75 mM inventory focus, STO-609 (CaMKK inhibitor, MedChemExpress, HY-19805) 0.5 mM inventory focus, Verapamil (T-/L-type Ca2+ channel inhibitor, MedChemExpress, HU-A0064) 100 mM inventory focus, KN-62 (CaMKII/IV inhibitor, MedChemExpress, HY-13290) 1 mM inventory focus, or in sterile water Mibefradil (T-/L-type Ca2+ channel inhibitor, MedChemExpress, HY-15553A) 2.5 mM inventory focus, Forskolin (adenylyl cyclase activator, MedChemExpress, HY-15371) 25 mM, SNAP (guanylyl cyclase activator, Tocris, 1561) 100 mM inventory concentrations. HEK293 cells have been incubated in DMEM cell tradition medium (Gibco, 1199506) containing 10% FBS (Gibco, 1009914) and 1% penicillin streptomycin (Gibco, 15140122) at 37°C, 5% CO2. Ten hours after transfection, the medicine have been added to the medium to their ultimate concentrations (as indicated within the determine legends or the particular strategies part). Cells have been then trypsinized and RNA was remoted as indicated for qRT-PCR. For fish embryos, we added the small molecules talked about above to E3 to the ultimate concentrations (200 nM RA, 13 μm 2-APB, or 24 μm STO until point out in any other case) and incubated the embryos 28°C. We decided the remedy instances and concentrations (offered within the determine legends or textual content) to be the simplest on the fin bud development and have the least impact on embryonic improvement. Grownup fish have been handled with 100 μm RA for six h, since we decided this focus confirmed reproducible results (S5F Fig). Vinpocetine (MedChemExpress, HY-13295) was dissolved in ethanol to make a ten mM inventory, Dibucaine (MedChemExpress, HY-B0552) was dissolved in DMSO to make a 500 mM inventory. For assessing the results of those inhibitors on depolarization, we handled embryos with 10 μm Vinpocetine and 40 μm Dibucaine for 4 h earlier than evaluation, since these concentrations confirmed continued reproducible inhibitory results. For fin bud development experiments, 10 μm Vinpocetine and 40 μm Dibucaine remedy began at 36 hpf for 12 h to maximise the results of the inhibitor on fin bud development by 48 hpf.
GCaMP6s measurements
HEK293 cells have been transfected with 1 μg of CMV-GCaMP6s, 1 μg of pcDNA-kcnk5b-MCherrry or pcDNA-mCherry or pcDNA-Kcnk5bMut. The pcDNA-Kcnk5bMut harbors 2 mutations: one altering the LEEP sequence within the first transmembrane area to VKKA and substituting S345 within the C-terminal tail with alanine. This generated a lifeless channel. HEK293 cells for Fig 6E have been transfected and sorted by SORP ARIA Fusion (BD Biosciences) for mCherry. GCaMP6s transgenic fish Tg[Cca.actb:GCaMP6s] [43] have been bought from Zebrafish Inventory Heart Wuhan, China (CZ1282 doi.org/10.3390/ijms22115551). One-celled embryos from the established Transgenic fish Tg[Cca.actb:GCaMP6s] [43] have been injected with Tg[hsp70–kcnk5b-mCherry] and have been warmth shocked at 37°C for 1 h at 42 hpf and imaged at 48 hpf. All of the samples have been imaged by Zeiss LSM 980 upright and analyzed by ZEN-Blue (Zeiss). To standardize depth measurements and restrict affect of variations in expression, we standardized all measurements to GCaMP6s cells missing mCherry expression within the spinal twine (physique) away from the fin bud. The outcomes are offered as GFP bud/GFP physique.
Patch clamping
Transfected HEK293T cells have been seeded on glass coverslips (Fisher Model) and incubated in cell tradition medium at 37°C, 5% CO2, 95% relative humidity for 8 to 10 h. The seeded coverslips have been transferred into Tyrode’s resolution (138 mM NaCl, 4 mM KCl, 2 mM CaCl2, 1 mM MgCl2, 0.33 mM NaH2PO4, 10 mM Glucose, 10 mM HEPES). Cells have been assessed within the ruptured-patch whole-cell configuration of the patch-clamp approach utilizing and EPC9 or EPC10 amplifier (HEKA) with borosilicate glass pipettes (Sutter Devices) with 3 to six MΩ resistance when stuffed with pipette resolution (130 mM glutamic acid, 10 mM KCl, 4 mM MgCl2, 10 mM HEPES, 2 mM ATP, pH to 7.2). For detecting potassium present, after gigaseal formation, cells have been voltage-clamped at −80 mV. Potassium conductance was elicited by check pulses from −100 mV to 70 mV (in 10 mV increments) of 600 ms length at a cycle size of 10 s. The ensuing tracings have been transformed into itx information by the ABF Software program (ABF Software program, RRID: SCR_019222) after which analyzed utilizing Clampfit Software program (Molecular Gadgets, RRID: SCR_011323). Currents have been measured on the finish of the check pulses.
Quantification and statistical analyses
To pick for correct two-component decay curve fittings of the FRET sensors, we used Chi-square analyses limiting suits 0.8>χ2<1.2 values, which was calculated throughout the SymPhoTime 64 software program, model 2.4 (Picoquant, Germany, RRID: SCR_016263). Graphs have been assembled and statistical analyses have been performed utilizing Microsoft Excel (Microsoft.com) or Prism9 (Graphpad.com). All statistical analyses for graphs have been performed utilizing the College students t check to evaluate the reproducibility of any variations between 2 teams throughout the information units of every experiment. P values variations of >0.05 have been indicated to be not vital “NS”; in any other case, the calculated P values have been offered within the determine.
Supporting info
S1 Fig. FLIM-FRET measurements with KIRIN1 sensor detects relative adjustments in intracellular Ok+.
(Aa) Patch-clamp experiments present elevated Ok+ currents from Ok+ leaking out of cells expressing CMV:kcnk5b-mCherry (blue) in comparison with cells expressing the management CMV:mCherry plasmid (pink). (Ab) FLIM-FRET measurements of the KIRIN1 sensor expressed in the identical cells that have been patched in Aa detected decreases in intracellular Ok+ in cells expressing CMV:kcnk5b-mCherry (blue) in comparison with cells missing the expression of the channel (CMV:mCherry, pink). (B) FLIM-FRET measurements for intracellular Ok+ ranges in HEK293 cells transfected with the CMV:KIRIN1 sensor and both CMV:mCherry or CMV:Kcnk5b-mCherry. Cells have been both handled with DMSO or FK506. (C) FLIM-FRET measurements for intracellular Ok+ ranges in HEK293 Cells transfected with the CMV:KIRIN1 sensor and both CMV:Kcnk5bS345A-mCherry, CMV:Kcnk5bWT-mCherry, or CMV:Kcnk5bS345E-mCherry. The measurements have been transformed to fold distinction by dividing their lifetime measurements with the lifetime measurements of the management cells expressing the KIRIN1 sensor and mCherry. Every experiment was repeated at the least 3 instances (N = 3). For cell patch clamping and FLIM measurements, every information level represents 1 cell (A, B, C). For fish embryo FLIM imaging, we measured 2 or 3 places in every tissue of 1 fin bud per embryo. P values symbolize statistical evaluation by Pupil’s two-tailed t check. Numerical information used on this determine are included in S8 Data.
https://doi.org/10.1371/journal.pbio.3002565.s015
(TIF)
S2 Fig. FLIM-FRET measurements not affected by adjustments in expression ranges or sedation technique.
(A) Warmth-shock technique for inducing expression of the transgenic Ok+ sensor on the indicated time factors for subsequent FLIM-FRET measurements. (B) Fluorescent picture of non-transgenic sibling 6 h after warmth shock at 48 hpf. (C) Fluorescent picture of transgenic Tg[hsp70:KIRIN1] 6 h after warmth shock at 48 hpf. (D) Consultant confocal aircraft by a growing pectoral fin bud of a Tg[hsp70:KIRIN1] transgenic fish exhibits expression of the Ok+-sensor transgene in cells aside from the nuclei. (E) Illustration of various decay (lifetime) curves and that regardless of variations in preliminary excitation ranges of the donor fluorophore (arrows), the decay charges are comparable (orange decay curves) until power is transferred from the donor to the acceptor fluorophore by FRET within the presence of Ok+, which is able to cut back the lifetime (blue decay curve). These particular variations in lifetime will be represented by particular colours alongside a rainbow scale to provide a picture that relates the lifetime worth of every pixel within the confocal aircraft to supply a spatial illustration of the distribution of relative Ok+ ranges within the fin bud. (F) 3D photos of density map for illumination depth (a, b) of pectoral fin buds at 32 hpf. Comparability between 2D aircraft of density map for illumination at a area of excessive depth (c) and the lifetime evaluation of the identical area within the ectoderm (d). Comparability between 2D aircraft of density map for illumination at a area of low depth (e) and the lifetime evaluation of the identical area within the ectoderm (f). Comparability between 2D aircraft of density map for illumination at a area of excessive depth (g) and the lifetime evaluation of the identical area within the mesenchyme (h). Comparability between 2D aircraft of density map for illumination at a area of low depth (i) and the lifetime evaluation of the identical area within the ectoderm (j). (ok) Graph of lifetime values of every area measured (d, f, h, j) exhibits that prime and low variations in depth (y-axis) don’t considerably alter the lifetime values (x-axis) of the ectoderm and the mesenchyme. (G) 3D photos of density map for illumination depth (a, b) of pectoral fin buds at 48 hpf. Comparability between 2D aircraft of density map for illumination at a area of excessive depth (c) and the lifetime evaluation of the identical area within the ectoderm (d). Comparability between 2D aircraft of density map for illumination at a area of low depth (e) and the lifetime evaluation of the identical area within the ectoderm (f). Comparability between 2D aircraft of density map for illumination at a area of excessive depth (g) and the lifetime evaluation of the identical area within the mesenchyme (h). Comparability between 2D aircraft of density map for illumination at a area of low depth (i) and the lifetime evaluation of the identical area within the ectoderm (j). (ok) Graph of lifetime values of every area measured (d, f, h, j) exhibits that prime and low variations in depth (y-axis) don’t considerably alter the lifetime values (x-axis) of the ectoderm and the mesenchyme. (H) 3D photos of density map for illumination depth (a, b) of pectoral fin buds at 56 hpf. Comparability between 2D aircraft of density map for illumination at a area of excessive depth (c) and the lifetime evaluation of the identical area within the ectoderm (d). Comparability between 2D aircraft of density map for illumination at a area of low depth (e) and the lifetime evaluation of the identical area within the ectoderm (f). Comparability between 2D aircraft of density map for illumination at a area of excessive depth (g) and the lifetime evaluation of the identical area within the mesenchyme (h). Comparability between 2D aircraft of density map for illumination at a area of low depth (i) and the lifetime evaluation of the identical area within the ectoderm (j). (ok) Graph of lifetime values of every area measured (d, f, h, j) exhibits that prime and low variations in depth (y-axis) don’t considerably alter the lifetime values (x-axis) of the ectoderm and the mesenchyme. (I) Consultant FLIM-FRET picture of the distribution of the lifetimes of a fin bud from a 32 hpf embryo immobilized with 200 nM Blebbistatin. (J) Consultant FLIM-FRET picture of the distribution of the lifetimes of a fin bud from a 32 hpf embryo immobilized with 4 μm MESAB (Tricane). (Ok) Consultant FLIM-FRET picture of the distribution of the lifetimes of a fin bud from a 48 hpf embryo immobilized with 200 nM Blebbistatin. (L) Consultant FLIM-FRET picture of the distribution of the lifetimes of a fin bud from a 48 hpf embryo immobilized with 4 μm MESAB. (M) Consultant FLIM-FRET picture of the distribution of the lifetimes of a fin bud from a 56 hpf embryo immobilized with 200 nM Blebbistatin. (N) Consultant FLIM-FRET picture of the distribution of the lifetimes of a fin bud from a 56 hpf embryo immobilized with 4 μm MESAB. (O) Graphed FLIM-FRET measurements of a number of fin buds of embryos immobilized both with MESAB or Blebbistatin from the indicated time factors exhibits no vital variations (P > 0.05) between the two therapies. For the FLIM measurements of Blebbistatin-treated and MESAB-treated embryos, experiments have been repeated at the least 3 instances (N = 3), and every repeat contained 3 or extra embryos. We measured 2 or 3 places in every tissue of 1 fin bud per embryo. Every measured worth is represented as an information level (O). P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values ≥0.05 are designated as “not vital” (NS). Scale bars equal 1 mm (B, C), 20 μm (D), 30 um (Ha-i), or as indicated within the panels. Numerical information used on this determine are included in S9 Data.
https://doi.org/10.1371/journal.pbio.3002565.s016
(TIF)
S3 Fig. Confocal planes of FLIM-FRET photos in fin buds at completely different developmental time factors.
(A) Confocal planes in fin bud from a 32 hpf embryo. The space between the primary aircraft and final aircraft is 15.2 μm. (B) Confocal planes in a fin bud from a 48 hpf embryo. The space between the primary aircraft and final aircraft is eighteen.36 μm. (C) Confocal planes in a fin bud from a 56 hpf embryo. The space between the primary aircraft and final aircraft is 10.3 μm. Numbers in decrease proper of every panel point out the order of the indicated confocal aircraft by the Z-stack. (D) FLIM measurements in fin buds of 56 hpf embryos of indicated cell classes from the transgenic KIRIN1 fish line Tg[hsp70:KIRIN1] mosaically expressing mCherry or kcnk5b-mCherry. “Adj” signifies cells adjoining to mCherry-positive (mCherry+) or kcnk5b-mCherry-positive (kcnk5b+) cells. Every experiment was repeated at the least 3 instances (N = 3). For fish embryo FLIM imaging, we measured 2 or 3 places in every tissue of 1 fin bud per embryo. Every measured worth is represented as an information level (D). Numerical information used on this determine are included in S10 Data.
https://doi.org/10.1371/journal.pbio.3002565.s017
(TIF)
S4 Fig. Overexpression of Ok+-leak channels and their results on pectoral fin bud development.
(A) qRT-PCR for kcnk5b-GFP expression after single warmth shock pulse of non-transgenic siblings and Tg[hsp70:kcnk5b-GFP] siblings. Every RNA pattern was remoted from fin buds of 40+ embryos at 48 hpf. (B) Expression of kcnk5b-GFP within the physique (a) and within the fin bud (b) of a consultant Tg[hsp70:kcnk5b-GFP] 56 hpf embryo 6 h after warmth shock. (C) Expression of kcnk10a-GFP within the physique (a) and within the fin bud (b) of a consultant Tg[hsp70:kcnk10a-GFP] 56 hpf embryo 6 h after warmth shock. (D) Brightfield picture of thorax area of a post-heat-shocked 48 hpf non-transgenic embryo. The realm of the fin bud (highlighted by a black-dotted line). The otic vesicle was used as a measurement customary (highlighted by a red-dotted line). (E) Brightfield picture of thorax area of a post-heat-shocked 48 hpf hsp70:kcnk5b-GFP transgenic embryo with the fin bud (black-dotted line) and otic vesicle (red-dotted line) used as a measurement customary. (F) Measurements of pectoral fin bud areas of heat-shocked teams of non-transgenic (Non-Tg) and Tg[hsp70:GFP] (GFP-Tg) as controls and Tg[hsp70:kcnk5b-GFP] transgenic fish line in addition to the Tg[hsp70:kcnk10a-GFP] transgenic fish line. Every measured bud space was standardized to the otic vesicle space in the identical embryo and every measurement is represented as a ratio of bud-area–to–otic-vesicle space. Every experiment was repeated at the least 3 instances (N = 3) and every repeat contained 3 for extra fish. Every information level represents 1 fin bud measurement per embryo. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values ≥0.05 are designated as “not vital” (NS). Numerical information used on this determine are included in S11 Data.
https://doi.org/10.1371/journal.pbio.3002565.s018
(TIF)
S5 Fig. Results of nuclear hormone therapies on kcnk5b, rcan2, and management genes in pectoral fin buds and grownup fins and sufficiency/necessity experiments for rcan2.
(A) qRT-PCR measurements of the transcription of kcnk5b within the growing pectoral fin bud with and with out retinoic acid stimulation. (B) qRT-PCR measurements of cyp26a expression in pectoral fin buds with or with out 200 nM retinoic acid remedy, a gene identified to be induced by retinoic acid remedy. (C) FLIM measurements from Ectoderm (Ecto) and Mesenchyme (Mesen) cells of buds at 32 hpf handled both with DMSO or 200 nM thyroid hormone for six h. (D) In situ staining depth measurements from the in situs of rcan2 expression within the embryonic fin buds after DSMO or RA remedy. (E) qRT-PCR for expression of rcan2 and dio3b in grownup caudal fin after remedy with DMSO or 500 nM thyroid hormone for twenty-four h. dio3b is a gene identified to be up-regulated by thyroid hormone stimulation within the grownup caudal fin. (F) RT-PCR for expression of cyp26a handled both with DMSO or 100 nM RA for six h in grownup caudal fin. (G) FLIM measurements of intracellular Ok+ ranges in grownup caudal fin cells after 6 h handled both with DMSO or 100 nM RA. (H) qRT-PCR measurements utilizing 2 completely different primer units for rcan2 in grownup caudal fins of indicated remedy teams. (I) Entire-mount in situ hybridization of a caudal fin 3 day submit amputation exhibits rcan2 expression within the distal blastemal however absent from the proximal blastema (a). Arrowheads point out amputation aircraft. Cryo cross sections by the three day submit amputation, distal tip of a regenerating grownup caudal fin within the ray (b) and interray tissues (c) after in situ hybridization for rcan2. The blue coloration signifies rcan2 expression. (J) Consultant western blot for Rcan2 and beta-actin proteins from lysates of regenerating grownup caudal fins on the indicated days submit amputation (dpa). (Ok) Measurements of Rcan2 protein expression after standardization to beta-actin expression on the indicated time factors. (L) Consultant brightfield picture of non-transgenic sibling 6 h after heat-shock stimulation. (M) Consultant mCherry fluorescence picture of non-transgenic sibling 6 h after heat-shock stimulation. (N) Consultant brightfield picture of transgenic Tg[hsp70:rcan2-mCherry] sibling 6 h after heat-shock stimulation. (O) Consultant mCherry fluorescence picture of transgenic Tg[hsp70:rcan2-mCherry] sibling 6 h after heat-shock stimulation. (P–R) Consultant amputated caudal fin of heat-shocked AB non-transgenic (non-Tg) fish at time 0 days submit amputation (dpa) (P) and regenerated 56 dpa (Q) along with its consultant fluorescence picture for mCherry (R). (S–U) Consultant caudal fins of Tg[hsp70:rcan2-mCherry] transgenic fish (rcan2-Tg) at time 0 dpa (S) and regenerated 56 dpa (T) along with its consultant fluorescence picture for mCherry (U). (V) Fin-to-body size ratios of non-transgenic and Tg[hsp70:rcan2-mCherry] fish on the indicated time factors throughout one-heat-shock per day routine. (W) Wild-type 48 h submit fertilization (hpf) larva (a). Sequence of rcan2 gene in exon 4. (b) The sgRNA goal website is indicated by the orange field. Of the full variety of injected embryos (45) assessed, solely 9% confirmed wild-type sequence. (X) 48 hpf larva of huge deletion group (a). Nearly all of focused embryos (64%) harbored an analogous giant deletion: indicated by the hole in sequence. The downstream sequence additionally confirmed scrambled nucleotide order (b). (Y) 48 hpf larva of small deletion group (a). Over 1 / 4 of the focused embryos (27%) harbored comparable small deletions in addition to scrambled downstream sequence (b). (Z) Consultant mCherry fluorescent picture of CRISPR rescue expression of rcan2*-mCherry mRNA (a) and CRISPR-targeted rcan2 allele from an rcan2* mRNA-mCherry-expressing embryo (b). rcan2*-mCherry mRNA has mutated wobble-position nucleotides of codons to impair interplay with the sgRNA to proceed sgRNA-mediated disruption of the rcan2 alleles whereas sustaining the integrity of the transgenic Rcan2 protein. (AA) Consultant photos of management (a) and CRISPR-targeted rcan2 KO (b) juvenile fish 76 days submit fertilization. (BB) Caudal fin-to-body size ratios between management AB and rcan2 KO juvenile fish. Scale bars equal 20 μm (I), 100 μm (L–NO, W–Z), 1 mm (P–U) and 500 μm (AA). For qRT-PCR experiments from fin buds, every experiment was repeated at the least 3 time (N = 3) and every repeat contained duplicate or triplicate samples. Every pattern contained 80+ fin bud isolations (A, B). For the FLIM measurements of embryos, we made 3 measurements per tissue in every fin bud. Not less than 3 embryos have been measured per group per experiment, and every experiment was repeated 3 instances. Every information level represents a measurement (C). For FLIM measurements from grownup fins, we measured 3 places within the mesenchyme of every unamputated grownup caudal fin: one at every of the two lobe ideas, and one within the midline between the two lobes on the midline of the fin. Every information level represents 1 measurement (G). For qRT-PCR and western blots experiments with grownup caudal fins, every level represents a separate pattern group from an isolation of the distal ideas with 2 segments from 30+ fish (H, I). For fin bud and grownup fin development measurements, every experiment was repeated 3 instances. The grownup fin measurements in S5V contained 10+ fish per group every time. The measurements in S5BB, every level represents 1 fish. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values ≥0.05 are designated as “not vital” (NS). Numerical information used on this determine are included in S12 Data.
https://doi.org/10.1371/journal.pbio.3002565.s019
(TIF)
S6 Fig. CRISPR kcnk5b KO technique, focusing on effectivity of endogenous locus and Kcnk5b mutants.
(A) Wild-type 48 hours submit fertilization (hpf) larva (a), and sequence of kcnk5b gene in exon 1 (b). Of 100 embryos focused, 12% don’t present gene defects. (B) 48 hpf embryo (a) harboring a big deletion within the kcnk5b exon 1 (b), and 44% harbor giant deletions. (C) 48 hpf embryo (a) harboring small deletions and sequence adjustments (b), and 44% harbor such small deletions. The sgRNA goal websites are indicated by the grey field labeled “goal.” (D, E) Consultant embryo with kcnk5b CRISPR knockout (kcnk5b KO) brightfield (D) and lack of inexperienced fluorescence (E) of CRISPR focused embryos. (F, G) Consultant embryo kcnk5b KO brightfield (F) and GFP from management GFP mRNA. (H, I) Consultant embryo kcnk5b KO rescued with expression of kcnk5b*-GFP mRNA, brightfield (H) and inexperienced fluorescence (I). kcnk5b*-GFP mRNA has mutated wobble-position nucleotides of codons to impair interplay with the sgRNA to proceed sgRNA-mediated disruption of the kcnk5b alleles whereas sustaining the integrity of the transgenic Kcnk5b protein. (J) Consultant disrupted kcnk5b allele sequence after CRISPR-targeting and kcnk5b*-GFP-expressing embryo. (Ok, L) Wild-type, uninjected embryo for GFP (Ok) and absence of mCherry (L) fluorescence. (M, N) Consultant transgenic embryo for kcnk5bS345-GFP (M) and rcan2-mCherry (N) mRNAs. (O, P) Consultant transgenic embryo for kcnk5bS345A-GFP mutant (O) and rcan2-mCherry (P) mRNAs. Scale bars equal 100 μm (A–C, D–I, Ok–P).
https://doi.org/10.1371/journal.pbio.3002565.s020
(TIF)
S7 Fig. Growth of embryos incubated with Vinpocetine, Dibucane, and DiSBAC2(3).
(A) Consultant picture of a 48 hpf AB non-transgenic fish after 1 warmth shock at 32 hpf and handled solely with the solvents ethanol (EtOH) and DSMO for 12 h beginning at 36 hpf. (B) Consultant picture of 48 hpf AB non-transgenic fish after 1 warmth shock at 32 hpf and handled with 10 μm Vinpocetine (Vin) and 40 μm Dibuciane (Dib) for 12 h beginning at 36 hpf. (C) Consultant picture of a 48 hpf Tg[hsp70:kcnk5b-GFP] fish heat-shocked as soon as at 32 hpf to induce expression of kcnk5b-GFP and handled for 12 h with the solvents EtOH and DMSO at 36 hpf. (D) Consultant picture of a 48 hpf Tg[hsp70:kcnk5b-GFP] fish heat-shocked as soon as at 32 hpf to induce expression of kcnk5b-GFP and handled with 10 μm Vinpocetine (Vin) and 40 μm Dibuciane (Dib) for 12 h beginning at 36 hpf. (E–J) Embryos incubated in 10 μm DiSBAC2(3) dye for 3 h present dye penetration within the embryos. mCherry fluorescence of embryo handled with EtOH and DMSO earlier than DiSBAC2(3) incubation (E). Brighfield picture of embryo handled with EtOH and DMSO after DiSBAC2(3) incubation (F). Fluorescence of DiSBAC2(3) in embryo handled with 10 μm Vin and 40 μm Dib earlier than DiSBAC incubation (I). mCherry fluorescence of embryo handled with Vin and Dib earlier than DiSBAC2(3) incubation (H). Brighfield picture of embryo handled with Vin and Dib after DiSBAC2(3) incubation (I). Fluorescence of DiSBAC2(3) in embryo handled with Vin and Dib after DiSBAC2(3) incubation (J).
https://doi.org/10.1371/journal.pbio.3002565.s021
(TIF)
S8 Fig. Results of drug therapies on tradition cells and embryos.
(A) FLIM-FRET measurements of intracellular cAMP ranges. Forskolin used as optimistic management for cAMP manufacturing. (B) FLIM-FRET measurements of intracellular cGMP ranges. SNAP used as optimistic management for cGMP manufacturing. (C) Patch-clamp experiment measuring the Ok+ leak from HEK293T cells transfected both with Kcnk5b-GFP (blue) or a mutated Kcnk5bMut-GFP (pink) that displayed virtually no channel exercise. (D) Diagram for T- and L-type Ca2+ channel inhibition by verapamil or mibefradil. (E) qRT-PCR of SHH expression in HEK293 cells after remedy with verapamil on the indicated concentrations. (F) qRT-PCR of SHH expression in HEK293 cells after remedy with Mibefradil on the indicated concentrations. (G) Brightfield photos of HEK293 cells transfected with CMV-mCherry and handled with DMSO (a) or transfected with CMV-Kcnk5b-mCherry and handled with DMSO (b) or with verapamil, a L-/T-channel inhibitor, at 50 μm (c), 100 μm (d), 140 μm (e), 200 μm (f). (H) Fluorescence of HEK293 cells transfected with CMV-mCherry and handled with DMSO (a), or transfected with CMV-Kcnk5b-mCherry and handled with DMSO (b) or with verapamil at 50 μm (c), 100 μm (d), 140 μm (e), 200 μm (f). (I) Brightfield photos of HEK293 cells transfected with CMV-mCherry and handled with DMSO (a) or transfected with CMV-Kcnk5-mCherry and handled with DMSO (b) or with Mibefradil, a L-/T-channel inhibitor, at 3.5 μm (c), 5 μm (d). (J) Fluorescent photos of HEK293 transfected with CMV-mCherry and handled with DMSO (a) or transfected with CMV-Kcnk5b-mCherry and handled with DMSO (b) or with Mibefradil at 3.5 μm (c) 5 μm (d). (Ok) Brightfield photos of HEK293 cells transfected with CMV-GFP and handled with DMSO (a) or transfected with CMV-Kcnk5b-GFP and handled with DMSO (b) or with the IP3 receptor inhibitor 2-APB at 30 μm (c), 75 μm (d), 105 μm (e), 150 μm (f). (L) Fluorescence photos of HEK293 cells transfected with CMV-GFP and handled with DMSO (a) or transfected with CMV-Kcnk5b-GFP and handled with DMSO (b) or the IP3 receptor inhibitor 2-APB at at 30 μm (c), 75 μm (d), 105 μm (e), 150 μm (f). Every experiment was repeated at the least 3 instances (N = 3) and every repeat contained duplicate or triplicate samples. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values ≥0.05 are designated as “not vital” (NS). Scale bars equal 20 μm (B–E, M, O) and 100 μm (F–Ok). Numerical information used on this determine are included in S13 Data.
https://doi.org/10.1371/journal.pbio.3002565.s022
(TIF)
S9 Fig. Results of drug therapies on tradition cells and embryos.
(A) qRT-PCR for SHH in HEK293 cells transfected both with GFP or Kcnk5b-GFP and handled with DMSO or the CaMKII,IV inhibitor KN-62 on the indicated concentrations. (B) Brightfield photos of HEK293 cells transfected with CMV-mCherry and handled with DMSO (a) or transfected with CMV-Kcnk5b-mCherry and handled with DMSO (b) or handled with KN-62 an inhibitor for CaMKII and CaMKIV at 2 μm (c) and three μm (d). (C) Fluorescence photos of HEK293 cells transfected with CMV-mCherry and handled with DMSO (a) or transfected with CMV-Kcnk5b-mCherry and handled with DMSO (b) or handled with KN-62 an inhibitor for CaMKII and CaMKIV at 2 μm (c) and three μm (d). (D) Brightfield photos of cells handled transfected with CMV-GFP and handled with DSMO (a) or transfected with CMV-Kcnk5b-GFP and handled with DSMO (b) or STO-609 an inhibitor of CaMKK at 0.2 μm (c), 0.5 μm (d), 1 μm (e), 1.5 μm (f). (E) Fluorescent photos of cells handled transfected with CMV-GFP and handled with DSMO (a) or transfected with CMV-Kcnk5b-GFP and handled with DSMO (b) or STO-609 an inhibitor of CaMKK at 0.2 μm (c), 0.5 μm (d), 1 μm (e), 1.5 μm (f). (F) Consultant 48 hpf kcnk5b transgenic Tg[hsp70:kcnk5b-GFP] embryo 12 h submit warmth shock and 6 h remedy in solvent focus of DMSO. (G) Enlarge view of kcnk5b-Tg embryo (F) exhibits some coronary heart edema related to heat-shock-induced expression of kcnk5b. (H) Consultant 48 hpf kcnk5b transgenic embryo 12 h submit warmth shock and 4 h remedy within the IP3R inhibitor 13 μm 2-APB. (I) Enlarge view of kcnk5b transgenic embryo in (H). (J) Consultant 48 hpf kcnk5b transgenic embryo 12 h submit warmth shock and 6 h remedy in 24 μm STO. (Ok) Enlarge view of kcnk5b transgenic embryo in (J). (L, M) Cells transfected with CMV-camkk1b-mCherry in consultant brightfield (L) and fluorescence (M) photos. Every experiment was repeated at the least 3 instances (N ≥ 3) and every repeat contained duplicate or triplicate samples. P values symbolize statistical evaluation by Pupil’s two-tailed t check. P values ≥0.05 are designated as “not vital” (NS). Scale bars equal 20 μm (D–I, Ok, M, N, P, Q, M, O) and 100 μm (R–W). Numerical information used on this determine are included in S14 Data.
https://doi.org/10.1371/journal.pbio.3002565.s023
(TIF)